2
Diagnosis, Ecology, and Exposure Pathways
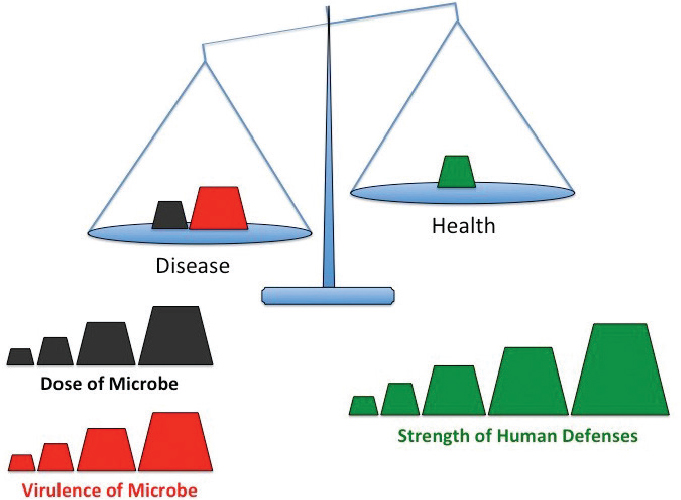
Humans coexist with an abundance of microbes, organisms so small they are invisible to the naked eye. The vast majority are benign and many are beneficial, yet everyone can name microbes that cause disease. Although it is convenient to classify microorganisms as either friend or foe, such a distinction masks more complex interactions that dictate whether the human–microbe encounter promotes disease or health. In general, the impact of exposure to a particular microbe depends on the balance of three factors: the quantity of microorganisms, their capacity to cause harm, and the strength of an individual human’s defenses (see Figure 2-1).
To tip the balance toward infection, all microbes, including Legionella, must surmount multiple challenges. First, microbes need to encounter a susceptible host. For Legionella, aerosols of contaminated water or soil can disperse these bacteria, exposing people in the vicinity. Second, the microbe has to enter the host. When inhaled or aspirated, aerosols (less than 10 µm in diameter) can carry Legionella into aveoli in the lungs. Third, to initiate an infection, microbes must breach inborn defensive barriers. In the case of Legionnaires’ disease, a respiratory tract damaged by cigarette smoke, for example, offers an increased opportunity to establish infection. Finally, to cause disease, the infecting microbe needs to inflict damage. The trauma may be direct, such as when a bacterial toxin punctures host cells, or an indirect effect of a hyperactive inflammatory response, for example. Legionnaires’ disease injures lung tissue and protective lung macrophages, a consequence of not only factors released by virulent bacteria, but also a robust inflammatory response that wreaks collateral damage. Bearing in mind these four prerequisites to infection, one can understand why combinations of exposure, Legionella loaded with an arsenal of virulence factors, and/or impaired human immune defense barriers create opportunities for some Legionella strains to establish severe lung infections. How the interplay among host defenses, Legionella pneumophila biology, and the ecology of engineered water systems alter the balance between health and disease is the focus of this chapter.
SOURCE: Adapted from Swanson et al. (2016). American Society for Microbiology, Copyright 2016; adapted with permission. No further reproduction or distribution is permitted without the prior written permission of the American Society for Microbiology.
HUMAN HOST
The majority of Legionnaires’ disease cases (from 80 to 90 percent in Europe and the United States) are linked to L. pneumophila (Beauté et al., 2013; Cross et al., 2016; Dooling et al., 2015; von Baum et al., 2008; Yu et al., 2002). However, since its discovery in the 1970s, more than 61 species and 3 subspecies of Legionella have been described, half of which have been isolated from patients (e.g., Hazel et al., 1987; Jaeger et al., 1988; Khodr et al., 2016; Vaccaro et al., 2016). In people with weakened immune systems, species other than L. pneumophila that are frequently isolated include L. micdadei, L. bozemanii, L. dumoffi and L. longbeachae (Cunha et al., 2016; Rucinski et al., 2018). In Oceania and parts of Asia, disease due to L. longbeachae approaches or exceeds that for L. pneumophila (Whiley and Bentham, 2011).
Human and financial burdens due to Legionella are substantial. Legionella species (spp.), particularly L. pneumophila, can cost from $26,000 to $38,000 per hospital admission (Collier et al., 2012). More importantly, Legionella infections can be morbid, leading to prolonged hospitalization and often intensive care admission. Mortality rates of legionellosis range between 2.9 and 33 percent (Burillo et al., 2017; Cunha et al., 2016; Gargano et al., 2017; Greenberg et al., 2006; Mykietiuk et al., 2005). Death is more likely for people who are immunocompromised (del Castillo et al., 2016; Han et al., 2015; Lanternier et al., 2017; Pedro-Botet et al., 1998; Sivagnanam et al., 2017), admitted to the intensive care unit (Chidiac et al., 2012; Cunha et al., 2016; von Baum et al., 2008), receive delayed antibiotics
(Heath et al., 1996), or who develop hospital-acquired legionellosis (Jesperson et al., 2010; Soda et al., 2015; Stout et al., 2003).
As introduced in Chapter 1 and further explained in Chapter 3, Legionella pneumonia is a significantly underreported disease (Beauté et al., 2013; Jong et al., 2010; Neil and Berkelman, 2008). Incidence varies even when legionellosis is reported, likely because of differences in local ecology, water sources, temperature and weather patterns (Fisman et al., 2005; Ricketts et al., 2018), testing availability (Pierre et al., 2017), and surveillance of and methods for reporting incident cases. In the Etiology of Pneumonia in the Community (EPIC) study evaluating community-acquired pneumonia, Legionella pneumonia was the eighth most common pathogen identified for patients requiring hospital admission in five hospitals in Nashville, Tennessee, and Chicago, Illinois (Jain et al., 2015). Even this high ranking was conservative, as the EPIC study data only included assessment of L. pneumophila serogroup 1 and excluded high-risk immunosuppressed patients or those with prior hospitalization. The Competence Network for Community-Acquired Pneumonia (CAPNETZ) study in Germany, which used culture, urinary antigen, and polymerase chain reaction (PCR) testing of samples, found that 97 out of 941 (9.6 percent) of all community-acquired pneumonia cases were due to legionellosis, the majority of which were attributed to L. pneumophila (von Baum et al., 2008). Non-pneumophila presentations of legionellosis, limited testing, use of empiric therapy, and lack of consensus for the epidemiological definition of presentations such as Pontiac fever (Tossa et al., 2006) further contribute to under-reporting (Whiley et al., 2014). Furthermore, rates of disease caused by non-pneumophila Legionella spp. and non-serogroup 1 L. pneumophila, for which there are more limited diagnostic modalities, are also thought to be underestimated (Benin et al., 2002; Lode et al., 1987; Muder and Victor, 2002). Serological testing of blood donors indicates that exposure to Legionella may be higher than generally appreciated, with seroprevalence ranging between 4 and 22 percent and up to approximately 40 percent in some cities or among those with high-risk occupations (Borella et al., 2008; Coniglio et al., 2009; Nadaraja et al., 1987; Rudbeck et al., 2008; Valcin¸ a et al., 2015).
Increasing Incidence
Studies worldwide have shown increasing incidence of Legionella cases (Beauté, 2017; Burillo et al., 2017; Neil and Berkelman, 2008). In the United States, reported cases increased six-fold from 2000 to 2018 (see Figure 1-2). The causes of this increase are not well characterized, but are thought to be multifactorial. Methods to detect Legionella in clinical samples are now both easier to perform (e.g., urinary assays) and more readily available outside of large academic laboratories, making their use increasingly common (Pierre et al., 2017). Community-acquired pneumonia national guidelines (Bradley et al., 2011; Mandell et al., 2007) and prediction tools (Cunha, 1998; Fiumefreddo et al., 2009; Miyashita et al., 2017) have both helped increase awareness of Legionella. However, these guidelines, aimed at community practioners, were created to streamline diagnostic work-up and antibiotic management. Indeed, most current guidelines recommend that even low-risk patients receive empiric antibiotics that include either a macrolide (in combination with a beta-lactam or cephalosporin or as the primary agent, depending on host risk factors) or a respiratory flouroquinolone (moxifloxacin or levofloxain) (Yu et al., 2004). These agents not only target many common causes of pneumonia, such as Streptococcus pneumonia and Moraxella catarhallis, but also cover atypical pneumonia pathogens including Mycoplasma pneumoniae and Legionella spp. Although such guidelines promote prompt administration of appropriate empiric therapy for patients with pneumonia, providers of such early antibiotic therapy may
actually be less apt to pursue diagnostic testing for legionellosis. This tendency is particularly true for mild cases, as some guidelines only recommend testing for patients who have severe disease or are admitted to the intensive care unit (Mandell et al., 2007).
