5
Biological Utilization of CO2 into Chemicals and Fuels
Harnessing the natural ability of microorganisms to capture and convert CO2 into chemicals and fuels has great potential for utilization of gaseous carbon waste. Because biological processes have unique carbon utilization resource requirements and product opportunities, these can be seen as complementary to chemical and mineral carbonation options. This chapter addresses opportunities, challenges, and research needs relevant to utilization technologies that convert CO2 into various products through photosynthetic and nonphotosynthetic means.
PHOTOSYNTHETIC APPROACHES TO CARBON DIOXIDE UTILIZATION
Approaches Based on Algae
While nonphotosynthetic processes for carbon waste gas utilization can produce high biomass yields, these systems suffer from poor life-cycle analyses when compared to photosynthetic methodologies (Vieira et al., 2013). Among photosynthetic processes, algal biomass provides high yields, with more than 30- to 50-fold improvement in oil yield in comparison to common agricultural crops (Singh and Gu, 2010). The term algae broadly refers to any photosynthetic prokaryotic microorganism (cyanobacteria) or eukaryotic microorganism (microalgae) that can be cultivated. Algae cultivation was introduced in the 1950s and has been practiced both academically and commercially for decades (Fisher and Burlew, 1953; Golueke and Oswald, 1959). Algae can be viewed as self-replicating machines that convert sunlight and CO2 into value-added products. Products of algae cultivation include a wide range of biofuels, dietary protein and food additives, commodities, and specialized chemicals.
Algal biofuels have long been seen as a potentially viable strategy for addressing dwindling petroleum reserves, and investments in algal biofuel technologies have tended to rise when crude oil prices rise. For example, the U.S. Department of Energy (DOE) Office of
Fuels Development funded the Aquatic Species Program, focused on the production of biodiesel from microalgae, from 1978 to 1996 (Sheehan et al., 1998), and in 2008 DOE renewed its support for algal biofuel research after crude oil prices topped $100/barrel. This most recent spike in research investment coincided with large investments in algal biofuel technologies by startups and established companies; however, a subsequent decrease in crude oil prices to below $30/barrel in 2016 created a major setback to industrial efforts. There are a wide range of estimates for the cost of production for algal biofuels (NRC, 2012) and it is difficult to predict how future policy and other factors may influence their competitiveness with traditional fossil resources; however, one challenge of algal cultivation stems from the cost and development of infrastructure for physical processing, such as dewatering and extraction, that continue to pose challenges for energy demand (Sander and Murthy, 2010). One obstacle to the growth of these methods will be adoption of support and infrastructure commonly available to other forms of domestic agriculture (Trentacoste et al., 2015). Multiple reports describe the state of the science and research challenges for algal biofuels (DOE BETO, 2016, 2017; DOE EERE, 2010, 2017; NRC, 2012; Sheehan et al., 1998).
Algae cultivation offers significant productivity improvements as compared to contemporary agriculture. Multiple studies have demonstrated up to 30-fold productivity improvements in oil production (Abishek et al., 2014; Wen and Johnson, 2009) and 50-fold improvements in protein production from algae when compared to soybean, canola, or corn (Bleakley and Hayes, 2017) per acre of land. Although not fully evaluated, protein derived from algal biomass has been proposed for both animal and human consumption, and some companies, for example, Qualitas,1 already produce specialized algae to enrich animal feeds. The competitive landscape for algal cultivation would shift should these commercial applications prove competitive with current agricultural products such as soy or corn.
Despite these advantages, there are also significant challenges to algae cultivation. Photosynthesis is inherently inefficient, as only 3-6 percent of total solar radiation energy is captured (Zelitch, 1975). In addition, a limitation of algae is the efficiency of the carbon dioxide conversion that limits the annual flux (Wilcox, 2012). To compensate for this low efficiency, cultivation ponds or bioreactors are often designed to maximize light exposure through large volume and surface area; as a result, algal cultivation can be land and water intensive. Theoretically, to capture all CO2 from a 10 kiloton/day power plant would require 25-37 acres of cultivation (Hazelbeck, D., personal communication, 2018). In addition, biomass cultivation presents capital and operations costs that do not have close parallels in existing large-scale industries.
On the other hand, a key benefit to biological systems is their inherent flexibility in terms of feedstocks and environments. There are opportunities to utilize nonarable land and saline water or wastewater for algae cultivation, thereby minimizing competition for natural
___________________
resources. In addition, biological systems can tolerate low CO2 concentrations and impurities in the carbon sources common to industrial power generation. These variables have a considerable impact on capital and operations costs for biological conversion technologies, and it is important to account for resource use and environmental impact when comparing algae cultivation to conventional agriculture or other activities.
Approaches Based on Green Algae
Green algae, a common term for eukaryotic single cellular photosynthetic organisms deriving from several phyla, are a highly diverse group of algae representing many thousands of identified species. Multiple strains have been validated in biomass applications, and their selection is commonly determined by culture conditions. For instance, algae from cold environments have adapted to these environments through modification of primary metabolism and physiology (Morgan-Kiss et al., 2006).
The majority of biomass cultivation efforts using green algae have focused on biofuel production. In addition, several co-products have been identified that can help make biofuel production more economically feasible. The following sections describe opportunities for using green algae to produce various biofuels and co-products, as well as opportunities and limitations involved in advancing such applications through genetic manipulation.
Biodiesel Production
Biodiesel, or fatty acid methyl ester, production from plant- and animal-based fats and oils (including those from green algae) is now considered part of a mature industry. While biodiesel can be used as drop-in diesel replacement in many diesel engines, its hygroscopicity, cloud point, and fouling properties have limited widespread adoption. Once considered an ultimate product of biomass production, biodiesel has more recently become relegated to small fuel markets and as an additive for ultra-low-sulfur petroleum diesel to provide added lubricity properties (Hazrat et al., 2015).
After biomass harvesting, lipids are extracted by one of several methods to prepare isolated neutral lipids, or triacylglyceride (TAG), or total lipids containing both polar and neutral lipids. The traditional process for biodiesel production from plant- and animal-based fats and oils typically requires a high-purity source of TAG; however, methods have been developed to produce biodiesels from total lipid extracts (Asikainen et al., 2015; Mubarak et al., 2015).
Renewable Diesel/Gasoline Production
The most broadly adoptable advances in algae biomass conversion to biofuels lies in the catalytic hydrogenation of lipid extracts, known as hydrotreatment, for the production of true diesel and gasoline components. Although the methods for these transformations have been
known for decades, the capital infrastructure required to carry out hydrotreatment on a commercial scale has limited its implementation to a few refining companies, such as Neste Oil and Chevron (Al-Sabawi and Chen, 2012; No, 2014).
One particular benefit of renewable diesel and gasolines is the flexibility with regard to the source of feedstock. Currently, industrial hydrotreatment processing can utilize fats and oils from any bioderived triacylglyceride source, including vegetable oils and animal fats. However, while algal lipids do contain triacylglycerides, extracts from many species also contain significant portions of polar lipids and hydrophobic pigments. New technology may be required for conversion of these species into renewable fuels by hydrotreatment facilities (Davis et al., 2013). The requirement of hydrogen gas for the catalytic hydrogenation of lipid extracts must also be considered.
In addition, liquefaction, pyrolysis, and gasification offer routes to produce a variety of fuels, including methane, ethanol, and fuel oils (Demirbas, 2010). Many of these have been proposed to be routed to more complex chemicals through reforming processes (Bhujade et al., 2017).
Valorization of Co-Products
In order to compete with fossil fuels, biofuels must provide attractive economics. This can be achieved by increasing the cost of petroleum products through pricing nonrenewable carbon (or other mechanisms), or the value of algae cultivation can be further increased by production of additional products. Biomass conversion to biofuels inherently creates waste products from unused biomass, primarily composed of protein and carbohydrates. These can be consumed by anaerobic digestion to produce biogas, a low-value end product. Or, they can be converted into a variety of more valuable products. Combined approaches that utilize photosynthetic, fermentative, and chemical methods can be employed to produce products with high value. For instance, a combined algal processing method capable of producing multiple products from algae biomass was recently demonstrated (Miara et al., 2014). In such systems, algal carbohydrates can be diverted to fermentative production of ethanol or other commodity chemicals, such as succinic acid (Raab et al., 2010) from metabolically engineered yeast, while the proteins may be applied to adhesive manufacturing (Roy et al., 2014). Examples of co-products that may be valorized to improve the economics of biofuel production from algae include dietary protein, polyunsaturated fatty acids, and pigments.
Dietary protein. Algae have long been viewed as an attractive source of dietary protein, both as animal feeds and for human consumption (Becker, 2007). Green algae contain between 40 and 70 percent of their dry weight as protein, and the amino acid profile of most algae compares favorably with common food proteins. A few studies have been performed to determine algae’s protein efficiency ratio (PER), which reflects an animal’s weight gain per unit
of protein consumed; one study found algae offered up to 80 percent PER compared to milk proteins (Becker, 2004) while incurring a substantially smaller footprint than milk production in terms of freshwater resources and arable land (Bleakley and Hayes, 2017). Protein productivity from algae has been estimated at up to 50 times that of soybeans per acre of land. This suggests green algae has the potential to supplement or replace crops for animal feed with vastly reduced demands for arable land, but studies have not been validated at scale.
Polyunsaturated fatty acids. Many green algae naturally produce polyunsaturated fatty acids (PUFAs) that are valuable for humans and animals as food additives. While they are typically harvested from fish, PUFAs from algae represent a viable and more sustainable set of target molecules (Adarme-Vega et al., 2012). In particular the omega-3 varieties eicosapentaenoic acid and docosahexaenoic acid are valuable for the supplementation of many farmed animals, particularly carnivorous fish such as salmon and tuna (Ahmed et al., 2012; Bimbo, 2007). Omega-3 feeding studies have also been performed on cattle and poultry (Nitsan et al., 1999; Ponnampalam et al., 2006). In addition, omega-3 fatty acids demonstrate a consumer market as neutraceutical products that is currently dominated by fish oils.
Pigments. Colorants are manufactured as additives to foods, drinks, cosmetics, and a host of other products in order to increase the appeal of the product to the consumer. Some pigments are manufactured utilizing fossil fuel sources such as coal. Algae are well known for their ability to produce a variety of pigments and could provide a more sustainable alternative to the utilization of fossil fuels (Kaur et al., 2009). One well-known example of the successful application of algae for the commercial production of pigments is astaxanthin (see Box 5-1). The manufacture of other pigments with algae remains a potential opportunity for further exploration.
Genetic Manipulation of Green Algae
The tools to facilitate genetic manipulation of green algae are far less advanced than tools for genetic manipulation of bacteria, yeast, and vascular plants. Key challenges include poor genome insertion, gene silencing, and unoptimized promoter systems. Nevertheless, some academic and industrial groups have attempted genetic modification of green algae for the purposes of improving photosynthetic efficiency, decreasing photodamage of the light harvesting complex, and optimizing the efficiency of carbon uptake and incorporation. Several products have also been sought from transgenic expression, including high-value therapeutic proteins, biohydrogen, lipids, and terpenoids (Gimpel, 2013).
Many species of green algae are genetically tractable, but Chlamydomonas reinhardtii has become the primary model system, with the majority of tools and techniques developed for this species (Rasala et al., 2013). Green algae have three separate genomes: nuclear,
chloroplast, and mitochondrial. In C. reinhardtii, each of these genomes has been sequenced, and tools and protocols for genetic manipulation have been developed for each. Commonly, nuclear and chloroplast genomes have been primarily targeted for transgene expression, and each location involves special tools, advantages, and challenges. Engineering the nuclear genome for protein production offers benefits similar to other eukaryotic expression systems, including regulated expression, cytoplasmic protein folding machinery, access to posttranslational modification (such as glycosylation), and protein secretion. Furthermore, like many green algae, C. reinhardtii is a haploid organism. As a result, nuclear genome modifications can be used in combination with sexual breeding to provide additional genetic control. However, nuclear transformation currently suffers from several drawbacks, including random gene insertion, low protein expression, and gene silencing (Cerutti et al., 1997; De Wilde et al., 2000; Wu-Scharf, 2000). These complications are being actively researched, and some limited solutions have been recently developed. For instance, gene silencing has been addressed by directly coupling transgene expression with antibiotic resistance gene expression, therefore imparting a strict selection method for gene retention (Rasala, 2012, 2013). In addition, experiments employing new technologies for gene integration and knockout, including CRISPR methodologies, are currently under way (Ferenczi et al., 2017; Shin et al., 2016). Nevertheless, nuclear expression remains limited in terms of gene size, number, and complexity.
Tools and methodologies for manipulating gene expression in the chloroplast have achieved significant advances in recent years. Unlike nuclear expression, chloroplast expression allows homologous recombination, and multiple vectors, selection, and transformation methods have been developed. The green algae chloroplast offers an environment for protein expression, including chaperones and posttranslational modification enzymes that can facilitate proper protein and expression folding, and heterologous protein expression has achieved remarkable levels, sometimes as high as 10 percent of total soluble protein (Mayfield, 2007). Despite these benefits, chloroplast expression remains limited in terms of gene size and number, challenging the needs for engineering large operons and metabolic pathway design.
In the broader scheme of genetic engineering tools, photosynthetic organisms have historically received a low level of research support, and algae represent a small subset of this group (Gao et al., 2012). However, tools developed in recent decades underscore the potential for green algae as an ideal photosynthetic host organism. New and improved genetic tools for green algae genetic manipulation remain a key need for the advancement of algae biomass applications. This includes development of genetic insertion (homologous recombination) technologies, identification of robust promoters for gene expression, development of synthetic operons for multiple gene incorporation, tools for engineering large genes and pathways, and novel selection methods.
Approaches Based on Cyanobacteria
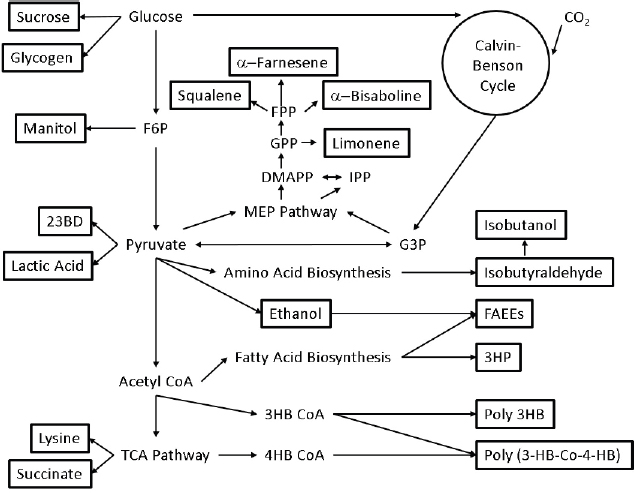
Cyanobacteria are being engineered to directly convert solar energy, carbon dioxide, and water to biofuels and other products. Cyanobacteria-based approaches possess advantages over traditional biological production systems based on plants, green algae, or heterotrophic organisms. For example, cyanobacteria are more amenable to genetic manipulation than are algae and can therefore be adapted for the production of a wider range of products (see Figure 5-1). The photosynthetic efficiency of cyanobacteria is two to four times higher than that of plants (Melis, 2009), and their cultivation does not compete with food crops for land usage. Cyanobacteria also do not have the sugar requirements of heterotrophic organisms such as Escherichia coli and yeast, although they have a reduced growth rate and fewer available synthetic biology tools compared to these hosts. While heterotrophic hosts may be adapted to utilize cellulosic feedstocks these technologies are still not well developed and as such efficiency can be low. Cyanobacteria allow us to skip this carbon-harvesting step and fix CO2.
There is a plethora of cyanobacterial species but only a few have been adapted to chemical production. The three predominant strains utilized for chemical production are Synechococcus elongatus PCC 7942 (7942), Synechocystis sp. PCC 6803 (6803), and Synechococcus sp. PCC 7002 (7002). These strains all have sequenced genomes, established culturing methods, and basic metabolic engineering tools (Berla et al., 2013; Markley et al., 2015; Nozzi et al., 2017; Yu et al., 2015), yet each strain presents its own unique advantages and challenges.
__________________
1 Reprinted from Carroll, Austin L., Anna E. Case, Angela Zhang, and Shota Atsumi. 2018. “Metabolic engineering tools in model cyanobacteria.” Metabolic Engineering. doi:10.1016/j.ymben.2018.03.014 with permission from Elsevier.
7942 is a freshwater unicellular cyanobacterium and studied for photosynthesis and genetic manipulation (Golden et al., 1987). 7002 is a marine species capable of growth in a variety of conditions (salt, temperature, and light) (Batterton and Van Baalen, 1971), making it more apt to utilize natural saltwater resources. However, metabolic engineering has not been as well studied in 7002 as in freshwater species and synthetic biology capabilities for 7002 lag behind that of 7942 and 6803. 6803 has a good selection of genetic tools and can grow photomixotrophically.
Cyanobacteria have been engineered to produce a wide range of fuels, fuel precursors, and commodity chemicals. The productivities and titers (mostly on the order of milligrams per liter) are generally too low to make commercialization of the technology appealing, however, and these technologies have been demonstrated primarily on small, academic research scales. To move these technologies toward commercialization, it will be critical to reduce the cost of production and further advance tools for genetic manipulation and metabolic engineering of cyanobacteria.
Fuel Production
Fuels and fuel precursors that can be produced with cyanobacteria include ethanol, butanol, fatty acids, heptadecane, limonene, and bisabolene.
Ethanol. Ethanol, a common biofuel candidate, has been produced in cyanobacteria by the transfer of two ethanol pathway genes from yeast. This production has been enhanced to form 5.5 g/L ethanol in Synechocystis sp. PCC 6803 (Dexter and Fu, 2009; Gao et al., 2012).
Butanol (n-butanol and isobutanol). Isobutanol has applications as a drop-in biofuel that could be integrated into current energy infrastructure. Production can be achieved by expressing heterologous 2-ketoisovalerate decarboxylase and aldehyde reductase that redirect carbon flux from the L-valine biosynthetic pathway to isobutanol resulting in a titer of 0.5 g/L in 7942 (Atsumi et al., 2009). n-butanol, which also has uses as a drop-in biofuel and is currently used in a variety of consumer products, can be produced in 7942 (Lan and Liao, 2011). The n-butanol biosynthetic pathway was constructed based on the natural n-butanol biosynthetic pathway found in Clostridium acetobutylicum. Examination and characterization of Coenzyme A-acylating (CoA) aldehyde dehydrogenases that are oxygen tolerant enabled production of butyraldehyde from butyryl-CoA. Final titers of 404 mg/L were achieved with productivity of 2 mg/L/h (Lan et al., 2013).
Fatty acids. Fatty acids can be used in the synthesis of biodiesels. Cyanobacteria natively produce free fatty acids in trace amounts. This production has been improved in 7942, 6803, and 7002. Free fatty acid overproduction leads to reduced cell fitness. Free fatty acid production in 7002 results in reduced negative impacts on the cell, and production can be further improved by the nonnative expression of RuBisCO leading to >130 mg/L free fatty acids being produced (Ruffing, 2014). When coupled with native free fatty acid production, the expression of a heterologous diacylglycerol acyltransferase in tandem with an ethanol production pathway results in a variety of fatty acid ethyl esters (FAEEs) in 7942. This platform leads to the accumulation of up to 7 mg/L of FAEEs (Lee et al., 2017a).
Heptadecane. Long-chain hydrocarbons such as heptadecane (see Table 5-1) can be readily used in fuel production for the synthesis of biodiesel. Heptadecane production can be achieved by capitalizing on the natural fatty acid production in the cell. Titers can be improved in strains that natively produce alkanes by overexpressing an acyl-ACP reductase/aldehyde-deformylating oxygenase (AAR/ADO) pathway. Expression of the AAR/ADO pathway from 7942 in a marine cyanobacterium results in up to 4.2 µg/g dry cell weight heptadecane (Yoshino et al., 2015).
TABLE 5-1 Summary of commodity chemicals and fuels that are currently synthesized from CO2 from cyanobacteria and their reported production.
| Compound | Strain | Reported Production |
|---|---|---|
| Ethanol | Synechocystis sp. PCC 6803 | 5.5 g/L |
| Isobutanol | Synechococcus elongatus PCC 7942 | 0.5 g/L |
| n-Butanol | Clostridium acetobutylicum | 2 mg/L/h |
| Fatty acids | Synechococcus elongatus PCC 7942 Synechocystis sp. PCC 6803 Synechococcus sp. PCC 7002 |
>130 mg/L |
| Synechococcus elongatus PCC 7942 | 4.2 µg/g dry cell weight | |
 |
Synechococcus elongatus PCC 7942 Synechococcus sp. PCC 7002 Synechocystis sp. PCC 6803 |
1 mg/L/OD730/day 4 mg/L 7 mg/L |
 |
Synechococcus sp. PCC 7002 | 0.6 mg/L |
 |
Synechococcus elongatus PCC 7942 | 2.4 g/L (after 21 days) 3 g/L (after 10 days, photomixotrophic conditions) 12.6 g/L (continuous lighting with glucose and CO2) 5.7 g/L (23-hour light cycling) |
 |
Synechococcus elongatus PCC 7942 | 0.3 g/L |
| Compound | Strain | Reported Production |
|---|---|---|
| Ethylene | Synechocystis sp. PCC 6803 | 2.5 mL/h/OD730 |
| Glycogen | Synechococcus sp. PCC 7002 | 3.5 g/L (after 7 days) |
| Lactate | Synechocystis sp. PCC 6803 Synechococcus elongatus PCC 7942 |
0.8 g/L (after 2 weeks) 1.4 g/L (after 10 days) |
 |
Synechococcus elongatus PCC 7942 Synechocystis sp. PCC 6803 |
0.8 g/L (after 6 days) |
| Isoprene | Synechococcus elongatus PCC 7942 | 1.3 g/L (after 21 days) |
 |
Synechococcus elongatus PCC 7942 | 50 mg/L |
 |
Synechococcus elongatus PCC 7942 | 5 mg/L |
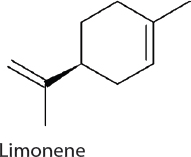
Limonene. Limonene has applications as both a biofuel and a solvent. Limonene can be produced with the expression of a single heterologous gene for limonene synthase (LS) selected from spearmint for its high selectivity for limonene production. The expression of LS capitalizes on carbon flow through the native methyl-D-erythritol 4-phosphate (MEP) pathway, commonly found in plants, and funnel flow to limonene (Wang et al., 2016a). Establishing high LS expression is critical in improving titers. Limonene production has been established in 7942 and 7002 at 1 mg/L/OD730/day (Wang et al., 2016a) and 4 mg/L (Davies et al., 2014), respectively. In 6803, carbon flux through the oxidative pentose phosphate (OPP) pathway is used to drive limonene production. Overexpression of two native enzymes
(ribose-5-phosphate isomerase and ribulose 5-phosphate 3-epimerase) and expression of a heterologous geranyl diphosphate synthase gene establishes limonene production at 7 mg/L (Lin et al., 2017).
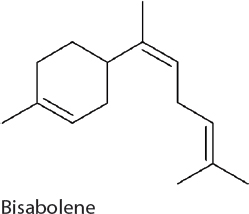
Bisabolene. Bisabolene has applications as a biodiesel candidate and production relies on carbon flow through the MEP pathway and the expression of a single heterologous gene, (E)-α-bisabolene synthase. Production has been established in 7002 resulting in 0.6 mg/L (Davies et al., 2014).
Commodity Chemicals Production
Commodity chemicals that can be produced with cyanobacteria include 2,3-butanediol, 1,3-propanediol, ethylene, glycogen, lactate, 3-hydroxypropanoic acid, 3-hydroxybutanoic acid, 4-hydroxybutanoic acid, isoprene, and farnesene.
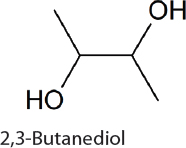
2,3-Butanediol. 2,3-Butanediol is a commodity chemical used to make synthetic polymers (van Haveren et al., 2008) and can readily be converted to methyl ethyl ketone, a fuel additive and solvent (Tran and Chambers, 1987). 2,3-Butanediol relies on carbon syphoned from central metabolism in the form of pyruvate. The condensation of two pyruvate molecules, followed by decarboxylation and reduction through heterologously expressed genes, results in 2.4 g/L 2,3-butanediol after 21 days of production (Oliver et al., 2013). 2,3-Butanediol was similarly produced in 6803 (Savakis et al., 2013) and 7002 (Nozzi et al., 2017). In 7942 the addition of glucose and expression of galP, encoding a hexose symporter, allows growth using both CO2 and glucose (McEwen et al., 2013). This modification permits continued 2,3-butanediol synthesis in both light and dark conditions, a potentially useful strategy for 24-hour metabolite production. Under photomixotrophic conditions 3 g/L 2,3-butanediol can be produced over 10 days (McEwen et al., 2016). Global metabolic rewiring of these strains for improved CO2 fixation increases 2,3-butanediol titers, glucose utilization efficiency, and carbon fixation in both lighted and diurnal conditions (Kanno et al., 2017). Under continuous lighting, rewired 7942 produces 12.6 g/L 2,3-butanediol (Kanno et al., 2017) when fed glucose and CO2. Under natural 24-hour (diurnal) light cycling in the rewired strain, 5.7 g/L 2,3-butanediol is produced.
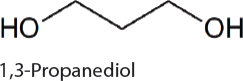
1,3-Propanediol. 1,3-Propanediol has a variety of uses including in polymers, paints, solvents, and antifreeze. Production in cyanobacteria is dependent on removal of carbon from the Calvin-Benson cycle in the form of dihydoxyacetone phosphate (DHAP). DHAP can be converted to 1,3-propanediol via a four-step heterologously expressed metabolic pathway. After the disruption of production bottlenecks 0.3 g/L 1,3-propanediol was produced in 7942 (Hirokawa et al., 2016).
Ethylene. Ethylene can be used to generate polyethylene, a widely used polymer, and in its gaseous state can be used to speed the ripening of produce. Production has been established in 6803 and relies on the expression of a single heterologous gene, efe, encoding ethylene forming enzyme (Efe). Efe converts 2-oxoglutarate from the tricarboxylic acid (TCA) cycle to ethylene. Partial deletion of ntcA, encoding the global transcription factor known for regulating carbon metabolism (Mo et al., 2017), exhibits a 23 percent increase in ethylene production and 1.5-fold increase in Efe activity in 6803 (Mo et al., 2017). Ethylene production in this strain reached 2.5 mL/h/OD730 (Mo et al., 2017).
Glycogen. Glycogen is a common form of energy storage and a potential carbon source for chemical production. It can also be converted into ethanol for use as a biofuel. 7002 naturally amasses glycogen under nitrogen-depleted conditions. Glycogen production can be improved by modifying culture conditions, including light intensity, CO2 concentration, and salinity (Aikawa et al., 2014). Improved production conditions result in 3.5 g/L glycogen after 7 days (Aikawa et al., 2014).
Lactate. Biological synthesis of lactic acid for biodegradable polymers has been established in 6803. Lactic acid production requires the heterologous expression of a single enzyme, lactate dehydrogenase, which pulls pyruvate from central metabolism to catalyze the conversion to lactic acid at a titer of 0.8 g/L after 2 weeks of cultivation (Angermayr et al., 2014). Further optimization of co-factor requirements leads to approximately 1.4 g/L lactate produced after 10 days in 7942 (Li et al., 2015).