Changing demographics, such as the aging population, may contribute to the rise in disease incidence as well. From 1970 to 2018, the median age in the United States has increased by nearly ten years, such that Americans aged 65 and older now make up a larger portion of the U.S. population (CDC, 2013). And the number of elderly Americans is predicted to increase. Immune senescence of both innate and adaptive immunity plays an important role in increased risk among the elderly population (Boe et al., 2017). Enhanced survival of high-risk patients (e.g., those with an underlying condition such as cancer, cardiac disease, or lung disease) may also contribute to the increased incidence trend. Likewise, the increasing number of patients with compromised immunity due to immunosuppressive therapies and prolonged survival among higher-risk immunosuppressed patients may contribute to increasing legionellosis incidence. Although likely an underestimate, current data suggest that at least 2.7 percent of Americans consider themselves immunosuppressed (Harpaz et al., 2016), indicating nearly 9 million at-risk individuals in the United States by this criterion alone. With the increasing number of agents that modify immune responses, improvements in survival, and the number of conditions that are currently treated with immunosuppressive therapy, this susceptible population is expected to increase. Specific immuncompromised populations are known to be growing. For example, the United Network of Organ Sharing reported a doubling of the number of patients receiving a solid organ transplant over the past 20 years.1 Likewise, the National Cancer Institute estimates that the number of cancer survivors will increase by 30 percent over the next decade.2
Increasing population density in cities served by aging, centralized water systems (and including more cooling towers) may elevate the risk of legionellosis. Indeed, the American Society of Civil Engineers gave the U.S. water infrastructure a “D” rating (ASCE, 2017), noting that many pipes laid in the mid-20th century are now beyond their expected lifespan, increasing their risk for main breaks and intrusion, corrosion-enhanced biofilm development, and colonization with Legionella. Exposure in daily life may be more frequent due to increasing contact with water products and water devices, particularly those ideally suited for Legionella growth (e.g., cooling towers; hosing, faucets, and showerheads; hot tubs, Jacuzzis, and spas [collectively called hot tubs in this report]; humidifiers; fountains).
Additionally, climatic changes, including increased rainfall and global temperatures, have been linked to increasing disease incidence, either directly or through increased use of water sources linked to legionellosis. Most climate work has focused on the effects of temperature and rainfall events. Clear seasonality to L. pneumophila exposure has been established by multiple studies (ECDC, 2013; Marston et al., 1994). For example, a PCR screen of more than 44,000 respiratory specimens over a recent eight-year period in Rochester, Minnesota, documented annual peaks during the warm, humid months (Rucinski et al., 2018). Likewise, risk of disease increased during warm, wet periods in the mid-Atlantic region of the United States, as well as The Netherlands, Spain, and Taiwan (Chen et al., 2014; Fisman et al., 2005; Garcia-Vidal et al., 2013, reviewed by Walker, 2018; Hicks et al., 2007; Karagiannis et al., 2009; Simmering et al., 2017). Cassell et al. (2018) suggest that precipitation is associated with the risk of sporadic Legionnaires’ disease, noting a 48 percent increased risk of legionellosis two weeks after a 5-mm average increase in rainfall. With increasing global
___________________
1 See https://optn.transplant.hrsa.gov/data/view-data-reports/national-data/#.
2 See https://www.cancer.gov/about-cancer/understanding/statistics.
temperatures, more precipitation and flooding in some regions, and rising sea levels, there is concern that Legionella and other waterborne infections will continue to increase.
Worldwide, the actual burden of Legionnaires’ disease is generally acknowledged to be underreported as a consequence of the generally low rate of diagnostic testing coupled with the reliance on a diagnostic test that is highly specific for a single serogroup of L. pneumophila (Dooling, 2015; Mercante and Winchell, 2015; Phin et al., 2014; St-Martin et al., 2013; von Baum et al., 2008). During a three-year period in Germany, about 10 percent of Legionnaires’ disease patients were infected with species other than L. pneumophila (von Baum et al., 2008). Over a ten-year period in Denmark, 40 percent of the Legionnaires’ disease cases that were confirmed by laboratory culture were caused by L. pneumophila that were not serogroup 1 (St-Martin et al., 2013). Similarly, in the United States, for the Legionnaires’ disease cases reported from 2011 to 2013 to the Centers for Disease Control and Prevention’s (CDC) Active Bacterial Core Surveillance network, 9 percent of the 140 culture-confirmed cases were due to non-serogroup 1 L. pneumophila (Dooling et al., 2015). Also troubling is the higher mortality among patients infected with these non-serogroup 1 strains compared with serogroup 1 L. pneumophila (Marston et al., 1994; Mercante and Winchell, 2015; St-Martin et al., 2013). These differences could be because of differences in those at risk for these pathogens or because of the lack of commonly available diagnostic tools, which delays identification. Finally, sole reliance on the urine antigen test hampers efforts to recognize and interrupt outbreaks, as epidemiological investigations require discriminatory genetic tests of clinical and environmental L. pneumophila isolates (Mercante and Winchell, 2015).
Risk Factors for Legionella Disease
Numerous factors are linked to increased risk of legionellosis, varying from host factors to exposure factors. Indeed, in large outbreaks not everyone who is exposed to Legionella develops disease (Bartram, 2007; Phin et al., 2014). Male adults are at higher risk (Cunha et al., 2016; MacIntyre et al., 2018; WHO, 2018). Indeed, in the United States the incidence of reported cases of legionellosis in men (2.31/100,000) was approximately 50 percent higher than in females (1.50/100k) in 2016. Age is also commonly identified as a risk factor for legionellosis, with most studies suggesting risk begins increasing at approximately age 40 to 50 years (Bartram, 2007; Farnham et al., 2014; Sopena et al., 2007). Data on children are more sparse, but suggest that children may be less likely to develop severe infections (Greenberg et al., 2006; Muldoon et al., 1981; Yu and Lee, 2010). In one review of case reports, the majority of children who became ill had other at-risk diseases (e.g., cancer) and more than half were under the age of one (Greenberg et al., 2006). More than 70 percent of pediatric legionellosis cases may be hospital-acquired, suggesting either less clinical disease or underdiagnosis in the community (Alexander et al., 2008). Neonates may be at highest risk for hospital-acquired legionellosis because of both increased exposures and their weaker immune status (Levy and Rubin, 1998). Although infrequently reported, water births have been linked to some cases of neonatal legionellosis (Franzin et al., 2001; Granseth et al., 2017).
Populations with jobs that increase occupational exposure to water are also at risk of legionellosis (Principe et al., 2017). Among these populations are water-service providers, maintenance workers, wastewater and cleaning personnel, and workers in industries that use industrial water sprayers (e.g., paper and textile mills, plastic molding factories).
Another classical demographic risk factor for legionellosis is impaired immunity, either through anatomic changes to the airway or weakened barriers to respiratory pathogens.
Indeed, age-related immune senescence is thought to be an important reason why rates are higher among older patients. Smoking is a clear dose-dependent risk factor and possibly contributes to the higher incidence of cases reported for males than females. Smoking also changes the airway epithelium, perturbs pulmonary cilia function, decreases airway clearance, and alters the aerodigestive microbiome, factors each thought to increase the risk for bacterial pneumonias, including those caused by Legionella spp. (Arcavi and Benowitz, 2004; Gao et al., 2014; Morris et al., 2013). Not only are those who smoke at higher risk than those who have never smoked, but those who have smoked more than 20 cigarettes per day for more than 20 years have a risk 25 times greater than that of non-smokers as well as a higher incidence of disease during outbreaks (Che et al., 2008). Risk and severity of illness may be further increased by smoking-related complications such as chronic obstructive pulmonary disease or emphysema (El-Ebiary et al., 1997). Even exposure to second-hand smoke has been suggested to increase risk (Wang et al., 1995).
Another population vulnerable to Legionella pneumonia is patients at increased risk of aspiration, such as those with neuromuscular diseases, the elderly, and neonates (Blatt et al., 1993; Wei, 2014). (Indeed, studies suggest that episodes of silent aspiration are significantly more common among elderly patients when compared to age-matched controls [Kikuchi et al., 1994]). Researchers have hypothesized that the “microaspiration” that occurs during drinking or with particular clinical conditions may deliver Legionella to the lung and cause pneumonia (Lee and Ryu, 2018; Marrie et al., 1991). The oropharynx (Jaresova et al., 2006) and dental plaques (Tesauro et al., 2018) may also be colonized with Legionella, which would increase the likelihood of aspiration. Still, exposure pathways related to aspiration are more difficult to investigate unless clearly linked to feeding tubes or other mechanical methods associated with tap water and aspiration (Dournon et al., 1982; Marrie et al., 1991; Muder et al., 1992; Venezia et al., 1994; Yu, 1993).