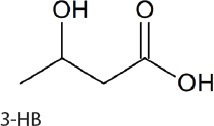
3-Hydroxypropionic acid, 3-hydroxybuterate, and 4-hydroxybuterate. 3-Hydroxypropionic acid (3-HP), 3-hydroxybuterate (3-HB), and 4-hydroxybuterate (4-HB) (Table 5-1) all have applications for the synthesis of polymers and plastics used in daily life. 3-HB was produced in 6803 and 7942 by the expression of a nonnative malonyl-CoA reductase, which converts malonyl-CoA to 3-HP. Malonyl-CoA natively feeds into fatty acid biosynthesis. This production can be aided by streamlining carbon flux to malonyl-CoA, expressing malonyl-CoA reductase, and optimizing nicotinamide adenine dinucleotide phophaste (NADPH) levels, resulting in 0.8 g/L after 6 days (Wang et al., 2016b). 7002 has been demonstrated to produce poly-3-hydroxybutyrate and poly-3-hydroxybutyrate-co-4-hydroxybutyrate through the introduction of a gene cluster from Chlorogloea fritschii PCC 9212 (Zhang et al., 2015). 3-HB production relies on native acetyl-CoA production, which can be converted to 3-HB in two consecutive steps. Alternatively, 4-HB can be produced by pulling carbon from the TCA cycle in the form of succinic semialdehyde. 3-HB can be produced by itself or in combination with 4-HB to create a copolymer. Production can reach 0.05 g/L of 3-HB or 4.5 percent total cell dry weight of the copolymer, with 4-HB accounting for 12 percent of the copolymer (Zhang et al., 2015).
Isoprene. Isoprene is commonly used for the production of synthetic rubbers. Overexpression of the native MEP pathway and the expression of plant isoprene synthase gene (ispS) in 7942 results in the production of isoprene. With this production platform 1.3 g/L isoprene was produced after 21 days (Gao et al., 2016).
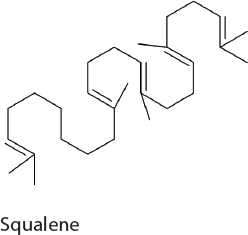
Squalene. Squalene is widely used in the food, personal care, and medical industries but its commercial production is unreliable and nonideal. Squalene production has been previously established in heterotrophic hosts, such as yeast. Similarly, squalene was produced in 7942 by the overexpression of the native MEP pathway in addition to the expression of a heterologous squalene synthase from yeast (Choi et al., 2017). Following optimization in carbon flux, the expression of this pathway results in 50 mg/L squalene.
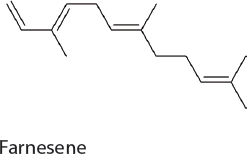
Farnesene. Farnesene has been used as a precursor for high-performance polymers and as a jet fuel candidate. Production in 7942 was achieved by the heterologous expression of a single gene encoding farnesyl synthase, which funnels carbon out of the MEP pathway in the form of farnesyl diphosphate. The engineered strain produced 5 mg/L of α-farnesene (Lee et al., 2017b).
Genetic Manipulation of Cyanobacteria
While some target fuels and chemicals can be produced through exploitation of naturally occurring cyanobacteria traits, manipulation of organisms’ genetic material can enable production of many other target molecules. An understanding of areas where desired genetic material may be inserted without interrupting essential functions (gene integration and plasmids), ways to control production of the desired chemical target (promoters and riboswitches), and ways to tell the cell where to start and stop generating the target molecule (ribosomal binding sites) are necessary in order to conduct genetic manipulations. CRISPR is another tool for reliable insertion of desired genetic material into host cells.
In addition to an enhanced ability to introduce and control novel pathways in cyanobacteria, a thorough understanding of the flow of carbon through the cell’s metabolism is key to creating an industrially relevant production system. Techniques for improving carbon flux have included the use of carbon sinks, disrupting side pathways, removing inhibitors, and protein fusions to improve rate. While these techniques have pushed cyanobacteria toward industrial relevance there is still much that is unknown and needs further study.
Metabolic Engineering of Cyanobacteria
One of the primary challenges of chemical production in cyanobacteria is the low titers that result from most pathways. Production is usually limited to the milligram per liter (mg/L) order. A great deal of effort has gone into engineering RuBisCO, the primary carbon
capture mechanism in cyanobacteria. While some improvements have been made (Whitney et al., 2011), RuBisCO has proven very difficult to improve, and alterations still remain below the required threshold for wide-scale chemical production.
An alternative to RuBisCO engineering is the supplementation of cyanobacteria with alternate carbon sources including glucose, glycogen, acetate, and xylose. While these carbon sources effectively improve titers, this additional carbon is not always directed toward chemical production and is often lost as biomass. Careful rewiring of carbon integration into metabolism can help improve this carbon utilization and act as a carbon sink to improve CO2 fixation (Oliver and Atsumi, 2015; van der Woude et al., 2014). However, this approach requires a detailed understanding of carbon metabolism in the cell and how an alternate carbon source is being incorporated. In addition, these alternate carbon sources can increase the risk of contamination. As a slow-growing organism, cyanobacteria in culture can quickly become overcome by competing organisms that can utilize sources such as glucose. The necessity of using stricter techniques to prevent contamination, combined with the added cost of the alternate carbon source, can significantly raise production costs and eliminate some of the advantages of using cyanobacteria as a host strain.
In order to optimize carbon flow, researchers must fully understand it. Genome-scale models (GSMs) are important tools for assessing and engineering metabolic systems. The models may be used to describe an organism’s entire metabolism utilizing genomic information (Broddrick et al., 2016; Kim et al., 2017; Shirai et al., 2016; Triana et al., 2014). It would be challenging to optimize production of a target chemical, identify bottlenecks, and target the best production system for the host without GSM. GSM-directed engineering has been successfully used to improve a variety of production platforms in E. coli, including 1,4-butanediol (Yim et al., 2011), lycopene (Alper et al., 2005), lactic acid (Fong et al., 2005), and succinate (Lee et al., 2005).
The transfer of GSMs from heterotrophs to photoautotrophs is a difficult process; one reason is that photosynthetic growth complicates simple factors, like energy input, making them complex and difficult to measure. The recent development of two GSMs (Broddrick et al., 2016; Triana et al., 2014) for 7942 allows for greater predictive power when making modifications to metabolism. New insights have been gained using a GSM developed for 7002 (Vu et al., 2013), which could help engineer the strain (Hendry et al., 2016). Based on this model it has been predicted that up to 10 percent of fixed carbon could be rerouted to production (Vu et al., 2013). These predictions have not been confirmed experimentally.
Construction of a comprehensive GSM for 6803 has attempted to resolve many of the problems that are seen in 7942 and 7002 with GSM development in cyanobacteria (Hucka et al., 2003; Knoop et al., 2010; Stanford et al., 2015). However, automated assembly of key GSM components is prevented by incongruent gene annotations and database nomenclature. GSM analysis uses 3,167 genes to study gene function, carbon metabolism, photosynthesis, and chemical production (Shirai et al., 2016; Yoshikawa et al., 2017). These models can iden-
tify deletions of native metabolic reactions that compete for reductive power required in targeted chemical production successfully (Yoshikawa et al., 2017). While traditional GSMs are limited to the native genes found in 6803, recent iterations incorporate nonnative metabolic reactions to construct hybrid phototrophic and heterotrophic cyanobacteria models (Saha et al., 2016; Shirai et al., 2016). These expanded models can predict new strategies for metabolic pathway construction and increase yields in targeted chemical production.
Synthetic biology tools are essential to utilize a strain as a production host. These tools can range from sites for the expression of nonnative genes to systems for controlling gene expression or tools for genetic manipulation (Albers et al., 2015; Immethun et al., 2017; Ueno et al., 2017). Heterotrophic hosts such as yeast and E. coli have well-developed synthetic biology toolboxes; however, these tools are often incompatible with photoautotrophic hosts such as cyanobacteria. In fact, tools developed for cyanobacteria are often difficult to transfer between strains, necessitating the development of a unique set of tools for gene integration and controlling gene expression on a transcriptional and translational level for each strain. Many of these tools have been developed for cyanobacteria (Berla et al., 2013; Golden et al., 1987; Markley et al., 2015). In addition to the difficulty of transferring tools between hosts, it can also be difficult to transfer pathways between host organisms. Gene expression, intermediates, and final products have varying toxicity across strains. Each strain can have a unique codon preference, altered enzyme activities, and constitutive promoters. One of the greatest challenges to metabolic engineering efforts for cyanobacteria is the careful tailoring of tools to each strain.
Key Considerations for Photosynthetic Approaches
Whether they involve the cultivation of green algae or cyanobacteria, photosynthetic approaches to carbon dioxide utilization raise a unique set of considerations relevant to their costs, benefits, environmental impacts, and social acceptability. Key considerations include the impact of the cultivation method used, restrictions in the use of genetically modified organisms, the degree of CO2 solvation, nutrient requirements and downstream burdens, impacts on water and land use, and availability and suitability of CO2 waste streams.
Impact of Cultivation Method
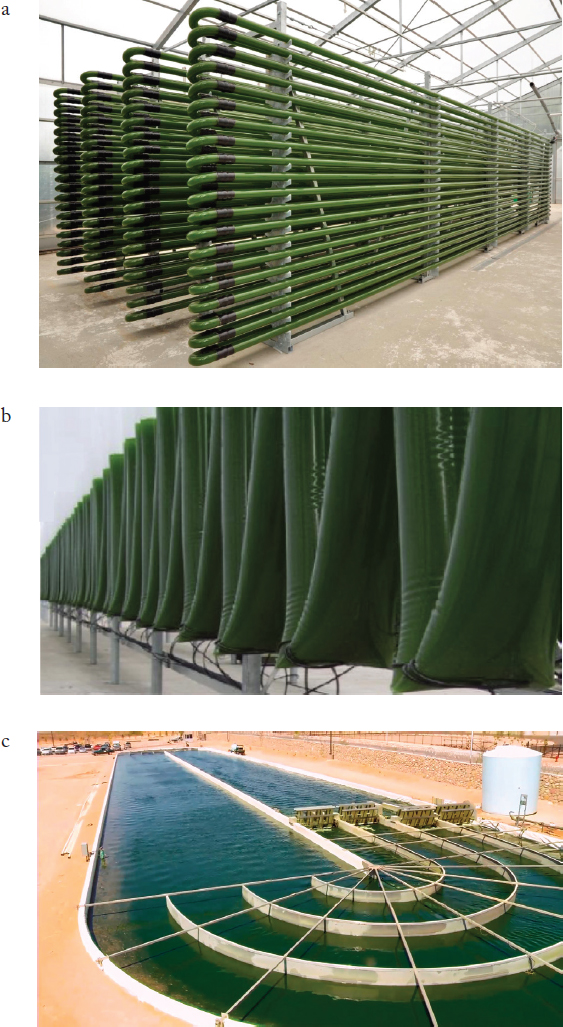
A key consideration for algae biomass production is the selection of cultivation method. Both open ponds and closed photobioreactor (PBR) designs have been implemented on academic and industrial scales (Figure 5-2). Open pond designs commonly employ oblong, “raceway” designs (Benemann et al., 1978) in which a high rate of water movement can be achieved with paddlewheels or air lift pumps. Raceway ponds have proven to be the dominant design, and their design has changed little in three decades, as their construction is believed to

__________________
1 Photo by IGV Biotech.
2 Huang et al., 2017.
3 See https://www.energy.gov/eere/bioenergy/production (accessed October 10, 2018).
be among the least expensive of reactor designs. Raceway ponds can vary in size from several square feet to a hectare or more, and typically employ polymer bed liners and paddlewheel-style impellers. Bed liners are a requirement in some jurisdictions to meet water treatment codes, and the use of any genetically modified species may also mandate such barriers. The cost of bed liners commonly dominates the capital costs of a raceway pond, particularly with increasing size (Rogers et al., 2014).
Closed PBRs can involve a variety of scales, materials, and designs (Gupta et al., 2015), many of which serve to optimize solar radiation upon the biomass culture. The simplest and most common PBR implements a “hanging bag” approach, where polyethylene tubes are suspended vertically with air and CO2 diffusers to provide agitation and carbon addition (Martínez-Jerónimo and Espinosa-Chávez, 1994). Multiple larger system designs have been introduced in recent years using a variety of materials. Inherent in the viability of any PBR system is temperature control, a common challenge of closed systems, and the ability to agitate the culture, typically with pumped liquids.
The capital and operational costs of PBRs are considerably higher than those for open ponds, and the adoption of each technology is dependent on multiple factors (Richardson et al., 2012). Closed PBRs have the benefit of tight control over cultivation conditions, including temperature, CO2 and O2 levels, sun exposure, contaminants, and evaporation. As a result, culture density can reach levels not obtainable in open ponds (Schoepp et al., 2014). The ability to exclude contaminants and exogenous species, including predators, offers additional benefits. As a result, closed PBRs have been favored for the production of high-value products, where product purity is favored over low-cost cultivation. Similarly, open ponds have been favored for the production of biofuels given the need to minimize production costs.
Use of Genetically Modified Organisms
Current U.S. law and regulations tightly constrain the use of genetically modified algae in outdoor open ponds due to concerns regarding containment and spread of genetically modified organisms (GMOs). A limited study was recently performed to evaluate the dispersal, colonization, and impact of GMO algae into native water bodies (Szyjka et al., 2017). The particular study found that while broad dispersal occurs, impact on indigenous species was viewed as negligible. Additional work in this area is needed.
Degree of CO2 Solvation
A major challenge to waste CO2 utilization in all aquatic systems is CO2 solvation. Simple sparging of gases in aqueous solutions results in costly and inefficient solubilization. Two methods have been developed: one utilizes an amine-based CO2 concentrator, followed by thermal stripping, and the second uses carbonate salts to deprotonate solvated CO2 into soluble bicarbonate. The latter method has also been shown to be improved with the use of exogenous
carbonic anhydrase, an enzyme that catalyzes CO2 solvation (Hernandez-Mireles et al., 2014). Each of these methods contains drawbacks, such as thermal input or high pH values. While some strains of algae can adapt to these conditions, more research is required to improve their integration into large-scale biomass cultivation (Könst et al., 2017). Some algal strains produce extracellular carbonic anhydrase to facilitate CO2 solvation (Huertas et al., 2000).
Nutrient Requirements and Downstream Burdens
As with all photosynthetic crops, cultivating algae requires the application of fertilizers such as phosphorus and nitrogen. This has raised concerns that algae cultivation could contribute to eutrophication (excessive algal growth due to the influx of nutrients) in freshwater and coastal zones. Eutrophication is already a serious environmental concern in many areas due to agricultural runoff and municipal wastewater. To prevent algae cultivation from further compounding this problem and even partially address eutrophication, some have advocated the use of municipal and agricultural wastewater as sources of nutrients for algal cultivation (Woertz et al., 2009), thus capturing these nutrients with biomass cultivation before they reach downstream water bodies (Abdel-Raouf et al., 2012; Benemann et al., 2003; Brune et al., 2003). These scenarios would require co-localization of algae cultivation and wastewater treatment, and model pilot studies show significant promise (Bohutskyi et al., 2016; Chekroun et al., 2014). Nutrient recycling is also an important area of research for algal cultivation both as a means to minimize waste and as part of efforts to conserve water use (Rösch et al., 2012).
Impacts on Water and Land Use
Algae biomass cultivation depends upon plentiful sources of water. This poses concerns related to competition for scarce freshwater resources. However, algae are naturally abundant in a variety of environmental conditions, including freshwater, saltwater, brackish water, and a number of extreme environments. Both freshwater and saltwater algal strains have been used for demonstration and pilot-scale programs. Saline groundwater can be sourced throughout the United States. Other viable sources are ocean water and municipal wastewater (Farooq et al., 2015). As a result, freshwater resources need not be threatened by algae cultivation. The recycling, treatment, and disposal of water used for algal biomass has also been a topic of research interest because the life-cycle implications of the type of water used and whether it is recycled are significant (Guieysse et al., 2013; Yang et al., 2011).
Land use is also an important consideration, given that converting CO2 waste from a single power plant would require dozens of hectares of biomass cultivation. Since algae cultivation does not require arable land, it will not compete with agriculture and can valorize regions with marginal or saline soils. Like traditional agriculture, the species selection will depend upon local climate. Endemic algae species have been studied for a variety of climates and environmental conditions.
Availability and Suitability of CO2 Waste Streams
To achieve maximal biomass production, CO2 must be supplied to algae cultivation. For many pilot-scale installations, sourcing CO2 has presented a challenge that has been overcome through on-site compressed CO2 storage and delivery. Economic studies have indicated that co-localization of power generation with biomass production can provide significant advantages (Kadam, 1997; Zeiler et al., 1995). Common CO2 concentrations of flue gas from power generation ranges from 12 to 15 mol%, and these concentrations can be efficiently assimilated by algae biomass. Several pilot-scale facilities have implemented this strategy, whereby flue gas is utilized at the site to provide CO2 to algal cultures (Chen et al., 2012; de Morais and Costa, 2007; Wang et al., 2008). Such co-localization strategies dominate technoeconomic analyses, as transportation of CO2 through pipelines or vehicles would require purification and concentration of CO2 from the source and fail to take advantage of the inherent flexibility of photosynthetic microorganisms to utilize a range of CO2 concentrations.
Biomass utilization of CO2 from industrial power plants provides the potential opportunity for large-scale carbon capture. Natural gas–fired power plants offer a waste stream with low sulfur and nitrogen content. The low levels of NOx and SOx present in the waste stream ultimately can be metabolized by most strains of algae and, as such, require considerations primarily for transport, heat exchange, and time of use (Radmann et al., 2011). Coal-fired power plants, on the other hand, present the challenge of additional contaminants. If not adequately purified, these waste streams can include contaminants known to be algecedic such as arsenic, cadmium, mercury, and selenium (Vocke et al., 1980). In addition to posing challenges for biomass cultivation, trace metals and other contaminants that derive from flue gas may limit potential applications of biomass products. For example, products derived from coal-fired flue gas are likely to contain more contaminants, making these sources better suited for biofuel applications than for the production of protein for animal feed.
NONPHOTOSYNTHETIC APPROACHES TO CARBON DIOXIDE UTILIZATION
Commercializing microbial production has always been an economic challenge due to the high cost of carbon feedstocks and low product yields. While significant work has been done to increase yields in recent years, the need to feed cultures with high-cost sugars is still an issue. Photosynthetic organisms like algae and cyanobacteria mitigate this problem by utilizing CO2 as their feedstocks, but they are slow growing and it is difficult to achieve industrially relevant productivity and scale up the production systems. Nonphotosynthetic organisms that can convert gases like CO2 or methane have become prime targets for microbial production due to their wide diversity of pathways and growth rates. A summary of nonphotosynthetic approaches is shown in Table 5-2.
TABLE 5-2 Summary of nonphotosynthetic approaches to carbon utilization products.
| Product | Route |
|---|---|
| Acetogens | Carbon dioxide fixation |
| Two-state integrated process | |
| Carbon monoxide fixation | |
| Acetate | Direct electron transfer from electrodes to microorganisms |
| Succinate | Direct electron transfer from electrodes to microorganisms |
| Alcohols | Indirect electron transfer via electrochemically synthesized electron donors |
| Pyruvate | Indirect electron transfer via electrochemically synthesized electron donors |
Nonphotosynthetic biological systems possess a number of potential advantages over photosynthetic systems. These include a wide variety of organisms, a larger range of potential target chemicals, and the ability to avoid the inefficiency of photosynthesis. Aerobic systems also have the advantage of high productivity, capacity for continuous cultivation, and compatibility with artificial photosynthesis. Some nonphotosynthetic organisms can take advantage of low-cost, low-emissions electrons from renewable energy sources, and many can be cultivated with cheap and ubiquitous gaseous carbon feedstocks comprised of hydrogen, CO, and CO2, such as industrial waste gas, biogas, or syngas.
There are also significant challenges that need to be addressed in order for technologies based on nonphotosynthetic organisms to reach maturity. These include many of the same challenges associated with photosynthetic systems, such as poor solvation of CO2 and H2, a lack of genetic tools and metabolic understanding, a limited number of strains that have been explored, and limited techniques for downstream processing of products. Nonphotosynthetic applications are at an earlier maturity level than photosynthetic ones, and support is warranted on these applications to advance understanding and the maturity of these systems.
Approaches Based on Chemolithotrophs
Whereas cyanobacteria derive their energy and carbon from the reduction of CO2, chemolithotrophs derive their energy from the oxidation of reduced inorganic compounds and their carbon from CO2 (Kelly, 1981). This allows chemolithotrophs to perform light-independent CO2 fixation, eliminating photosynthetic production issues like cell shading. However, cultivating chemolithotrophs is more complex as two inputs are required instead of one. Previous studies have characterized and tested the CO2 fixation capabilities and commercial applications of selected chemolithotrophs (Schiel-Bengelsdorf and Durre, 2012).
Use of Acetogens with a Feedstock of CO2
Acetogens are a well-studied subset of mixotrophic chemolithotrophs that operate strictly under anaerobic conditions (Ragsdale and Pierce, 2008). One unique characteristic of these bacteria is the native Wood-Ljungdahl pathway (WLP), which fixes two CO2 molecules into one acetyl-CoA with less energy than other carbon fixation pathways (Ragsdale, 2008). This gives acetogens an advantage as microbial production platforms because they can bypass carbon loss under certain fermentation conditions.
During traditional heterotrophic fermentation the conversion of sugar into acetyl-CoA results in one-third of all carbon lost to CO2 production, limiting the maximum theoretical yield of products to 67 percent or less. During nonphotosynthetic mixotrophic fermentation the WLP reassimilates the CO2 generated from sugar conversion, resulting in three acetyl-CoA and one adenosine triphosphate (ATP) for every molecule of glucose or hexose sugar. The amount of CO2 reassimilated is dependent on the availability of NAD(P)H and, therefore, is inversely dependent on the target product’s degree of reduction. If the product is highly reduced, less NAD(P)H is available for CO2 fixation and vice versa.
Mixotrophic production has many advantages relevant to industrial viability. Using syngas in addition to sugar fermentation provides added reducing power for CO2 reassimilation. Sugar catabolism, specifically glycolysis, provides ample ATP, which would otherwise be severely limited in WLP gas-only production. Mixotrophic production also potentially enables larger product yields by dividing carbon utilization between biomass and product formation.
Carbon catabolite repression (CCR), in which one feedstock is preferred over the other (Bertsch and Muller, 2015), could present challenges for mixotrophic production. The concern with acetogens is that the sugar feedstock is preferred, thus reducing or eliminating the utilization of syngas for energy production or carbon fixation. However, nonphotosynthetic mixotrophic fermentation with syngas has been demonstrated in a variety of acetogenic microbes without CCR (Jones et al., 2016). In that study, carbon labeling fermentation resulted in Clostridium ljungdahlii with 73-80 percent and C. autoethanogenum with 51-58 percent of its acetate derived from syngas, demonstrating concurrent utilization of both feedstocks with little CO2 lost from glycolysis. The study was also able to engineer Clostridium ljungdahlii to produce acetone, a commodity with a market of $8 billion.2 Acetone anabolism does not require additional NAD(P)H downstream of acetyl-CoA, allowing for greater CO2 fixation. Mixotrophic production from a high-density continuous fermenter resulted in a titer of 10 g/L and a productivity of 2 g/L/h, 92 percent of the theoretical mixotrophic maximum compared to 138 percent of the theoretical heterotrophic maximum. However, metabolic engineering was not applied to the host strain, and reactor conditions were not extensively optimized, especially for scale-up. More research is necessary to overcome the low degree of
___________________
2 See https://www.hexaresearch.com/press-release/global-acetone-market.
reduction required for the target products. The capability to engineer acetogens is currently limited by knowledge gaps and insufficient engineering tools (Bertsch and Muller, 2015).
Use of Two-Stage Integrated Process
Given the limitations of engineering acetogens, a viable alternative is to use other well-studied organisms to produce the target chemicals using carbon provided by acetogen CO2 fixation. In a demonstration of this approach, Hu et al. created a two-stage integrated process that converts syngas to hydrocarbons (Hu et al., 2016). In the first stage CO2 and CO or H2 are converted anaerobically to acetic acid by the acetogen Moorella thermoacetica. Acetic acid is then fed into the second stage where it is converted aerobically into lipids by Yarrowia lipolytica. Accounting for the fact that certain products are not readily formed under anaerobic conditions, this two-stage bioreactor process allows for the nonphotosynthetic fixation of CO2 followed by acetate-dependent production of aerobic products. The fermentation conditions of M. thermoacetica and Y. lipolytica were optimized for acetate and lipid production, respectively. M. thermoacetica growth and production were divided into two phases. A COdependent growth phase was established under CO2/CO conditions; then the reactor was switched to H2/CO2 to increase production of acetate. This allowed for a high cell density and high specific productivity, resulting in 0.9 g of acetate/L/h. Cell recycling was used in Y. lipolytica fermentation to decouple growth and lipogenesis from acetic acid consumption. This too allowed for a high cell density and high lipid productivity. When combined into the two-stage reactor system the results were inferior to the individual reactors, producing 18 g/L of C16-C18 triacylglcerides from syngas with an overall productivity of 0.2 g/L/h and 36 percent lipid content. Despite this shortcoming, the CO2 fixation rate exceeded the CO2 generation rate, demonstrating the feasibility of a two-stage bioprocess to convert syngas into useful commodities.
Use of Acetogens with a Feedstock of CO
Some acetogens are able to efficiently convert CO into useable CO2 via a carbon monoxide dehydrogenase. Of these acetogens, a few produce products other than acetate from acetyl-CoA, including ethanol, butanol, and butyrate (Kopke et al., 2011). Researchers have investigated the fermentation conditions required to boost ethanol production, such as by optimizing metal cofactor concentrations, pH, and trace nutrients, and sought to optimize bioreactor designs for peak gas-to-liquid volumetric transfer efficiencies (Daniell et al., 2012). This research has paved the way for the commercialization of acetogens in the production of various high-value commodities. For example, LanzaTechTM, a company focused on converting fuel gases into useful commodities,3 currently operates a 100,000 gallon per year commer-
___________________
cial plant that utilizes steel flue gas for ethanol production. Their production scheme employs acetogens, such as Clostridium autoethanegenum, which can utilize CO2 and H2 or CO, with or without H2.4 Nonphotosynthetic fixation of carbon dioxide through gas fermentation in the presence of hydrogen is a scalable mechanism of nonphotosynthetic fixation of carbon dioxide. The company also has laboratory-scale pilots of butadiene, propylene, and 2,3-butanediol (being developed as a precursor to jet fuel).
Approaches Based on Bioelectrochemical Systems
Expansion of renewable energy has provided an avenue for the sustainable production of commodities through bioelectrochemical systems. Bioelectrochemical systems carry out artificial photosynthesis by providing microorganisms with electrons that the cells use to reduce CO2 into small organic compounds. In a basic bioelectrochemical setup that draws electricity from a photovoltaic system, electrons are generated at the anode from water, an inexpensive source of electrons, or organic waste and sulfides, which increase the setup complexity but allow for electron recovery at lower potentials (Lovely and Nevin, 2013). Microbes carry out the reduction of CO2 into organic products at the cathode. This typically occurs under an anaerobic environment to prevent oxygen reduction from depleting available electrons and generating toxic by-products, such as H2O2.