Patients with known impaired immune function, including those with organ dysfunction, are also at increased risk for legionellosis. Patients undergoing treatment for cancer and those who have received a solid organ transplant may be at highest risk because of the depth and length of immunosuppression required (del Castillo et al., 2016; Jacobson et al., 2008; Lanternier et al., 2017; Sivagnanam and Pergam, 2016). These patients are also at increased risk for non-pneumophila legionellosis and non-serogroup 1 L. pneumophila infections (Ampel and Wing, 1990; Dowling et al., 1984; Knirsch et al., 2000; Muder and Victor, 2002; Singh et al., 2004). Particular immunosuppressive agents have been linked to legionellosis, chiefly glucocorticoids such as prednisone (Htwe and Khardori, 2017). Patients receiving tumor necrosis factor inhibitors for rheumatologic and inflammatory bowel diseases are also at increased risk for legionellosis (Lanternier et al., 2013). Likewise, patients with renal dysfunction (including those on dialysis), liver disease, lung disease, and known cardiac dysfunction are more susceptible to legionellosis (Chidiac et al., 2012; Ongut et al., 2003; Viasus et al., 2013). Immunosuppressed patients also have increased severity of disease including intensive care unit (ICU) admission, intubation, and death.
Genetic predisposition may account for enhanced susceptibility to disease from Legionella spp. in those with or without other risk factors (Berrington and Hawn, 2013). Legionellosis is linked to genetic polymorphisms in three human Toll-like receptors (i.e., TLR-4, TLR-5, and TLR-6), components of the innate immune system that recognizes pathogen-associated molecular pattern (PAMP) molecules (Hawn et al., 2003, 2005; Misch et al., 2013). Also associated with an increased risk for legionellosis is a common haplotype of the GMP-AMP synthase-stimulator of interferon genes (STING) pathway (HAQ TMEM173/STING), which is central for innate immune sensing of bacterial infections (Ruiz-Moreno et al., 2018).
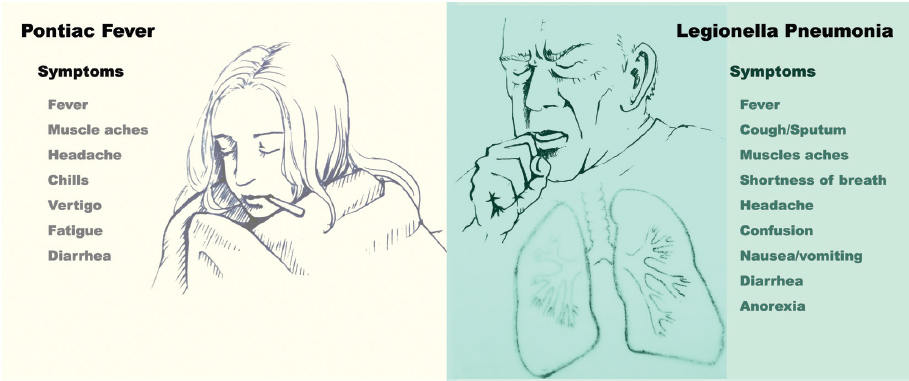

SOURCE: Courtesy of Kyoko Kurosawa.
Clinical Manifestations of Legionellosis
Legionella spp. cause clinically significant disease among susceptible human hosts. The most common manifestations are pneumonia (i.e., classical Legionnaires’ disease) and Pontiac fever. Diagnosis can be challenging as many clinical signs and symptoms typical of legionellosis are often found in both infectious and non-infectious diseases (see Figure 2-2) (Cunha, 1998; Phin et al., 2014). More rarely, Legionella spp. have been associated with skin and soft-tissue infections, bacteremia, endocarditis, and septic arthritis (Banderet et al., 2017; Heriot et al., 2014; Kilborn et al., 1992; Pearce et al., 2011; Qin et al., 2002).
Worldwide, Legionella pneumonia is the most common manifestation of legionellosis. Legionella pneumonia is often classified as an “atypical pneumonia” along with those caused by bacterial pathogens such as Chlamydia pneumoniae and Mycoplasma pneumoniae. When compared with common bacterial pneumonia agents, atypical pneumonias may present common symptoms and radiologic findings and less common presentations (e.g., diffuse interstitial patterns on chest x-ray). Clinical and radiologic findings cannot distinguish between these pathogens and other causes of community-acquired pneumonia. However, they respond to antibiotic classes and agents that primarily target intracellular infections (Sharma et al., 2017). Following exposure, L. pneumophila has an incubation period of approximately two to ten days (Bartram, 2007). About 10 percent of cases have an incubation period longer than ten days, such that case information should be collected for a minimum of 14 days prior to onset of symptoms for community cases.3 Likewise, incubation periods may be longer for hospitalized populations, which often include immunosuppressed hosts (Bargellini et al., 2013). Finally, the incubation period may also be dose-dependent (Prasad et al., 2017).
Initial symptoms of Legionnaires’ disease typically include fever, cough, and myalgias. Other commonly reported symptoms include headache, confusion, shortness of breath, sputum production, anorexia, nausea, and diarrhea; patients with community-acquired
___________________
pneumonia who present with neurologic and gastrointestinal symptoms may be more likely to have legionellosis (CDC, 2017; Cunha, 1998). Some patients with Legionella pneumonia present with acute respiratory failure, hypotension, and sepsis-like signs that can mimic other common causes of bacterial pneumonia (e.g., Streptococcus pneumoniae). In contrast, patients who are immunosuppressed may present without fever, cough, or other more typical symptoms (del Castillo et al., 2016; Sivagnanam and Pergam, 2016).
Pontiac fever presentation is less specific and therefore less frequently reported. It is often described as a “flu-like” illness, with fever, headaches, and myalgias as the primary symptoms; chills, vertigo, diarrhea, and physical weakness or lack of energy (asthenia) are other symptoms (Tossa et al., 2006). Pontiac fever cases are defined in part by their absence of pneumonia. Since many other illnesses resemble Pontiac fever, the diagnosis usually relies on the recognition of typical clinical features during an outbreak situation; therefore, sporadic cases are likely to be missed (Murdoch, 2003). Pontiac fever in particular may be underdiagnosed in children, whose febrile illnesses are frequent and often self-limited (Qin et al., 2002).
It is unclear why patients develop Pontiac fever rather than pneumonia; consequently, the pathogenesis of the disease remains unclear. Several pathways for Pontiac fever have been hypothesized, including bacterial toxins, allergic responses, and exposure and reaction to Legionella-carrying ameobae (Edelstein, 2007). A self-limiting disease, Pontiac fever does not require treatment with antibotic therapy, leading some to hypothesize that the disease is not directly related to infection by these bacteria. At the same time, there have been outbreaks where patients who develop Pontiac fever have positive urinary antigen testing, suggesting that the disease is associated with ingestion or inhalation of either live or dead microorganisms (Burnsed, 2007). Mechanism of exposure may also play a role, as some recreational outbreaks have been linked to both Legionella pneumonia and Pontiac fever, whereas others are tied only to Pontiac fever (Euser et al., 2010; Leoni et al., 2018).
Not all Legionella spp. are thought to cause Pontiac fever, as the disease is most frequently linked to L. pneumophila exposures. Similar to pneumonia, however, non-pneumophila species such as L. feelii, L. micdadei, and L. bozemanii can also cause Pontiac fever (Cramp et al., 2010; Fentersheib et al., 1990; Fields et al., 2001; Herwaldt et al., 1984; Huhn et al., 2002). Only one study has suggested Pontaic fever was associated with exposure to water sources with higher concentrations of Legionella spp. (i.e., more than 103 colony-forming units per liter [CFU/L]) (Remen et al., 2011). Interestingly, in the same study, younger nursing staff were at higher risk, suggesting immune responses to prolonged or prior exposures may provide protection from this form of disease. Species-specific strains or particular bacterial activities themselves may be critical to disease outcomes. For example, a L. fellii serogroup with a monopolar flagellum associated with Legionella pneumonia showed a higher cell infection rate, stronger internalization by host cells, and greater cytotoxicity in vitro than a different L. fellii serogroup without a flagellum that was associated with Pontiac fever (Wang et al., 2015a). The lower risk associated with Pontiac fever, the limited diagnostic work-up, and the rarity of documented positive cultures linked to the disease make studies of pathogenesis difficult. Regardless of the outstanding questions concerning the pathogenesis of Pontiac fever, it is clear that exposure to water or soil contaminated with Legionella spp. is required for the development of the disease.
Diagnosis of Legionellosis
Presenting Laboratory Findings
Beyond clinical symptoms, laboratory findings may point to a diagnosis of Legionella pneumonia. Patients can present with either leukocytosis or leukopenia, hyponatremia,
elevated liver enzymes, and renal dysfunction. Non-specific blood tests that suggest inflammation (e.g., C-reactive protein) can also be elevated (Fiumefreddo et al., 2009). Clinical prediction tools, such as the Winthrop-University Hospital criteria (Cunha et al., 1998), Community-Based Pneumonia Incidence Study Group score (Fernández-Sabé et al., 2003), Japan Respiratory Society score (Yanagihara et al., 2001), and a six-parameter clinical score developed by Fiumfreddo and colleagues (2009) are thought to have poor sensitivity for legionellosis, but may be useful for their negative predictive value (Miyashita et al., 2017).
Radiology
There are no unique radiologic findings specific for legionellosis. For Legionella pneumonia, chest radiographs often demonstrate focal infiltrates or consolidations consistent with pneumonia (Poirier et al., 2017). Computed tomography (CT) findings may show multi-lobar or air-space disease with associated ground-glass opacities; and lymphadenopathy and pleural effusions may be seen as well (Mittal et al., 2017). Among the immunocompromised, Legionella can present as pulmonary nodules with or without cavitation (del Castillo et al., 2016; Mittal et al., 2017). Patients with Pontiac fever are defined by their lack of findings on radiologic imaging.
Legionella-Specific Diagnostics
Culture. Cultures can be collected from pulmonary and extra-pulmonary sites (Mercante and Winchell, 2015). Legionella requires special culture media, most commonly buffered charcoal yeast extract agar (see Descours et al., 2014, for a recent analysis of different media). Growth usually occurs within three to five days, although two weeks may be required, as antibiotics used to reduce background respiratory microbiota can inhibit Legionella growth (Pierre et al., 2017). Growth from clinical samples may be limited or delayed if, prior to specimen collection, patients have been given antibiotics targeting Legionella spp. Non-pneumophila Legionella spp. tend to be more fastidious, may require longer incubation times (Mercante and Winchell, 2015), may be inhibited by some culture media (Lee et al., 1993), and require considerable technical laboratory expertise to culture (Lucas et al., 2011). Most culture-based systems are optimized for L. pneumophila and may limit growth of non-pneumophila species. Another complication arises when patients have multiple strains of Legionella during an active infection (Coscolla et al., 2014; Zhang et al., 2014). Because of these challenges, most hospital-based laboratories do not routinely test for Legionella spp. by culture. Yet, cultures are critically important to epidemiologic investigations, as they allow for analysis of relatedness within clusters and between clinical and environmental samples. In the United States culture diagnosis has declined from greater than than 60 percent in the early 1990s to 5 percent from 2005 to 2009 (Mercante and Winchell, 2015). In Europe, where cultures are more often utilized, 79 percent of culture-confirmed cases were reported as L. pneumophila serogroup 1 (ECDC, 2019).
Urinary Assay. The Legionella urinary antigen test (UAT) is the most frequently used method for legionellosis diagnosis in the United States (Mercante and Winchell, 2015). The UAT is routinely available on-site at 25 percent of acute-care hospitals (Garrison et al., 2014) and commercial laboratories. The test is popular because of the rapid turn-around time for on-site laboratories, ease of use, high sensitivity, and the ability to identify L. pneumophila serogroup 1, the most prevalent Legionella spp. associated with clinical disease, without the need for invasive procedures. However, the UAT can be negative very early in the disease and is of limited value in patients who cannot produce urine (anuric), e.g., due to kidney
failure. On the other hand, the UAT can remain positive for months after an infection, particularly in immunosuppressed patient populations (Kashuba and Ballow, 1996; Kohler et al., 1984; Munoz et al., 2009). A major limitation of diagnosis strategies that focus on the UAT alone is their failure to detect important non-serogroup 1 L. pneumophila infections and non-pneumophila infections.
Serology. Acute and convalescent titers for Legionella may be helpful in documenting Legionella exposures, but have limited sensitivity for confirmation of Legionnaires’ disease (Botelho-Nevers et al., 2016; Plouffe et al., 1995). Indeed, up to 20 to 30 percent of patients with proven Legionella may not mount an antibody response sufficient for diagnosis (Benz-Lemoine et al., 1991). For patients with altered immunity, the sensitivity and the specificity of seroconversion to non-pneumophila Legionella spp. is unclear (Reller et al., 2003). Even with the presense of high levels of antibody, one cannot differentiate recent versus past exposure, limiting the use of serology in acute infections (Mercante and Winchell, 2015). Serological testing for Legionella can also cross-react with Coxiella burnetti (the agent of Q fever) and Mycoplasma pneumoniae, among others (Boswell et al., 1996; Musso and Raoult, 1997).
Serology can provide important information for epidemiologic investigations. In a large outbreak in Norway, acute-phase Legionella tests (i.e., culture, UAT, and PCR) detected about 56 cases, whereas serology detected an additional 47 cases (Simonsen et al., 2015). Thus, serology may identify individuals with less severe disease and symptoms (e.g., Pontiac fever) who might otherwise be missed during large industrial exposures and outbreaks. Nevertheless, the use of serology diagnosis for either sporadic disease or outbreak investigation has declined (Mercante and Winchell, 2015).
Direct Fluorescent Antibody Testing. Direct fluorescent antibody (DFA) testing is infrequently used to diagnose Legionella. The sensitivity of DFA can be very low (11 to 40 percent) (Hayden et al., 2001; She et al., 2007), and it often does not detect non-pneumophila Legionella spp. (Reller et al., 2003). Although some argue DFA has high specificity when positive, caution is needed as a positive DFA in the absence of other supportive evidence is thought to be insufficient for Legionella diagnosis (Haldane et al., 1993; Reller et al., 2003).
Molecular Testing. PCR and other nucleic acid amplification tests are highly sensitive assays for lower respiratory tract specimens but are primarily available in referral or research laboratories. Most published studies utilize PCR testing that targets the gene encoding the macrophage infectivity potentiator (mip) surface protein of L. pneumophila (Phin et al., 2014). One study found a four-fold increase in Legionella case detection with PCR testing of lower-respiratory specimens compared to culture (Murdoch et al., 2013), and another found only 40 percent of PCR-positive specimens were also culture-positive (Rucinski et al., 2018). Compared to UAT, PCR tests may be more sensitive, detecting an additional 18 to 30 percent of cases (Avni et al., 2016). PCR may have the advantage for Legionella diagnosis in patients already on empiric therapy with Legionella active antibiotics, which limit bacterial growth by culture.
Most PCR methods currently detect serogroup 1 strains but cannot distinguish among L. pneumophila serogroups (Benitez and Winchell, 2013), and most probes target L. pneumophila specifically. There are newer PCR assays that target common non-pneumophila species including L. longbeachae, L. micdadei, and others (Cross et al., 2016). Broader Legionella spp. PCR tests have been developed (Benitez and Winchell, 2013; Chen et al.,
2015), but to date only one test has been approved by the U.S. Food and Drug Administration (FDA) for testing of clinical samples. The BioFire® FilmArray® Pneumonia Panel is a multiplex PCR respiratory panel that detects 33 different respiratory pathogens, including L. pneumophila. The assay was approved in November 2018 for sputum, endotracheal aspirates, and bronchoalveolar lavage (or mini-BAL) lower-respiratory tract samples. PCR assays have been licensed in Europe, but most available assays are primarily laboratory-developed assays of variable sensitivity (Ricci et al., 2018). Of note, PCR is less sensitive for non-respiratory samples (e.g., blood; Avni et al., 2016; von Baum et al., 2008).
Pharmaceutical Therapy
Legionella spp. are intracellular pathogens that not only avoid phagosome–lysosome fusion and degradation, but also replicate within alveolar macrophages and epithelial cells (Newton et al., 2010). Therefore, the mainstay of drug therapy for legionellosis are antibiotics that target the intracellular space. Guidelines in the United States and in Europe recommend macrolides and fluoroquinolones as first-line therapy (Mandel et al., 2007; Pea, 2018; Woodhead et al., 2011). Macrolides work primarily by disrupting the 50S subunit of bacterial ribosomes, thereby inhibiting protein synthesis, which is critical for the microbe’s survival. Most early studies evaluated the macrolides erythromycin and clarithromycin, but azithromycin is the preferred agent as it is better tolerated (Langley et al., 2004), associated with fewer side effects than other macrolides, and is the most effective macrolide in animal models (Fitzgeorge et al., 1990; Plouffe et al., 2003).
Compared with macrolides, fluoroquinolones, which inhibit bacterial DNA gyrase and topoisomerase IV enzymes, are more potent against Legionella spp. in both in vitro and in vivo models of infection (Pedro-Botet and Yu, 2009). Currently, there is no randomized clinical trial comparing macrolides to fluoroquinolones for antibiotic therapy of legionellosis. However, in non-randomized, observational studies, fluoroquinolones were more effective than macrolides (i.e., erythromycin and clarithromycin) in fever resolution and length of hospitalization (Burdet et al., 2014; Garcia-Vidal et al., 2017; Griffin et al., 2010). Despite therapy, mortality rates for cases treated with both drug classes remain around 10 percent (Burdet et al., 2014).
Beyond macrolides and fluoroquinolones, other agents such as rifampin/rifampicin, tetracyclines, and trimethoprim-sulfa are used, sometimes in combination (Pedro-Botet and Yu, 2009). Although data are limited, some experts recommend that patients with severe clinical illness may benefit from combination therapy (Nakamura et al., 2009; Varner et al., 2011).