Bioelectrochemical systems have the potential to be more productive than biological systems, especially photosynthesis-based systems. Currently, plant solar conversion efficiency of photonic energy caps at 3-4 percent at peak growth, whereas photosynthetic microbes grown in optimized bioreactors can reach 5-7 percent efficiency (Blankenship et al., 2011). By contrast, photovoltaic devices, which can utilize a greater extent of incident solar energy, have 14-18 percent efficiency. Coupling photovoltaic devices with biological CO2 fixation can result in a more efficient production platform overall.
There are several methods of integrating electrocatalysis into microbial production; the differences between these methods lie in how the electrons are transferred into the biological system (Lovely, 2011). Early research focused on providing electron shuttles that would transfer electrons from the electrodes to electron carriers in the cell. The issue with this method is that the electron shuttles, such as neutral red, are expensive, add complexity to product recovery, and are unstable and toxic within the host organism. Current research is focused on two main approaches: directly transferring electrons from electrodes to microorganisms and indirectly transferring electrons via electron donors. It is unclear whether the direct or indirect approach will prove to be more scalable for commercialization; each method has inherent challenges that need to be overcome to become industrially viable. The integration of electro-
___________________
4 See http://www.lanzatech.com/wp-content/uploads/2014/05/LanzaTech_Ex_Summary_8.5x11_05_09_2014.pdf.
catalysis and microbial production is a relatively new concept with research still in its infancy, and engineering bioreactors for electrocatalysis will be a substantial challenge.
Direct Electron Transfer from Electrodes to Microorganisms
The direct transfer of electrons from electrodes into microorganisms requires linking intercellular reductions with extracellular electrons. Electrotrophs typically transfer electrons out of the cell via conduit cytochromes and their accompanying transporters. To achieve bioelectrochemical production these cellular mechanisms need to be reversed to drive electrons into the cell. This reversion would allow electricity to push biosynthetic pathways toward producing high-value chemicals and fuels. Such approaches have been demonstrated in the production of succinate and acetate.
Succinate. Ross et al. (2011) investigated cathodic electron uptake via electrocatalysis experiments with the bacterial host Shewanella oneidensis. Since it is difficult to measure the transfer of electrons in cells, the experiments were performed under anaerobic conditions in which there was only one functioning fumarate reductase available, thus making formate reduction electrode dependent. The electron transfer conduit in S. oneidensis is known as the multiheme c-type cytochromes (Mtr) respiratory pathway. To test the reversibility of the Mtr pathway and understand the role of its individual components, multiple gene deletions were made. The results indicated that reverse electron flux requires the outer membrane and periplasmic components of Mtr. Interestingly, the electrons needed to pass into the menaquinone pool in the cytoplasmic membrane before reentering the periplasm to reduce formate. While the study was able to demonstrate evidence for the reversibility of the Mtr pathway in S. oneidensis, it also underscored the substantial amount of research needed to investigate electron transport pathways before modifications can be made.
Acetate. Because the direct bioelectrochemical model does not utilize electron shuttles, the host is required to be in immediate contact with the electrode in order for outer membrane electron transfer proteins to interact with the electrode surface. This represents a critical challenge of this method as there are physical limitations to the amount of electrode surface area that can be made available. While some researchers (such as Ross et al.) have focused on elucidating the roles of electron transfer pathways, others have focused on improving efficiency through improved reactor designs and operations. For example, LaBelle and May sought to improve microbial electrosynthesis of acetate. Converting H2 and CO2 into acetate is advantageous as creating a single C–C bond is less challenging and acetate has substantial commercial applications as a feedstock (LaBelle and May, 2017). Previous research demonstrated that increased biomass leads to higher acetate production; in their study, LaBelle and May sought to implement that concept in acetogensis electrocatalysis by increasing biomass production
at the cathode. They accomplished this by switching to a galvanic operation, which was able to provide a higher volumetric current density, thus providing more electrons and better access to the microbes. They also moved to a continuous flow reactor, which reduced nutrient limitations and alleviated product inhibition. These changes not only considerably increased the production and efficiency of acetate production but also decreased the operating costs. While production was boosted to 0.78 g/L/h, significantly more research is needed to further advance bioelectrochemical reactor systems, in particular to improve product recovery.
Indirect Electron Transfer via Electrochemically Synthesized Electron Donors
Electrons can be indirectly transferred from electrodes to microorganisms via electrochemically synthesized electron donors such as H2, formate, ammonia, sulfide, or iron (Lovely and Nevin, 2013). The low redox potentials of H2 and formate allow a favorable thermodynamic reduction of CO2 into organic compounds. Ammonia, sulfide, and iron are less thermodynamically favorable because they have higher redox potential. They also require an electron acceptor, such as oxygen, which adds complexity to the electrocatalysis setup and makes these three electron donors less favorable. Of the two more favorable electron donors, formate is a prime target for bioelectrochemical production. H2 has a low solubility, a low mass transfer rate in cells, and significant safety issues due to combustion concerns inside of pressurized reactors. Formate, by contrast, has high solubility and is readily converted into carbon and energy by cells. However, formate adds complexity to product recovery and can also accumulate and degrade at the anode, reducing yield. To be a viable electron donor, formate needs to be consumed at a rate equal to its production. Formate-facilitated electron transfer has been demonstrated in the production of alcohols and pyruvate.
Alcohols. Li et al. designed a bioelectrochemical production platform producing higher alcohols from CO2-derived formate in Ralstonia eutropha, a lithoautotrophic bacterium (Li et al., 2012). They identified three major hurdles to overcome in their production scheme: (1) engineering Ralstonia to produce alcohols, (2) producing formate under culturing conditions, and (3) engineering Ralstonia to withstand electricity. Previously developed pathways for isobutanol and 3-methyl-1-butanol were successfully installed in Ralstonia. Using an In foil cathode and a Pt anode they were able to produce formate, but culture growth was inhibited. They identified O2– and NO as stress triggers and solved the issue by placing a ceramic cup around the anode, shielding the culture by quenching the reactive species. Fed solely on electricity and CO2 the Ralstonia demonstrated healthy growth and a production of 140 mg/L mixed alcohols.
Pyruvate. In another demonstration, Tashiro et al. developed a CO2 and formate fixation pathway in E. coli that converts two formate and one CO2 into pyruvate (Tashiro et al., 2018).
This pathway, named the reductive glycine pathway (RGP), is theoretically the most efficient route for formate assimilation. RGP has many advantages: (1) it is a linear pathway, (2) its reactions are thermodynamically favorable, and (3) there are no oxygen-sensitive pathways involved. The researchers tested various RGP modules against cellular growth in auxotrophs and confirmed their products via carbon labeling. They evaluated the electrochemical cultivation system with indium, tin, lead, carbon, and copper electrodes. Indium had the highest efficiency for formate formation and performed well during bacterial cultivation. Similar to Ralstonia, the formation of reactive oxygen species threatens E. coli growth and productivity, something that is solved by growing under anaerobic conditions. The RGP was successfully constructed in E. coli, but additional research is needed to continue strain modifications and link pyruvate, a central metabolite, to the production of chemicals and fuels.
A RESEARCH AGENDA FOR BIOLOGICAL UTILIZATION OF CARBON DIOXIDE
While some methods for biological conversion of CO2 into fuels and chemicals are well developed, others are in their infancy. All existing and emerging approaches have technical challenges and limitations, as well as barriers to commercial-scale implementation. Social acceptability and regulations also will play a role in the viability of these technologies. Stages of development and key barriers for various biological utilization approaches are summarized in Figure 5-3 and Table 5-3.
Priority Research Areas
Despite the barriers, there are numerous opportunities for advancing biological utilization of CO2. Methods based on photosynthetic green algae are relatively mature; while they may be optimized to improve their cost competitiveness, there are opportunities, assuming the appropriate policy and social environment, to create valuable products. Cyanobacteria are a particularly promising platform for biological conversion because of their photosynthetic efficiency and genetic manipulability, but additional research is needed in order to improve titers and make cyanobacteria-based processes scalable and economically viable. Nonphotosynthetic biological carbon utilization systems avoid the use of high-cost carbohydrates and can be manipulated to create a wide variety of products. Compared to sugar cultivation, these cultures can be continuously cultivated for photosynthetic and nonphotosynthetic carbon dioxide fixation and methane conversion. Bioelectrochemical systems also hold promise because the electricity can help overcome the inherent inefficiency of photosynthesis. Priority research areas are described below.
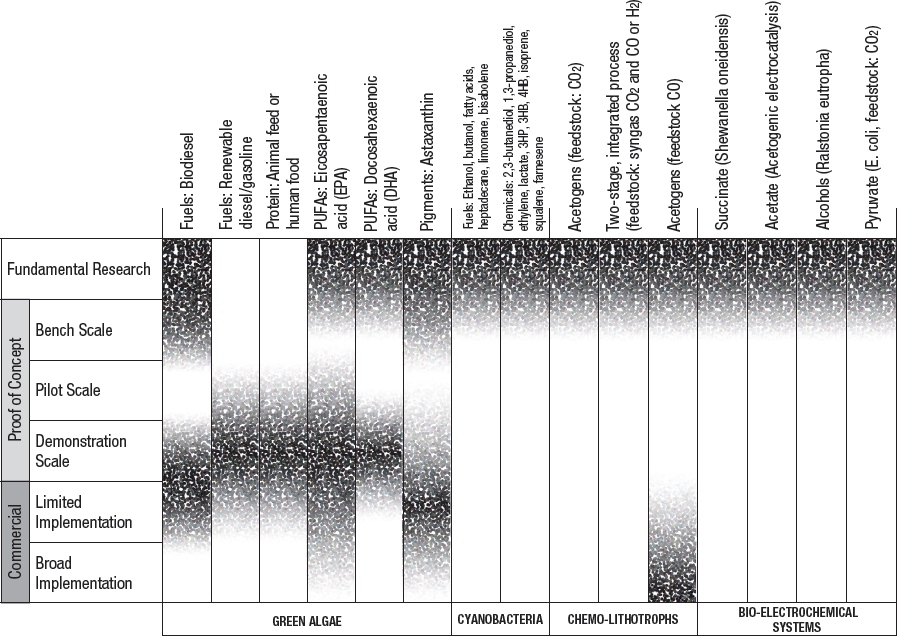
NOTE: Fundamental research is defined as observation and reporting of fundamental principles of a scientific or engineering process, and formulation of a technology concept. The three proof-of-concept stages are defined as progressively larger-scale reactions to produce product. These include bench-scale processes where critical functions are proved and components or systems are validated in a laboratory environment and at a laboratory scale. Pilot plant scale is defined as a system validated in a relevant environment and at an engineering scale. Demonstration plant scale is defined as a full-scale system demonstrated in a relevant environment. Commercial-limited is defined as an actual system operating at a stage where product is being sold in the market in limited areas with specific advantageous geographical, regulatory, or other factors. Commercial-broad is defined as an actual system operating at a stage where product is sold in the market with opportunities not limited to specifically advantaged locations (see Box 9-1).
TABLE 5-3 Key barriers for commercialization of the products from Figure 5-3.
| Platform | Product | Key Barriers |
|---|---|---|
| Green algae | Resource requirements (land, location, water) | |
| Fuels: Biodiesel | Hygroscopicity, cloud point, fouling | |
| Fuels: Renewable diesel/gasoline | Scalability and cost of hydrotreatment, renewable hydrogen | |
| Protein: Animal feed or human food | Scalability, social acceptance (particularly with regard to the use of waste gas) | |
| PUFAs: Eicosapentaenoic acid (EPA) | ||
| PUFAs: Docosahexaenoic acid (DHA) | Scalability, social acceptance | |
| Pigments: Astaxanthin | Limited market size | |
| Cyanobacteria | Growth rate, smaller number of synthetic biology tools | |
| Fuels: Ethanol, butanol, fatty acids, heptadecane, limonene, bisabolene | Low productivity and titers, lack of genetic manipulation tools, scale-up | |
| Chemicals: 2,3-butanediol, 1,3-propanediol, ethylene, lactate, 3-HP, 3-HB, 4-HB, isoprene, squalene, farnesene | Low productivity and titers, lack of genetic manipulation tools, scale-up | |
| Chemolithotrophs | Acetogens (feedstock: CO2) | Low degree of reduction |
| Two-stage, integrated process (feedstock: syngas CO2 and CO or H2) | Hydrogen requirement, low productivity and titers, scale-up | |
| Acetogens (feedstock CO) | Scalability | |
| Bioelectrochemical systems | Succinate (Shewanella oneidensis) | Electrode surface, reactor design and operations improvements |
| Acetate (Acetogenic electrocatalysis) | Product recovery, scalability | |
| Alcohols (Ralstonia eutropha) | Engineering organism to produce alcohols, produce formate under culturing conditions, and withstand electricity; scalability | |
| Pyruvate (E. coli, feedstock: CO2) | Strain modifications, link to production of chemicals and fuels, scalability | |
Bioreactor and Cultivation Optimization
Research is needed to improve bioreactor system design for efficient carbon dioxide solvation, mass transfer, dewatering and harvesting, and management and recycling of water and nutrients. This may include development of better computational and modeling tools for optimizing cultivation processes. Advancement of nonphotosynthetic methods may require novel bioreactor design in order to incorporate new feedstocks or hybrid fermentative systems. This could improve culture monitoring technologies and facilitate scale-up of utilization.
Analytical and Monitoring Tools
Research is needed to improve culture monitoring technologies. This could facilitate scale-up.
Genome-Scale Modeling and Improvement of Metabolic Efficiency
Research is needed to develop and improve methods for in-depth computational modeling, genetic manipulation, biochemical validation, and fermentative demonstration. This could improve metabolic flux, including carbon dioxide uptake and incorporation, photosynthetic efficiency, metabolic streamlining, and product accumulation.
Bioprospecting
Research is needed to accelerate the identification and characterization of organisms or biological systems with unique attributes such as carbon dioxide uptake, various product profiles, photosynthetic efficiency, and environmental tolerance. This could enhance the ability to produce target products in diverse geographic locations.
Valorization of Co-Products
Research is needed to develop feed and food uses for co-products of biological conversion, including studies in product safety and acceptability. This could improve the efficiency of energy and materials use and increase the economic value of biological conversion technologies.
Genetic Tools
Research is needed to enhance engineering of photosynthetic and nonphotosynthetic organisms, including expansion of tools for genetic incorporation, selectable markers, promoter elements, protein folding and stability, and posttranslational control. This could improve efficiency and rates of biomass production and selective product formation.
Pathways to New Products
Research is needed to identify biological pathways to produce nontraditional products and new products for unmet needs in commodity and specialty chemicals. This could expand the portfolio of products made via carbon utilization.
FINDINGS AND RECOMMENDATIONS
Finding 5-1 Biological conversion has a significant advantage over other technologies in that it does not require purified CO2 streams; however, co-localization with CO2 sources would be required due to barriers posed by transportation. Photosynthetic algal biomass production has some significant advantages compared with conventional crops in terms of its land footprint, water use footprint, and protein content. Photosynthesis is inherently inefficient though, requiring compensation with scale, making it land intensive.
Finding 5-2 When developing any type of photosynthesis-based system, it is important to consider the impact of the cultivation method, restrictions in the use of genetically modified organisms, the degree of CO2 solvation, nutrient requirements and downstream burdens, impacts on water and land use, and availability and suitability of CO2 waste streams.
Finding 5-3 Biofuels have potential to advance the circular carbon economy and reduce reliance on and environmental impacts of fossil resource extraction. Renewable fuel production has been limited in its implementation due to its high cost compared with fossil sources and the capital infrastructure necessary to catalytically hydrogenate lipid extracts.
Finding 5-4 Algal protein has the potential to supplement or replace conventional crops as a source of animal and/or human food, but studies have not been validated at scale. Some carbon waste streams, such as those with heavy-metal contaminants, will be inappropriate for animal or human food applications.
Finding 5-5 Polyunsaturated fatty acids are a promising and potentially lucrative product of algal biomass. Pigments may be another valuable algae-based product. These applications may have the additional benefit of reducing pressure on conventional fish-based sources of these products.
Finding 5-6 Cyanobacteria possess many advantages over other biological systems because they can easily be manipulated genetically to create different products, they are more efficient photosynthetically, and they can use CO2 directly. Challenges with cyanobacteria include slow growth rate and limited availability of synthetic biology tools.
Finding 5-7 Nonphotosynthetic microbial systems that can use CO2 rather than sugars hold promise for utilization of gaseous carbon waste feedstocks. Of particular interest may be mixotrophic acetogens.
Recommendation 5-1 Researchers should improve bioreactor system design for efficient carbon dioxide solvation, mass transfer, dewatering and harvesting, and management and recycling of water and nutrients.
Recommendation 5-2 Researchers should improve culture monitoring technologies to facilitate scale-up.
Recommendation 5-3 Researchers should develop and improve methods for in-depth computational modeling, genetic manipulation, biochemical validation, and fermentative demonstration.
Recommendation 5-4 Researchers should enhance the ability to produce target products in diverse geographic locations by accelerating the identification and characterization of organisms or biological systems with unique attributes.
Recommendation 5-5 Researchers should develop feed and food uses for co-products of biological conversion.
Recommendation 5-6 Researchers should enhance engineering of photosynthetic and nonphotosynthetic organisms to improve efficiency and rates of biomass production and selective product formation.
Recommendation 5-7 Researchers should identify biological pathways to produce nontraditional products and new products for unmet needs in commodity and specialty chemicals.
Recommendation 5-8 Research into reducing the capital infrastructure of hydrotreatment of algal lipids or alternative pathways should be conducted by researchers in order to reduce the costs associated with biofuels. Sources of renewable hydrogen are also needed.
Recommendation 5-9 Research into large-scale biomass cultivation should be considered by researchers, given the potential for algal biomass to supplement or replace animal feed and possibly human food. The life-cycle implications of necessary high-purity contaminants present in carbon waste streams will need to be considered.
Recommendation 5-10 Research into nonphotosynthetic technologies should be considered by researchers, given the promising opportunities that are complementary to photosynthetic technologies.
REFERENCES
Abdel-Raouf, N., A. A. Al-Homaidan, and I. B. M. Ibraheem. 2012. Microalgae and wastewater treatment. Saudi Journal of Biological Sciences 19:257-275.
Abishek, M. P., J. Patel, and A. P. Rajan. 2014. Algae oil: A sustainable renewable fuel of future. Biotechnology Research International 2014:272814. doi: 10.1155/2014/272814.
Adarme-Vega, T. C., D. K. Y. Lim, M. Timmins, F. Vernen, Y. Li, and P. M. Schenk. 2012. Microalgal bio-factories: A promising approach towards sustainable omega-3 fatty acid production. Microbial Cell Factories 11:96.
Ahmed, F., Y. Li, and P. M. Schenk. 2012. Algal biorefinery: Sustainable production of biofuels and aquaculture feed? In: Gordon, R. and Seckbach, J., Eds., The Science of Algal Fuels. Springer: Netherlands, 21-41.
Aikawa, S., A. Nishida, S. H. Ho, J. S. Chang, T. Hasunuma, and A. Kondo. 2014. Glycogen production for biofuels by the euryhaline cyanobacteria Synechococcus sp. strain PCC 7002 from an oceanic environment. Biotechnology for Biofuels 7:88.
Al-Sabawi M., and J. Chen. 2012. Hydroprocessing of biomass-derived oils and their blends with petroleum feedstocks: a review. Energy & Fuels. 26:5373-5399.
Albers, S. C., V. A. Gallegos, and C. A. Peebles. 2015. Engineering of genetic control tools in Synechocystis sp. PCC 6803 using rational design techniques. Journal of Biotechnology 216:36-46.
Alper, H., Y.-S. Jin, J. F. Moxley, and G. Stephanopoulos. 2005. Identifying gene targets for the metabolic engineering of lycopene biosynthesis in Escherichia coli. Metabolic Engineering 7:155-164.
Angermayr, S. A., A. D. van der Woude, D. Correddu, A. Vreugdenhil, V. Verrone, and K. J. Hellingwerf. 2014. Exploring metabolic engineering design principles for the photosynthetic production of lactic acid by Synechocystis sp. PCC6803. Biotechnology for Biofuels 7:99.
Asikainen, M., T. Munter, and J. Linnekoski. 2015. Conversion of polar and non-polar algae oil lipids to fatty acid methyl esters with solid acid catalysts–A model compound study. Bioresource Technology 191:300-305.
Atsumi, S., W. Higashide, and J. C. Liao. 2009. Direct photosynthetic recycling of carbon dioxide to isobutyraldehyde. Nature Biotechnology 27:1177-1180.
Batterton, J. C., and C. Van Baalen. 1971. Growth responses of blue-green algae to sodium chloride concentration. Archiv für Mikrobiologie 76:151-165.
Becker, E.W. 2007. Micro-algae as a source of protein. Biotechnology Advances 25:207-210.
Becker, E.W. 2004. Microalgae in human and animal nutrition. In: Richmond, A., Ed., Handbook of Microalgal Culture. Biotechnology and Applied Phycology. Oxford: Blackwell Science, 312-351.
Benemann, J. R., P. Pursoff, and W. J. Oswald. 1978. Engineering Design and Cost Analysis of a Large-Scale Microalgae Biomass System, Final Report to the U.S. Energy Department, NTIS #H CP/T1605-01 UC-61, 91 pp.
Benemann, J. R., J. C. Van Olst, M. J. Massingill, J. A. Carlberg, J. C. Weissman, and D. E. Brune. 2003. The controlled eutrophicatino process: using microalgae for CO2 utilization and agricultural fertilizer recycling. Proceedings of the 6th International Conference on Greenhouse Gas Control Technologies 1 – 4 October 2002, Kyoto, Japan 2:1433-1438.
Berla, B. M., R. Saha, C. M. Immethun, C. D. Maranas, T. S. Moon, and H. B. Pakrasi. 2013. Synthetic biology of cyanobacteria: Unique challenges and opportunities. Frontiers in Microbiology 4:246.
Bertsch, J., and V. Muller. 2015. Bioenergetic constraints for conversion of syngas to biofuels in acetogenic bacteria. Biotechnology for Biofuels 8:210 doi: 10.1186/s13068-015-0393-x. eCollection 2015.
Bhujade, R., M. Chidambaram, A. Kumar, and A. Sapre. 2017. Algae to economically viable low-carbon-footprint oil. Annual Review of Chemical and Biomolecular Engineering 8: 335-357.
Bimbo, A. P. 2007. Current and future sources of raw materials for the long-chain omega-3 fatty acid market. Lipid Technology 19:176-179.
Blankenship, R. E., D. M. Tiede, J. Barber, G. W. Brudvig, G. Fleming, M. Ghirardi, M. R. Gunner, W. Junge, D. M. Kramer, A. Melis, T. A. Moore, C. C. Moser, D. G. Nocera, A. J. Nozik, D. R. Ort, W. W. Parson, R. C. Prince, and R. T. Sayre. 2011. Comparing photosynthetic and photovoltaic efficiencies and recognizing the potential for improvement. Science 332:805-809.
Bleakley, S., and M. Hayes. 2017. Algal proteins: Extraction, application, and challenges concerning production. Foods 6(5):33. doi: 10.3390/foods6050033.
Borowitzka, M. A. 2013. High-value products from microalgae—their development and commercialisation. Journal of Applied Phycology 25(3):743-756.
Broddrick, J. T., B. E. Rubin, D. G. Welkie, N. Du, N. Mih, S. Diamond, J. J. Lee, S. S. Golden, and B. O. Palsson. 2016. Unique attributes of cyanobacterial metabolism revealed by improved genome-scale metabolic modeling and essential gene analysis. Proceedings of the National Academy of Sciences of the United States of America 113:E8344-E8353.
Brune, D. E., H. W. Yen, G. Schwartz, J. R. Benemann, M. J. Massingill, J. C. Van Olst, and J. A. Carlberg. 2014. The controlled eutrophication process; microalgae for biofuels production and fertilizer recycling at the Salton Sea, California. National Energy Technology Laboratory. Available at https://www.netl.doe.gov/publications/proceedings/03/carbon-seq/PDFs/034.pdf.
Buhutskyi, P., S. Chow, B. Ketter, S. C. Fung, D. Yacar, Y. Tang, M. Zivojnovich, M. J. Betenbaugh, and E. J. Bouwer. 2016. Bioresource Technology 222:294-308.
Cerutti, H., A. M. Johnson, N. W. Gillham, and J. E. Boynton. 1997. Epigenetic silencing of a foreign gene in nuclear transformants of Chlamydomonas. Plant Cell 9:925-945.
Chekanov, K., E. Schastnaya, A. Solovchenko, and E. Lobakova. 2017. Effects of CO2 enrichment on primary photochemistry, growth and astaxanthin accumulation in the chlorophyte Haematococcus pluvialis. Journal of Photochemistry and Photobiology B: Biology 171:58-66.
Chekroun, K. B., E. Sanchez, and M. Baghour. 2014. The role of algae in bioremedication of organic pollutants. International Research Jounral of Public and Environmental Health 1:19-32.
Chen, H. W., T. S. Yang, M. J. Chen, Y. C. Chang, C. Y. Lin, I. Eugene, C. Wang, C. L. Ho, K. M. Huang, C. C. Yu, and F. L. Yang. 2012. Application of power plant flue gas in a photobioreactor to grow Spirulina algae, and a bioactivity analysis of the algal water-soluble polysaccharides. Bioresource Technology 120:256-263.
Choi, S. Y., J. Y. Wang, H. S. Kwak, S. M. Lee, Y. Um, Y. Kim, S. J. Sim, J. I. Choi, and H. M. Woo. 2017. Improvement of squalene production from CO2 in Synechococcus elongatus PCC 7942 by metabolic engineering and scalable production in a photobioreactor. ACS Synthetic Biology 6:1289-1295.
Daniell, J., M. Köpke, and S. D. Simpson, 2012. Commerical biomass syngas fermentation. Energies 5:5372-5417.
Davies, F. K., V. H. Work, A. S. Beliaev, and M. C. Posewitz. 2014. Engineering limonene and bisabolene production in wild type and a glycogen-deficient mutant of Synechococcus sp. PCC 7002. Frontiers in Bioengineering and Biotechnology 2:21.
Davis, R., and M. Biddy. 2013. Algal lipid extraction and upgrading to hydrocarbons technology pathway. National Renewable Energy Laboratory Technical Report NREL/TP-5100-58049, PNNL-22315.
de Morais, M. G., and J. A. Costa. 2007. Isolation and selection of microalgae from coal fired thermoelectric power plant for biofixation of carbon dioxide. Energy Conversion and Management 48:2169-2173.
Demirbas, A. 2010. Use of algae as biofuel sources. Energy Conversion and Management 51:2738-2749.
de Wilde, C., H. Van Houdt, S. De Buck, G. Angenon, G. De Jaeger, and A. Depicker. 2000. Plants as bioreactors for protein production: Avoiding the problem of transgene silencing. Plant Molecular Biology 43:347-359.
Dexter, J., and P. Fu. 2009. Metabolic engineering of cyanobacteria for ethanol production. Energy & Environmental Science 2:857-864.