Selection for antibiotic-resistant Legionella is considered very limited, as humans are a dead end in the life cycle of Legionella spp. Indeed, just one case of suspected person-to-person tranmission has been reported in more than 40 years (Correia et al., 2016). Although infrequent, there are data demonstrating both clinical failures and Legionella that, under macrolide pressure, have documented macrolide resistance primarily through mutations in the bacterial ribosome (Descours et al., 2017; Dowling et al., 1985; Nielsen et al., 2000). To date, fully resistant strains have not been observed in wild-type isolates (Vandewalle-Capo et al., 2017). There is also concern that widespread use of azithromycin for other conditions among hospitalized patients could lead to increased resistance in Legionella (Torre et al., 2018). Similar to macrolides, rare resistance to fluoroquinolones has been identified, primarily through changes in bacterial DNA gryrase genes (Almahmoud et al., 2009; Bruin et al., 2014; Jonas et al., 2003).
LEGIONELLA SPECIES AND STRAINS
The Legionella genus is in constant flux, blurring the boundaries between species. Variation is generated by high rates of genetic exchange among species as well as by homologous recombination between exogenous DNA and the bacterial chromosome (Gomez-Valero et al., 2014, 2019; Joseph et al., 2016; Sanchez-Buso, 2014). Within the genus, both genome size and guanine-cytosine (GC) content vary widely (see Figure 2-3). Furthermore, among the pangenome of 80 sequenced Legionella strains representing 58 species, only 6 percent of the genes were encoded by each genome examined (Gomez-Valero et al., 2019). For example, although the flagellar regulon contributes to L. pneumophila infection of macrophages and amoebae (reviewed by Appelt and Heuner, 2017), flagella genes were absent in 23 of 80 sequenced genomes from 58 Legionella spp. (Gomez-Valero et al., 2019).
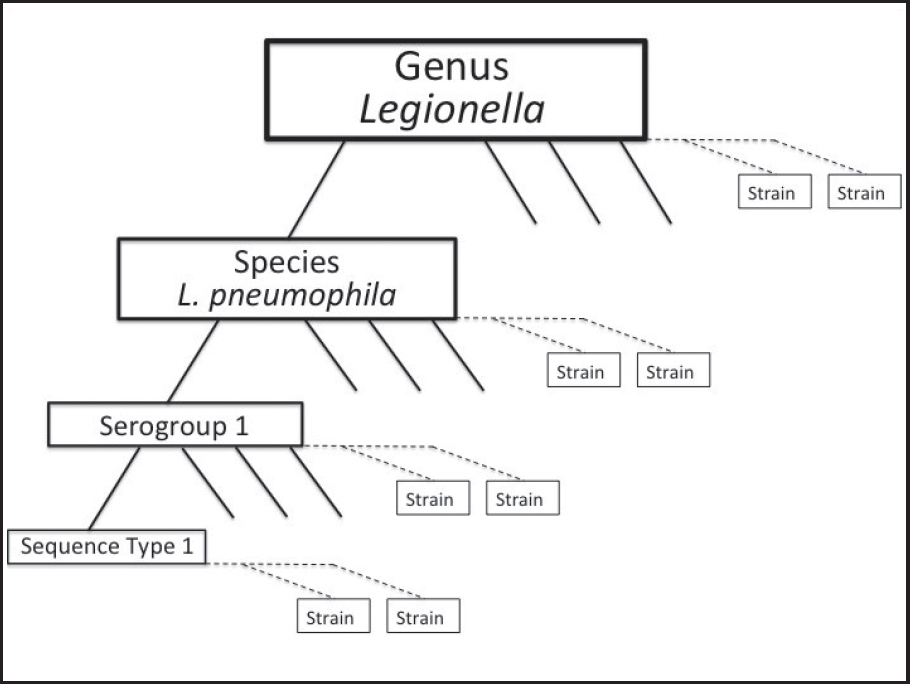
To further guide diagnosis and identification of the environmental source of the infectious bacteria, L. pneumophila strains can be classified by several techniques. Terms commonly used to describe particular isolates of bacteria are explained in Figure 2-4.
L. pneumophila can be divided into serogroups according to the structure of its lipopolysaccharide (LPS), a component of the outer membrane of these Gram-negative bacteria and the predominant antigen recognized by the human immune system (Ciesielski et al., 1986; Conlan and Ashworth, 1986). Although 16 distinct “serogroups” of L. pneumophila have been identified, the majority of disease cases are caused by serogroup 1 strains. For example, in Europe from 2011 to 2015, serogroup 1 strains were associated with 83 percent of the cases confirmed by culture (Beauté, 2017), a frequency similar to that reported by a 2002 international prospective study (Yu et al., 2002). More rarely, strains of other serogroups are isolated from Legionnaires’ disease patients, and the pattern varies by region. In two international studies, serogroups 3, 4, 5, 6, and 10 were each associated with less than 2 percent of culture-confirmed cases (Beauté, 2017; Yu et al., 2002). In Denmark from 1993 to 2006, serogroup 3 was identified in 23 percent and serogroup 6 in 5 percent of 419 culture-confirmed cases (St Martin et al., 2013). In contrast, in Ontario, Canada, serogroup 6 L. pneumophila accounted for 47 percent of the disease cases not caused by serogroup 1 strains (Khan et al., 2013). In Michigan, serogroup 6 L. pneumophila also predominated in a surveillance study of premise plumbing one year after the 2014-2015 Legionnaires’ disease outbreaks in Flint (Byrne et al., 2018).
L. pneumophila is the most prevalent reported species of Legionella in building water systems, whereas non-pneumophila strains show more geographic variation. Recent surveillance studies of hundreds of buildings in Germany and Hungary identified 58 to 84 percent of the isolates as L. pneumophila (Barna et al., 2016; Dilger et al., 2018; Kruse et al., 2016). Temperature is likely one factor that influences the prevalence of particular species. For example, in a study of more than 13,000 warm-water systems in southern Germany, L. anisa composed 18 percent of the total isolates obtained from water at 20°C but only 8 percent at 60°C (Dilger et al., 2018).
The clinical predominance of L. pneumophila serogroup 1 does not simply reflect a relative abundance in the environment (Doleans et al., 2004; reviewed by Mercante and Winchell, 2015). It also appears to grow better in humans than other Legionella species. For example, compared with seven other serogroups, the composition of serogroup 1 LPS equips L. pneumophila to resist killing by the alternate complement pathway (Khan et al., 2013), a key barrier of the human innate immune system. The serogroup 1 LPS O-antigen is also extremely hydrophobic (Zähringer et al., 1995), which may contribute to L. pneumophila survival in the human lung via evasion of toxic lysosomes (Fernandez-Moreira et al., 2006) or perhaps within aerosols. Two other observations suggest that the serogroup 1 LPS
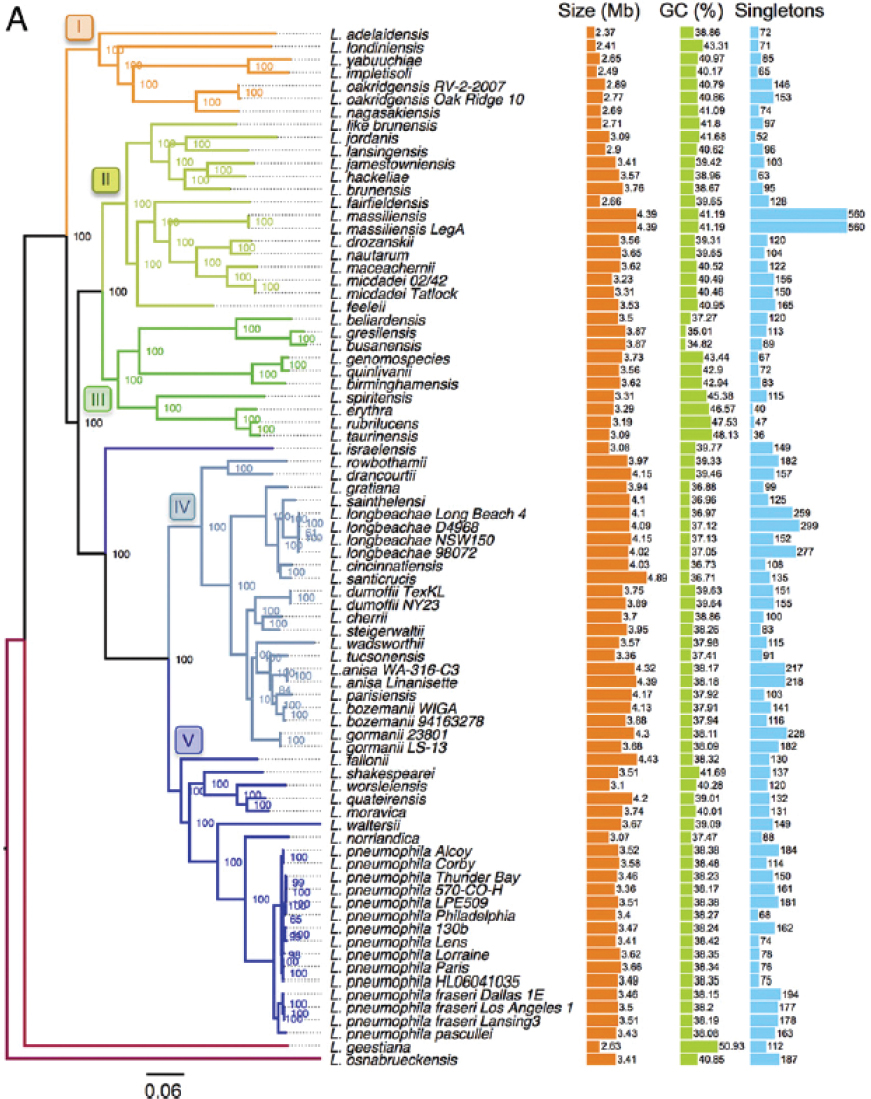
SOURCE: Gomez-Valero et al. (2019).