DOE BETO (Department of Energy Bioenergy Technologies Office). 2016. National Algal Biofuels Technology Review.
DOE BETO. 2017. Biofuels and Bioproducts from Wet and Gaseous Waste Streams: Challenges and Opportunitites.
DOE EERE (Department of Energy Office of Energy Efficiency & Renewable Energy). 2010. National Algal Biofuels Technology Roadmap.
DOE EERE. 2017. Algae Cultivation for Carbon Capture and Utilization Workshop Summary Report.
Farooq, W., W. I. Suh, M. S. Park, and J. W. Yang. 2015. Water use and its recycling in microalgae cultivation for biofuel application. Bioresource Technology 184:73-81.
Ferenczi, A., D. E. Pyott, A. Xipnitou, and A. Molnar. 2017. Efficient targeted DNA editing and replacement in Chlamydomonas reinhardtii using Cpf1 ribonucleoproteins and single-stranded DNA. Proceedings of the National Academy of Sciences of the United States of America 114:13567-13572, doi:10.1073/pnas.1710597114.
Fisher, A. W., Jr., and J. S. Burlew. 1953. Nutritional value of microscopic algae. Pp. 303-310 in Algal Culture from Laboratory to Pilot Plant. Washington, DC: Carnegie Institution.
Fong, S. S., A. P. Burgard, C. D. Herring, E. M. Knight, F. R. Blattner, C. D. Maranas, and B. O. Palsson. 2005. In silico design and adaptive evolution of Escherichia coli for production of lactic acid. Biotechnology for Bioengineering 91:643-648.
Gao, X., F. Gao, D. Liu, H. Zhang, X. Nie, and C. Yang. 2016. Engineering the methylerythritol phosphate pathway in cyanobacteria for photosynthetic isoprene production from CO2. Energy & Environmental Science 9:1400-1411.
Gao, Z., H. Zhao, Z. Li, X. Tan, and X. Lu. 2012. Photosynthetic production of ethanol from carbon dioxide in genetically engineered cyanobacteria. Energy & Environmental Science 5:9857-9865.
Gimpel, J. A., E. A. Specht, D. R. Georgianna, and S. P. Mayfield. 2013. Advances in microalgae engineering and synthetic biology applications for biofuel production. Current Opinion in Chemical Biology 17:489-495.
Golden, S. S., J. Brusslan, and R. Haselkorn. 1987. Genetic engineering of the cyanobacterial chromosome. Methods in Enzymology 153:215-231.
Golueke, C. G., and W. J. Oswald. 1959. Biological conversion of light energy to the chemical energy of methane. Applied Microbiology 7(4):219-227.
Guieysse, B., Q. Béchet, and A. Shilton. 2013. Variability and uncertainty in water demand and water footprint assessments of fresh algae cultivation based on case studies from five climatic regions. Bioresource Technology 128:317-323.
Gupta, P. L., S. M. Lee, and H. J. Choi. 2015. A mini review: Photobioreactors for large scale algal cultivation. World Journal of Microbiology and Biotechnology 31:1409-1417.
Hazrat, M. A., M. G. Rasul, and M. M. Khan. 2015. Lubricity improvement of the ultra-low sulfur diesel fuel with the biodiesel. Energy Procedia 75:111-117.
Hendry, J. I., C. B. Prasannan, A. Joshi, S. Dasgupta, and P. P. Wangikar. 2016. Metabolic model of Synechococcus sp. PCC 7002: Prediction of flux distribution and network modification for enhanced biofuel production. Bioresource Technology 213:190-197.
Hernandez-Mireles, I., R. van der Stel, and E. Goetheer. 2014. New methodologies for integrating algae with CO2 capture. Energy Procedia 63:7954-7958.
Hirokawa, Y., Y. Maki, T. Tatsuke, and T. Hanai. 2016. Cyanobacterial production of 1,3-propanediol directly from carbon dioxide using a synthetic metabolic pathway. Metabolic Engineering 34:97-103.
Hu, P., S. Chakraborty, A. Kumar, B. Woolston, H. Liu, D. Emerson, and G. Stephanopoulos. 2016. Integrated bioprocess for conversion of gaseous substrates to liquids. Proceedings of the National Academy of Sciences of the United States of America 113:3773-3778.
Hucka, M., A. Finney, H. M. Sauro, H. Bolouri, J. C. Doyle, H. Kitano, A. P. Arkin, B. J. Bornstein, D. Bray, A. Cornish-Bowden, A. A. Cuellar, S. Dronov, E. D. Gilles, M. Ginkel, V. Gor, I. I. Goryanin, W. J. Hedley, T. C. Hodgman, J. H. Hofmeyr, P. J. Hunter, N. S. Juty, J. L. Kasberger, A. Kremling, U. Kummer, N. Le Novere, L. M. Loew, D. Lucio, P. Mendes, E. Minch, E. D. Mjolsness, Y. Nakayama, M. R. Nelson, P. F. Nielsen, T. Sakurada, J. C. Schaff, B. E. Shapiro, T. S. Shimizu, H. D. Spence, J. Stelling, K. Takahashi, M. Tomita, J. Wagner, J. Wang, and SBML Forum. 2003. The systems biology markup language (SBML): A medium for representation and exchange of biochemical network models. Bioinformatics 19:524-531.
Huertas, I. E., B. Colman, G. S. Espie, and L. M. Lubian. 2000. Active transport of CO2 by three species of marine microalgae. Journal of Phycology 36:314-320.
Immethun, C. M., D. M. DeLorenzo, C. M. Focht, D. Gupta, C. B. Johnson, and T. S. Moon. 2017. Physical, chemical, and metabolic state sensors expand the synthetic biology toolbox for Synechocystis sp. PCC 6803. Biotechnology and Bioengineering 114:1561-1569.
Jones, S. W., A. G. Fast, E. D. Carlson, C. A. Wiedel, J. Au, M. R. Antoniewicz, E. T. Papoutsaki, and B. P. Tracy. 2016. CO2 fixation by anaerobic non-photosynthetic mixotrophy for improved carbon conversion. Nature Communications 7:12800. doi: 10.1038/ncomms12800.
Kadam, K. L. 1997. Power plant flue gas as a source of CO2 for microalgae cultivation: economic impact of different process options. Energy Conversion and Management. 38:S505-S510.
Kanno, M., A. L. Carroll, and S. Atsumi. 2017. Global metabolic rewiring for improved CO2 fixation and chemical production in cyanobacteria. Nature Communications 8:14724.
Kaur, G., D. P. Singh, J. I. S. Khattar, and J. Nadda. 2009. Microalgae: A source of natural colours. In: Khattar, J. I. S., Singh, D. P., and Kaur, G., Eds., Algal Biology and Biotechnology. I. K. International Publishing House Pvt. Ltd. New Delhi, 129-150.
Kelly, D. P. 1981. Introduction to the chemolithotropihc bacteria. In: Starr, M. P., Stolp, H., Truper, H. G., Balows, A., and Schlegel, H. G., Eds., The Prokaryotes. Springer: Berlin, Heidelberg.
Kim, W. J., H. U. Kim, and S. Y. Lee. 2017. Current state and applications of microbial genome-scale metabolic models. Current Opinion in Systems Biology 2:10-18.
Könst, P., I. H. Mireles, R. van der Stel, P. van Os, and E. Goetheer. 2017. Integrated system for capturing CO2 as feedstock for algae production. Energy Procedia 114:7126-7132.
Köpke, M., C. Mihalcea, J. C. Bromley, and S. D. Simpson. 2011. Fermentative production of ethnol from carbon monoxide. Current Opinion of Biotechnology 22:320-325.
Knoop, H., Y. Zilliges, W. Lockau, and R. Steuer. 2010. The metabolic network of Synechocystis sp. PCC 6803: Systemic properties of autotrophic growth. Plant Physiology 154:410-422.
LaBelle, E. V., and H. D. May. 2017. Energy efficiency and productivity enhancement of microbial electrosynthesis of acetate. Frontiers in Microbiology 8:756 doi: 10.3389/fmicb.2017.00756. eCollection 2017.
Lan, E. I., S. Y. Ro, and J. C. Liao. 2013. Oxygen-tolerant coenzyme A-acylating aldehyde dehydrogenase facilitates efficient photosynthetic n-butanol biosynthesis in cyanobacteria. Energy & Environmental Science 6:2672-2681.
Lee, H. J., J. Choi, S. M. Lee, Y. Um, S. J. Sim, Y. Kim, and H. M. Woo. 2017a. Photosynthetic CO2 conversion to fatty acid ethyl esters (FAEEs) using engineered cyanobacteria. Journal of Agricultural and Food Chemistry 65:1087-1092.
Lee, H. J., J. Lee, S. M. Lee, Y. Um, Y. Kim, S. J. Sim, J. I. Choi, and H. M. Woo. 2017b. Direct conversion of CO2 to alpha-farnesene using metabolically engineered Synechococcus elongatus PCC 7942. Journal of Agricultural and Food Chemistry 65:10424-10428.
Lee, S. J., D. Y. Lee, T. Y. Kim, B. H. Kim, J. Lee, and S. Y. Lee. 2005. Metabolic engineering of Escherichia coli for enhanced production of succinic acid, based on genome comparison and in silico gene knockout simulation. Applied and Environmental Microbiology 71:7880-7887.
Li, C., F. Tao, J. Ni, Y. Wang, F. Yao, and P. Xu. 2015. Enhancing the light-driven production of D-lactate by engineering cyanobacterium using a combinational strategy. Scientific Reports 5:9777.
Li, H., P. H. Opgenorth, D.G. Wernick, S. Rogers, T.-Y. Wu, W. Higashide, P. Malati, Y.-X. Huo, K. M. Cho, and J. C. Liao. 2012. Integrated electromicrobial conversion of CO2 to higher alcohols. Science 335:1596.
Lin, P. C., R. Saha, F. Zhang, and H. B. Pakrasi. 2017. Metabolic engineering of the pentose phosphate pathway for enhanced limonene production in the cyanobacterium Synechocysti s sp. PCC 6803. Scientific Reports 7:17503.
Lovley, D. R. 2011. Powering microbes with electricity: Direct electron transfer from electrodes to microbes. Environmental Microbiology Reports 3:27-35.
Lovley, D. R., and K. P. Nevin. 2013. Electrobiocommodities: Powering microbial production of fuels and commodity chemicals from carbon dioixde with electricity. Current Opinion in Biotechnology 24:385-390.
Maeda, Y., T. Yoshino, T. Matsunaga, M. Matsumoto, and T. Tanaka. 2018. Marine microalgae for production of biofuels and chemicals. Current Opinion in Biotechnology 50:111-120.
Markley, A. L., M. B. Begemann, R. E. Clarke, G. C. Gordon, and B. F. Pfleger. 2015. Synthetic biology toolbox for controlling gene expression in the cyanobacterium Synechococcus sp. strain PCC 7002. ACS Synthetic Biology 4:595-603.
Martínez-Jerónimo, F. and F. Espinosa-Chávez. 1994. A laboratory-scale system for mass culture of freshwater microalgae in polyethylene bags. Journal of Applied Phycology 6:423-425.
Mayfield, S. P., A. L. Manuell, S. Chen, J. Wu, M. Tran. D. Siefker, M. Muto, J. Marin-Navarro. 2007. Chlamydomonas reinhardtii chloroplasts as protein factories. Current Opinion in Biotechnology 18: 126-133.
McEwen, J.T., I. M. Machado, M. R. Connor, and S. Atsumi. 2013. Engineering Synechococcus elongatus PCC 7942 for continuous growth under diurnal conditions. Applied and Environmental Microbiology 79:1668-1675.
McEwen, J. T., M. Kanno, and S. Atsumi. 2016. 2,3-Butanediol production in an obligate photoautotrophic cyanobacterium in dark conditions via diverse sugar consumption. Metabolic Engineering 36:28-36.
Melis, A. 2009. Solar energy conversion efficiencies in photosynthesis: Minimizing the chlorophyll antennae to maximize efficiency. Plant Science 177:272-280.
Miara, A., P. T. Pienkos, M. Bazilian, R. Davis, and J. Macknick. 2014. Planning for algal systems: An energy-water-food nexus perspective. Industrial Biotechnology 10:202-211.
Mo, H., X. Xie, T. Zhu, and X. Lu. 2017. Effects of global transcription factor NtcA on photosynthetic production of ethylene in recombinant Synechocystis sp. PCC 6803. Biotechnology for Biofuels 10:145.
Morgan-Kiss, R. M., J. C. Priscu, T. Pocock, L. Gudynaite-Savitch, and N. P. Huner. 2006. Adaptation and acclimation of photosynthetic microorganisms to permanently cold environments. Microbiology and Molecular Biology Reviews 70:222-252.
Mubarak, M., A. Shaija, and T. V. Suchithra. 2015. A review on the extraction of lipid from microalgae for biodiesel production. Algal Research 7:117-123.
Nitsan, Z., S. Mokady, and A. Sukenik. 1999. Enrichment of poultry products with ω3 fatty acids by dietary supplementation with the alga Nannochloropsis and mantur oil. Journal of Agricultural and Food Chemistry 47:5127-5132.
No, S.Y. 2014. Application of hydrotreated vegetable oil from triglyceride based biomass to CI engines–A review. Fuel 115:88-96.
Nozzi, N. E., and S. Atsumi. 2015. Genome engineering of the 2,3-butanediol biosynthetic pathway for tight regulation in cyanobacteria. ACS Synthetic Biology 4:1197-1204.
Nozzi, N. E., A. E. Case, A. L. Carroll, and S. Atsumi. 2017. Systematic approaches to efficiently produce 2,3-butanediol in a marine cyanobacterium. ACS Synthetic Biology 6:2136-2144.
NRC (National Research Council). 2012. Sustainable Development of Algal Biofuels in the United States. Washington, DC: The National Academies Press. doi: 10.17226/13437.