enhances L. pneumophila virulence. Unlike environmental strains, the majority of clinical isolates display a distinct LPS epitope encoded by the lag-1 gene and recognized by particular monoclonal antibodies (MAb2 and MAb3/1), a structural feature that is used for diagnosis and risk-assessment (Kozak et al., 2009). It is also striking that the only genes shared by greater than 200 serogroup 1 strains comprise a locus dedicated to LPS biosynthesis (Cazalet et al., 2008). A cluster of these serogroup 1 LPS genes can spread horizontally among the legionellae (Cazalet et al., 2008; Merault et al., 2011), expanding the genetic diversity of serogroup 1 strains. Moreover, the LPS locus of at least two L. pneumophila serogroup 1 lineages exhibits an elevated rate of genetic exchange (David et al., 2017), further expanding their diversity. Whether serogroup 1 strains are better equipped to survive in aerosols or to colonize engineered water systems warrants investigation.
Because the L. pneumophila genome is dynamic, related strains of the same serogroup must be distinguished using molecular methods. Since 2005, the international Legionella community has applied a multi-locus DNA sequence typing scheme and online tools (Gaia et al., 2005; Ratzow et al., 2007; Underwood et al., 2006). Currently, more than 2,000
L. pneumophila sequence types (STs) can be distinguished based on the nucleotide sequence of seven alleles. By combining sequence-based typing with classical epidemiology, investigators can perform trace-back studies to identify outbreak sources, as well as assess regional patterns and persistence in particular environments. For example, Kozak-Muizniek and colleagues (2016) conducted a comprehensive phylogenetic analysis of hundreds of L. pneumophila strains collected in the United States from outbreaks, sporadic clinical cases, and environmental surveillance, all of which were submitted to the CDC over a 30-year period.
Worldwide, the most prevalent sequence type is ST1 within serogroup 1. In the United States from 1982 to 2012, this sequence type accounted for 49 percent of the L. pneumophila strains obtained from water and 25 percent of the sporadic disease isolates (Kozak-Muizniek et al., 2016). Likewise, in China over a seven-year period, 49 percent of environmental isolates were ST1 L. pneumophila (Qin et al., 2014). Multi-year surveillance studies in England, France, and Wales identified ST1 L. pneumophila as composing approximately 20 percent of environmental strains (Cassier et al., 2015; Harrison et al., 2009), and a Japanese analysis discovered that 74 percent of L. pneumophila isolates from cooling towers were ST1 (Amemura-Maekawa et al., 2012). Nevertheless, even among ST1 serogroup 1 L. pneumophila genomes, substantial diversity is generated by genetic recombination, as evident from whole genome sequence analysis of more than 200 ST1 strains (David et al., 2017). Thus sequence-typing alone is not sufficient to establish epidemiological associations of ST1 strains.
Nonetheless, sequence-based typing of large strain collections has identified regional variations in the endemic L. pneumophila populations and infection patterns. For example, ST1 strains caused multiple outbreaks in Belgium from 2000 to 2010 (Vekens et al., 2012), but the first outbreak in the United States was not recorded until 2012 (Kozak-Muiznieks et al., 2014). And although ST222 strains of L. pneumophila appear to be endemic in the northeastern regions of the United States and Canada, this type has been reported only recently and just once in three other countries (Byrne et al., 2018; Kozak et al., 2009; Kozak-Muiznieks et al., 2016; Tijet et al., 2010). In Western Europe, the L. pneumophila strain type most frequently isolated from patients is ST47 (also known as the Lorraine strain), yet no Asian or U.S. case has been attributed to this genotype (reviewed by Kozak-Muizniek et al., 2016). The biological basis for either the geographic distribution or the clinical prevalence of particular strains of legionellae remains to be determined. Knowledge of the genetic attributes that increase L. pneumophila fitness in distinct environments could guide detection and remediation strategies for particular geographic regions.
Despite the diversity of legionellae, research on the virulence and resilience mechanisms is focused on L. pneumophila serogroup 1 strains. Indeed, a literature search for titles and abstracts that contain “pneumophila” and “serogroup 1” identified more than 450 research articles; in contrast, “pneumophila” and “serogroup 6” identified just 64. Even more striking, a title search for “pneumophila” identified approximately 3,000 articles, whereas “micdadei” returned only 101. With the rapidly expanding opportunities to integrate epidemiological studies, genomics, and laboratory-based tests of virulence and persistence, the field is poised to advance knowledge of how, when, and where these common aquatic and soil microbes threaten human health.
L. pneumophila Life Cycle
As an environmental microbe, L. pneumophila adapts to fluctuating conditions by altering its physiology. Multiple distinct cell types have been defined based on a combination of morphological, biochemical, genetic, and molecular features (see Box 2-1; Robertson et al.,
2014). Depending on its environment, L. pneumophila can differentiate between replicative, transmissive, filamentous, mature infectious forms, and viable-but-not-culturable-like (VBNC-like) cells (see Figure 2-5). L. pneumophila cells obtained from solid media are a mix of replicating, transmissive, and filamentous cell types, whereas more homogenous cell populations can be isolated from broth cultures. Each specialized cell type differs in its capacity to infect host cells and tolerate antibiotics, biocides, and other environmental stresses, as discussed below.
Replicative and transmissive forms are two cell types observed when culturing L. pneumophila in rich bacteriological media at 37°C with aeration. Replicative cells are isolated during the exponential growth phase, and the transmissive cell type is generated in the stationary phase (Brüggemann et al., 2006; Byrne and Swanson, 1998). To alternate between replicative and transmissive states, L. pneumophila reprograms its gene expression and metabolic profile (Brüggemann et al., 2006; Dalebroux et al., 2009; Faucher et al., 2011, reviewed by Oliva et al., 2018). For example, when nutrients are plentiful in broth, amoebae, or macrophages, L. pneumophila rely on the regulatory protein CsrA to repress production of not only flagella but also virulence factors that promote transmission between host cells, while activating pathways that catabolize serine and glucose and generate energy storage granules, or “inclusions” of polyhydroxybutyrate (PHB; Eylert et al., 2010; Gillmaier et al., 2016; Häuslein et al., 2016, 2017; James et al., 1999; Sahr et al., 2017). Once nutrients become limiting, CsrA repression is relieved, and L. pneumophila begin to catabolize PHB, utilize glycerol as a precursor for biosynthesis, modify their LPS and surface composition, acquire stress resistance, assemble a flagella, produce numerous virulence factors for export by a Type IV secretion system, and become competent to evade phagosome-lysosome fusion (Eylert et al., 2010; Fernandez-Moreira et al., 2006; Harada et al., 2010; Häuslein et al.,
2016, 2017; Mendis et al., 2018; Nevo et al., 2014; Sahr et al., 2017; Trigui et al., 2015). Thus, replicative phase cells specialize in generating progeny, whereas transmissive cells are primed to escape one host cell, survive and disperse in the environment, and establish a protected replication niche in a new host cell.
In water and in lungs, a filamentous cell type of L. pneumophila has been observed (reviewed by Robertson et al., 2014). In general, filamentation is observed after exposure to environmental stress, including scarce nutrients, high temperatures, ultraviolet (UV) light, or antibiotics. Relatively few studies have focused on filamentous forms, and the regulatory controls that govern filamentation are not known. Nevertheless, some fitness advantages have been described. When L. pneumophila are cultured without aeration in rich media at 37°C, their elongated morphology likely enhances attachment to biofilm, as thick meshworks of multinucleate, filamentous cells readily adhere to glass (Piao et al., 2006). When co-cultured with epithelial cells and macrophages, filamentous L. pneumophila are internalized within phagolysosomal compartments that fail to seal; thus, these leaky host vacuoles are less toxic to the elongated prey (Prashar et al., 2012, 2013). Filamentous cells can revert to the typical short rod morphology once static cultures are aerated (Piao et al., 2006) or after ingestion by macrophages (Prashar et al., 2013). Additional research is needed to illuminate the molecular mechanisms that govern filamentation and when and where this multinucleate elongated cell type increases L. pneumophila fitness.
In amoebae and ciliates, intracellular L. pneumophila can further differentiate into hardy mature infectious forms (MIF; Faulkner and Garduño, 2002; reviewed by Robertson et al., 2014). Such MIF cells are characterized by thick cell walls, abundant PHB storage granules, and metabolic dormancy. Compared with cells obtained from either stationary phase broth or macrophage cultures, MIF L. pneumophila are more resistant, by orders of magnitude, to basic pH, detergents, chlorine, and antibiotics, and they adhere to macrophages more readily (Abdelhady and Garduño, 2013). MIFs can also be released from amoebae and ciliates within membrane-bounded vesicles, which provide an additional layer of protection. Thus, MIF cells appear to be well equipped for transmission within aerosols to the human lung (Robertson et al., 2014).