Oliver, J. W., and S. Atsumi. 2015. A carbon sink pathway increases carbon productivity in cyanobacteria. Metabolic Engineering 29:106-112.
Oliver, J. W., I. M. Machado, H. Yoneda, and S. Atsumi. 2013. Cyanobacterial conversion of carbon dioxide to 2,3-butanediol. Proceedings of the National Academy of Sciences of the United States of America 110:1249-1254.
Ponnampalam, E. N., N. J., Mann, and A. J. Sinclair. 2006. Effect of feeding systems on omega-3 fatty acids, conjugated linoleic acid and trans fatty acids in Australian beef cuts: Potential impact on human health. Asia Pacific Journal of Clinical Nutrition 15:21.
Raab, A. M., G. Gebhardt, N. Bolotina, D. Weuster-Botz, and C. Lang. 2010. Metabolic engineering of Saccharomyces cerevisiae for the biotechnological production of succinic acid. Metabolic Engineering 12:518-525.
Radmann, E. M., F. V. Camerini, T. D. Santos, and J. A. Costa. 2011. Isolation and application of SOX and NOX resistant microalgae in biofixation of CO2 from thermoelectricity plants. Energy Conversion and Management 52:3132-3136.
Ragsdale, S. W. 2008. Enzymology of the Wood-Ljungdahl pathway of acetogenesis. Annals of the New York Academy of Sciences 1125:129-136.
Ragsdale, S. W., and E. Pierce. 2008. Acetogenesis and the Wood-Ljungdahl pathway of CO2 fixation. Biochimica et Biophysica Acta 1784:1873-1898.
Rasala, B. A., D. J. Barrera, J. Ng, T. M. Plucinak, J. N. Rosenberg, D. P. Weeks, G. A. Oyler, T. C. Peterson, F. Haerizadeh, and S. P. Mayfield. 2013. Expanding the spectral palette of fluorescent proteins for the green microalga Chlamydomonas reinhardtii. Plant Journal 74:545-556.
Rasala, B. A., P. A. Lee, Z. Shen, S. P. Briggs, and S. P. Mayfield. 2012. Robust expression and secretion of Xylanase1 in Chlamydomonas reinhardtii by fusion to a selection gene and processing with the FMDV 2A peptide. PLoS One 7(8):e43349. doi:10.137/journal.pone.0043349.
Richardson, J. W., M. D. Johnson, and J. L. Outlaw. 2012. Economic comparison of open pond raceways to photo bio-reactors for profitable production of algae for transportation fuels in the Southwest. Algal Research 1:93-100.
Rogers, J. N., J. N. Rosenberg, B. J. Guzman, V. H. Oh, L. E. Mimbela, A. Ghassemi, M. J. Betenbaugh, G. A. Oyler, and M. D. Donohue. 2014. A critical analysis of paddlewheel-driven raceway ponds for algal biofuel production at commercial scales. Algal Research 76-88.
Rösch, C., J. Skarka, and N. Wegerer. 2012. Materials flow modeling of nutrient recycling in biodiesel production from microalgae. Bioresource Technology 107:191-199.
Ross, D. E., J. M. Flynn, D. B. Baron, J. A. Gralnick, and D. R. Bond. 2011. Towards electrosynthesis in Shewanella: Energetics of reversing the Mtr pathway for reductive metabolism. PLoS One 6(2): e16649. https://doi.org/10.1371/journal.pone.0016649.
Roy, J. J., L. Sun, and L. Ji. 2014. Microalgal proteins: A new source of raw material for production of plywood adhesive. Journal of Applied Phycology 26:1415-1422.
Ruffing, A. M. 2014. Improved free fatty acid production in cyanobacteria with Synechococcus sp. PCC 7002 as host. Frontiers in Bioengineering and Biotechnology 2:17.
Saha, R., D. Liu, A. Hoynes-O’Connor, M. Liberton, J. Yu, M. Bhattacharyya-Pakrasi, A. Balassy, F. Zhang, T. S. Moon, C. D. Maranas, and H. B. Pakrasi. 2016. Diurnal regulation of cellular processes in the cyanobacterium Synechocystis sp. strain PCC 6803: Insights from transcriptomic, fluxomic, and physiological analyses. mBio 7(3):e00464-16.
Sander, K., and G. S. Murthy. 2010. Life cycle analysis of algae biodiesel. International Journal of Life Cycle Assessment 15(7):704-714.
Savakis, P. E., S. A. Angermayr, and K. J. Hellingwerf. 2013. Synthesis of 2,3-butanediol by Synechocystis sp. PCC6803 via heterologous expression of a catabolic pathway from lactic acid- and enterobacteria. Metabolic Engineering 20:121-130.
Schiel-Bengelsdorf, B., and P. Durre. 2012. Pathway engineering and synthetic biology using acetogens. FEBS Letters 586:2191-2198.
Schoepp, N. G., R. L. Stewart, V. Sun, A. J. Quigley, D. Mendola, S. P. Mayfield, and M. D. Burkart. 2014. System and method for research-scale outdoor production of microalgae and cyanobacteria. Bioresource Technology 166:273-281.
Sheehan, J., T. Dunahay, J. Benemann, and P. Roessler. 1998. A Look Back at the U.S. Department of Energy’s Aquatic Species Program: Biodiesel from Algae. NREL/TP-580-24190. Golden, CO: National Renewable Energy Laboratory. Available at https://www.nrel.gov/docs/legosti/fy98/24190.pdf.
Shin, S.-E., J.-M. Lim, H. G. Koh, E. K. Kim, N. K. Kang, S. Jeon, S. Kwon, W.-S. Shin, B. Lee, K. Hwangbo, J. Kim, S. H. Ye, J.-Y. Yun, H. Seo, H.-M. Oh, K.-J. Kim, J.-S. Kim, W.-J. Jeong, Y. K. Chang, and B.-r. Jeong. 2016. CRISPR/Cas9-induced knockout and knock-in mutations in Chlamydomonas reinhardtii. Nature 6:27810 doi:10.1038/srep27810.
Shirai, T., T. Osanai, and A. Kondo. 2016. Designing intracellular metabolism for production of target compounds by introducing a heterologous metabolic reaction based on a Synechosystis sp. 6803 genome-scale model. Microbial Cell Factories 15:13.
Singh, J., and S. Gu. 2010. Commercialization potential of microalgae for biofuels production. Renewable and Sustainable Energy Reviews 14(9):2596-2610.
Stanford, N. J., K. Wolstencroft, M. Golebiewski, R. Kania, N. Juty, C. Tomlinson, S. Owen, S. Butcher, H. Hermjakob, N. Le Novere, W. Mueller, J. Snoep, and C. Goble. 2015. The evolution of standards and data management practices in systems biology. Molecular Systems Biology 11:851.
Szyjka, S. J., S. Mandal, N. G. Schoepp, B. M. Tyler, C. B. Yohn, Y. S. Poon, S. Villareal, M. D. Burkart, J. B. Shurin, and S. P. Mayfield. 2017. Evaluation of phenotype stability and ecological risk of a genetically engineered alga in open pond production. Algal Research 24:378-386.
Tashiro, Y., S. Hirano, M. M. Matson, S. Atsumi, and A. Kondo. 2018. Electrical-biologcal hybrid system for CO2 reduction. Metabolic Engineering 47:211-218.
Tran, A. V., and R. P. Chambers. 1987. The dehydration of fermentative 2,3-butanediol into methyl ethyl ketone. Biotechnology and Bioengineering 29:343-351.
Trentacoste, E. M., A. M. Martinez, and T. Zenk. 2015. The place of algae in agriculture: Policies for algal biomass production. Photosynthesis Research 123(3):305-315.
Triana, J., A. Montagud, M. Siurana, D. Fuente, A. Urchueguia, D. Gamermann, J. Torres, J. Tena, P. F. de Cordoba, and J. F. Urchueguia. 2014. Generation and evaluation of a genome-scale metabolic network model of Synechococcus elongatus PCC7942. Metabolites 4:680-698.
Ueno, K., Y. Sakai, C. Shono, I. Sakamoto, K. Tsukakoshi, Y. Hihara, K. Sode, and K. Ikebukuro. 2017. Applying a riboregulator as a new chromosomal gene regulation tool for higher glycogen production in Synechocystis sp. PCC 6803. Applied Microbiology and Biotechnology 101:8465-8474.
van der Woude, A. D., S. A. Angermayr, V. Puthan Veetil, A. Osnato, and K. J. Hellingwerf. 2014. Carbon sink removal: Increased photosynthetic production of lactic acid by Synechocystis sp. PCC6803 in a glycogen storage mutant. Journal of Biotechnology 184:100-102.
van Haveren, J., E. L. Scott, and J. Sanders. 2008. Bulk chemicals from biomass. Biofuels, Bioproducts and Bio-refining 2:41-57.
Vieira, E. D., G. Andrietta Mda, and S. R. Andrietta. 2013. Yeast biomass production: A new approach in glucose-limited feeding strategy. Brazilian Journal of Microbiology 44(2):551-558.
Vocke, R. W., K. L. Sears, J. J. O’Toole, and R. B. Wildman. 1980. Growth responses of selected freshwater algae to trace elements and scrubber ash slurry generated by coal-fired power plants. Water Research 14:141-150.
Vu, T. T., E. A. Hill, L. A. Kucek, A. E. Konopka, A. S. Beliaev, and J. L. Reed. 2013. Computational evaluation of Synechococcus sp. PCC 7002 metabolism for chemical production. Biotechnology Journal 8:619-630.
Wang, B., Y. Li, N. Wu, and C.Q. Lan. 2008. CO2 bio-mitigation using microalgae. Applied Microbiology and Biotechnology 79:707-718.
Wang, X., W. Liu, C. Xin, Y. Zheng, Y. Cheng, S. Sun, R. Li, X.-G. Zhu, S. Y. Dai, P. M. Rentzepis, and J. S. Yuan. 2016a. Enhanced limonene production in cyanobacteria reveals photosynthesis limitations. Proceedings of the National Academy of Sciences of the United States of America 113:14225-14230.
Wang, Y., T. Sun, X. Gao, M. Shi, L. Wu, L. Chen, and W. Zhang. 2016b. Biosynthesis of platform chemical 3-hydroxypropionic acid (3-HP) directly from CO2 in cyanobacterium Synechocystis sp. PCC 6803. Metabolic Engineering 34:60-70.
Wen, Z., and M. B. Johnson. 2009. Microalgae as a Feedstock for Biofuel Production. Virginia Cooperative Extension, Publication 442-886. Available at http://pubs.ext.vt.edu/content/dam/pubs_ext_vt_edu/422/442-886/442-886_pdf.pdf.
Whitney, S. M., R. L. Houtz, and H. Alonso. 2011. Advancing our understanding and capacity to engineer nature’s CO2-sequestering enzyme, Rubisco. Plant Physiology 155:27-35.
Wilcox, J. 2012. Carbon Capture. New York: Springer-Verlag.
Woertz, I., A. Feffer, T. Lundquist, and Y. Nelson. 2009. Algae grown on dairy and municipal wastewater for simultaneous nutrient removal and lipid production for biofuel feedstock. Journal of Environmental Engineering 135:1115-1122.
Wu-Scharf, D., B. Jeong, C. Zhang, and H. Cerutti. 2000. Transgene and transposon silencing in Chlamydomonas reinhardtii by a DEAH-box RNA helicase. Science 290:1159-1162.
Yang, J., M. Xu, X. Zhang, Q. Hu, M. Sommerfeld, and Y. Chen. 2011. Life-cycle analysis on biodiesel production from microalgae: Water footprint and nutrients balance. Bioresource Technology 102:159-165.
Yim, H., R. Haselbeck, W. Niu, C. Pujol-Baxley, A. Burgard, J. Boldt, J. Khandurina, J. D. Trawick, R. E. Osterhout, R. Stephen, J. Estadilla, S. Teisan, H. B. Schreyer, S. Andrae, T. H. Yang, S. Y. Lee, M. J. Burk, and S. Van Dien. 2011. Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol. Nature Chemical Biology 7:445-452.
Yoshikawa, K., Y. Toya, and H. Shimizu. 2017. Metabolic engineering of Synechocystis sp. PCC 6803 for enhanced ethanol production based on flux balance analysis. Bioprocess and Biosystems Engineering 40:791-796.
Yoshino, T., Y. Liang, D. Arai, Y. Maeda, T. Honda, M. Muto, N. Kakunaka, and T. Tanaka. 2015. Alkane production by the marine cyanobacterium Synechococcus sp. NKBG15041c possessing the alpha-olefin biosynthesis pathway. Applied Microbiology and Biotechnology 99:1521-1529.
Yu, J., M. Liberton, P. F. Cliften, R. D. Head, J. M. Jacobs, R. D. Smith, D. W. Koppenaal, J. J. Brand, and H. B. Pakrasi. 2015. Synechococcus elongatus UTEX 2973, a fast growing cyanobacterial chassis for biosynthesis using light and CO2. Scientific Reports 5:8132.
Zeiler, K.G., D.A. Heacox, S.T. Toon, K.L. Kadam, and L.M. Brown. 1995. The use of microalgae for assimilation and utilization of carbon dioxide from fossil fuel-fired power plant flue gas. Energy Conversion and Management 36(6-9):707-712.
Zelitch, I. 1975. Improving the efficiency of photosynthesis. Science 188:626-633.
Zhang, S., Y. Liu, and D. A. Bryant. 2015. Metabolic engineering of Synechococcus sp. PCC 7002 to produce poly-3-hydroxybutyrate and poly-3-hydroxybutyrate-co-4-hydroxybutyrate. Metabolic Engineering 32:174-183.








