Once either transmissive or MIF L. pneumophila encounter adequate nutrients within host amoebae or macrophages, they return to a replicative form that generates numerous progeny. For example, transmissive L. pneumophila will persist as single cells within macrophages or amoebae vacuoles until adequate threonine is acquired; subsequently, the bacteria switch to the replicative form (Sauer et al., 2005). Within amoebae, robust L. pneumophila replication ensues when arginine relieves transcriptional repression by the arginine repressor, ArgR (Hovel-Miner et al., 2009, 2010). Transporter proteins embedded in the host vacuolar and bacterial membranes equip replicative L. pneumophila to obtain amino acids needed for bacterial replication (Schunder et al., 2014).
After prolonged environmental stress, L. pneumophila is thought to differentiate into a VBNC-like cell type. Described for many bacteria, such a VBNC adaptation increases bacterial resistance but also impedes environmental surveillance strategies that rely on culture (reviewed by Li et al., 2014). Vibrio vulnificus is a marine Gram-negative pathogen whose VBNC state has been analyzed in molecular detail. After four days exposure to 5°C artificial seawater, culturability of V. vulnificus declines 8 logs. Once the cell suspension is returned to 22°C, within less than 10 hours culturability rapidly increases, by greater than 6 logs (Whitesides and Oliver, 1997). This efficient revival is coordinated by a quorum sensing signal transduction system (Ayrapetyan et al., 2014). Thus, the aquatic pathogen V. vulnificus is genetically programmed to respond to shifts in temperature and cell density by efficiently alternating between replication-competent and VBNC states.
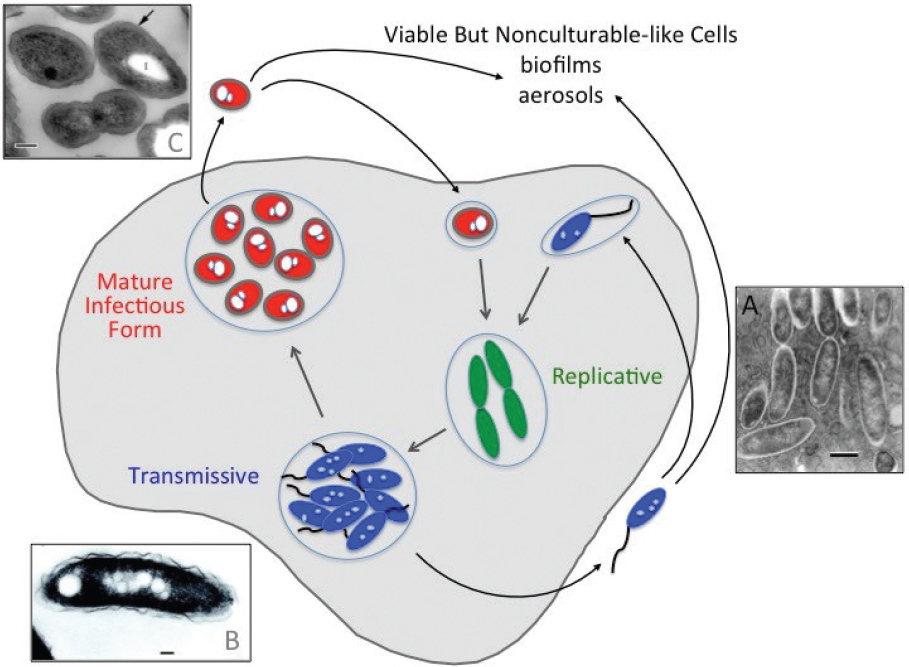
SOURCES: Faulkner et al. (2008); Faulkner and Garduño (2002); Garduño et al. (2002).
Compared to V. vulnificus, the capacity of Legionella spp. to alternate between replication and a resilient VBNC-like state remains enigmatic. When MIF L. pneumophila are held for weeks in hot, nutrient-poor water, the majority acquire features of VBNC cells (Al-Bana et al., 2014). For example, 30 days after exposure to 45°C double-deionized water, the capacity of MIFs to form colonies on rich media steadily declines by more than 6 logs; yet, greater than 80 percent of the cells have an intact cell membrane, as judged by cell membrane integrity stains. These MIF-derived, VBNC-like cells are resistant to detergent and remain intact when co-cultured with the ciliate hosts Tetrahymena tropicalis and T. thermophila, as judged by qualitative electron microscopy studies. However, MIF-derived, VBNC-like bacteria do not readily resume replication when cultured on rich medium or when ingested by a range of host phagocytes known to support robust L. pneumophila replication, including Acanthamoebae castellanii, T. tropicalis, T. thermophila, and human monocytic mouse fibroblast cell lines (Al-Bana et al., 2014). Therefore, whether MIF-derived, VBNC-like L. pneumophila pose a risk to public health remains an open question.
Compared to MIF cells, transmissive L. pneumophila exhibit a different profile after prolonged exposure to 45°C tap water (Al-Bana et al., 2014). After 45 days, culturability declines more than 8 logs, but only about 20 percent of the population maintains viability, as judged by either vital staining or qualitative ultrastructure analysis. Although these VBNC-like cells remain intact after treatment with detergent or ingestion by A. castellani, only about 1 in 105 of this population resumes replication in this permissive amoebae host.
Several investigations have reported that nutrient, chemical, or temperature stress triggers L. pneumophila to differentiate into a VBNC-like cell type (Alleron et al., 2008; Dietersdorfer et al., 2018; Ducret et al., 2014; García et al., 2007; Hwang et al., 2006; Kirschner, 2016; Schrammel et al., 2018). In general, after prolonged stress, stationary phase L. pneumophila lose culturability, yet bind a vital stain or retain intact ribosomes (Dietersdorfer et al., 2018; Epalle et al., 2015; Ohno et al., 2003; Steinert et al., 1997). Numerous experimental factors alter generation and resuscitation of these VBNC-like L. pneumophila, including water composition, bacterial strain, growth phase, cell density, and host cell type. Quantification of VBNC-like populations is also affected by the method used to prepare control dead cells, the medium in which cells are suspended, the density of the cell suspension, and the sensitivity of instruments used to detect fluorescence markers that distinguish between live and dead cells (Braun et al., 2019). In contrast to our mechanistic understanding of VBNC V. vulnificus and several other bacteria, neither the environmental conditions nor the regulatory pathways that stimulate L. pneumophila to alternate efficiently between VBNC-like and replicative or MIF cell types have been delineated.
A major outstanding question is whether VBNC-like L. pneumophila cause infections in humans. In early studies that utilized a guinea pig model to assess virulence of VBNC-like L. pneumophila, no viable bacteria were recovered after infection (Steinert et al., 1997). When co-cultured with amoebae, some VBNC-like cells appear to resume replication; however, the quantitative data provided make it difficult to rule out that a minor population of culturable cells survived the initial stress treatment and subsequently initiated the amoebae infection. In a recent comprehensive study, VBNC-like L. pneumophila obtained 221 days after starvation were 20- to 100-fold less infectious for human monocyte-derived macrophages than bacteria obtained from broth cultures (Dietersdorfer et al., 2018). Thus, more research is required to determine whether, or under what conditions, VBNC-like L. pneumophila can establish infections in humans.
A limitation of current laboratory research practices and disinfection studies is the failure to take into account the distinct cell types of L. pneumophila. Environmental surveillance largely relies on culture-based detection (ISO, 1998), which does not detect VBNC-like cells. Genomic DNA tests applied in the clinic or in the field cannot distinguish among dead, live, VBNC, or infectious bacteria. Sensitivity to detergents, biocides, antibiotics, and other stressors differs dramatically for replicative, transmissive, and MIF cells (Abdelhady and Garduño, 2013; reviewed by Robertson et al., 2014). The capacity of specialized L. pneumophila cell types to survive in aerosols and consequently gain access to the human lung has not been analyzed. The biochemical and environmental conditions that trigger development or resuscitation of VBNC-like L. pneumophila are not yet understood. Therefore, development and application of a standardized set of molecular markers specific to each cell type would advance the L. pneumophila field and ultimately guide clinical treatment and environmental remediation practices. Markers are currently available for replicative and transmissive forms of Legionella (Sauer et al., 2005) and for MIF (Abdelhady and Garduño, 2013), but not for VBNC-like Legionella or Legionella residing in biofilms. In the meantime, it is imperative that clinical, epidemiological, and research investigators be cognizant of the L. pneumophila life cycle and move toward newer approaches that can identify the cell type(s) in their samples.
MICROBIAL ECOLOGY OF LEGIONELLA
The microbial ecology of legionellae has been poorly studied since L. pneumophila was first cultivated from the environment. Although the genus Legionella contains 61 species and DNA has been identified for several addditional Legionella spp. that have not yet been cultivated (Edagawa et al., 2008; Parthuisot et al., 2010; Wery et al., 2008; Wullings and van der Kooij, 2006), what is known about legionellae ecology is almost exclusively based on studies with L. pneumophila. The original isolation and identification of L. pneumophila from water environments and environmental aerosols led to the biased consideration of this pathogen as a planktonic aquatic bacterium. While it is certainly present in water environments, current understanding points to its growth along with other pathogenic legionellae within various free-living protozoan hosts that feed on bacteria associated with biofilms (i.e., surface-attached microbes and their extracellular matrix) (Hilbi et al., 2011). Indeed, several Legionella spp. are thought to have developed various virulence factors as defense against predation by free-living protozoa (i.e., protozoa that grow in the environment on natural organic matter and microbes, not obligate parasites that only grow within another living organism) (Cianciotto, 2015), as have various other genera of so-called amoeba-resisting bacterial pathogens (Kebbi-Beghdaji and Greub, 2014). This section discusses principles of Legionella ecology as well as the two growth habitats for pathogenic legionellae—its primary reservoir in nature and its secondary habitat in engineered environments that may generate infectious doses delivered via aerosols to humans.
Principles of Legionella Ecology
Biofilms consist of microorganisms that grow attached to moist soil, sediment, decaying organic matter, or other solid surfaces. They are largely hydrated gels composed of extracellular polymeric substances, consisting of carbohydrates, fats, protein, and nucleic acids excreted by bacteria (Flemming et al., 2016). Gradients of nutrients, pH, and oxygen within the biofilm matrix support the varying needs of different microorganisms in the heterogeneous biofilm community. Various protozoa, and later on microinvertebrates, will naturally develop and feed on biofilms, further influencing the microbial diversity of mature biofilms. In oligotrophic environments such as drinking water, a “mature” biofilm consisting of a relatively stable microbial community composition can take several years to develop (Martiny et al., 2003). In contrast, most published biofilm experiments are typically undertaken after a limited period of development (Storey et al., 2004a, 2008).
L. pneumophila may form a mono-species biofilm in the laboratory and grow necrophilically on other decaying cells (Temmerman et al., 2006). Moreover, free, inactive Legionella persist within biofilms (Hindré et al., 2008). However, the primary growth habitat of L. pneumophila is within amoebae (Kuiper et al., 2004) or other free-living protozoa that are associated with biofilms (Buse et al., 2012; Hellinga et al., 2015). Within these protozoan hosts associated with fixed or free-floating biofilms (Hsu et al., 2011), pathogenic legionellae can replicate to problematic levels (Ashbolt, 2015; Declerck et al., 2010; Hamilton et al., 2019).
As illustrated in Figure 2-6 and Figure 2-7, large numbers of L. pneumophila can be released from the biofilm environment within fragments of biofilm, within protozoan trophozoites and cysts, or within expelled amoebal vesicles (membrane-bound structures containing undigested food and microorganims). These released L. pneumophila cells then disperse in the bulk water phase among other planktonic microorganisms. The dependency of L. pneumophila growth on protozoan hosts that graze on biofilms associates the ecology
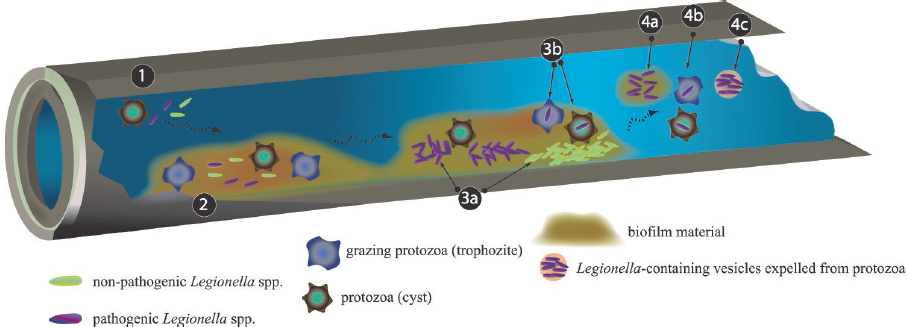
SOURCE: Lau and Ashbolt (2009).
of L. pneumophila indirectly to the ecology of its host protozoan and supporting biofilm microorganisms.
Role of Free-Living Protozoa in Growth of Legionellae
Rowbotham (1980) first showed that L. pneumophila multiply in different protozoan hosts. Moreover, L. pneumophila multipled in sterile water when amoebae were added, but not when amoebae were removed by filtration (Kuiper et al., 2004; Nahapetian et al., 1991; Wadowsky et al., 1988). Subsequently, many free-living protozoa (amoebae and ciliates) have been identified that can serve as hosts for growth of Legionella (see Table 2-1). It is likely that other higher animal organisms can also serve as growth and/or transport hosts for L. pneumophila (Hellinga et al., 2015). Of note, L. pneumophila cells will not grow within the dry, dormant cysts of protozoa, although the bacteria are preserved and show increased resistance to disinfection proceses such as heat, chemicals, sonication, and UV (Cervero-Arago et al., 2014; Declerck et al., 2010; Storey et al., 2004b).
Not all protozoa will serve as hosts under all conditions or for all strains and species of Legionella (e.g., Wadowsky et al., 1991), perhaps due to novel non-coding RNA expressed during predator–prey interactions (Weissenmayer et al., 2011). For example, some protozoa may be resistant to infection with L. pneumophila because they digest L. pneumophila (Amaro et al., 2015); these eukaryotes could potentially serve as biological control agents (Maita et al., 2018, as suggested by Wang et al., 2013). Some protozoa have a preference for other bacterial prey (Shaheen et al., 2019). Finally, some species of amoeba contain symbionts that do not allow Legionella to replicate within the host (Maita et al., 2018; Okubo et al., 2018).
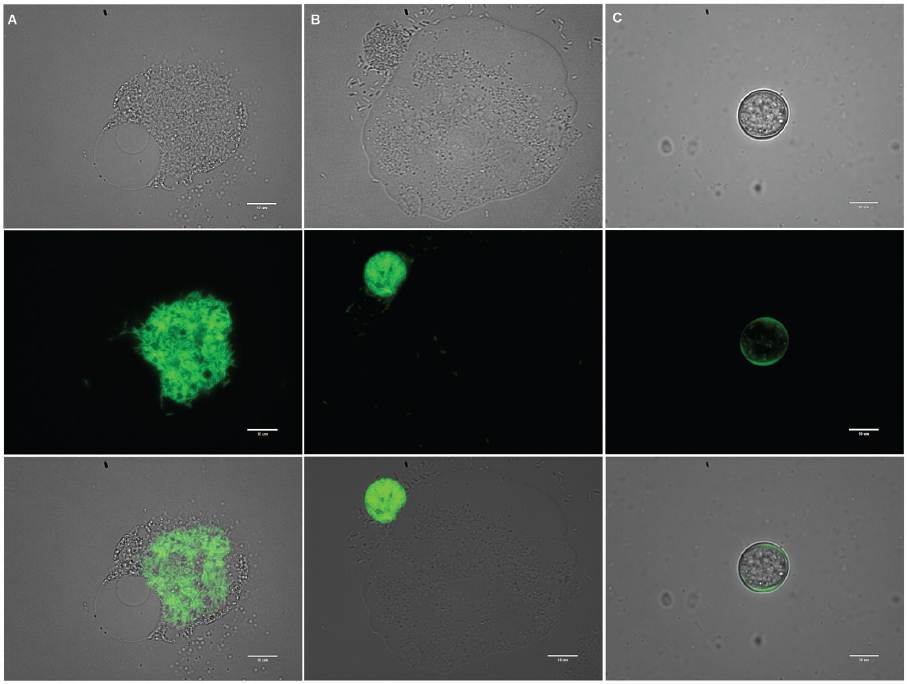
SOURCE: Courtesy of Mohamed Shaheen.
The protozoan hosts for L. pneumophila graze on microorganisms present in the biofilm or sediments. By phagocytosis, host cells engulf and internalized prey microbes within cell membranes known as phagosomes (Abu Kwaik et al., 1998). Normally, the phagocytosed prey is delivered to lysosomes where it is digested by the acidic pH and lysosomal enzymes. Nutrients liberated through this process fuel the protozoan cell. However, many bacterial species, including pathogenic Legionella spp., have evolved strategies to escape digestion by the protozoan cell and persist in vacuoles within the protozoan host (Vandenesch et al., 1990). Within these vacuoles, these bacteria may multiply, especially at temperatures greater than 30°C (Buse et al., 2017; Caicido et al., 2018). Bacterial proliferation eventually kills or lyses the protozoan cell, releasing the intracellular progeny into the aquatic or soil environment (Kuiper et al., 2004). This strategy likely enables fastidious bacteria such as L. pneumophila to persist and compete with other bacteria in otherwise low-nutrient (oligotrophic) and inhospitable environments (King et al., 1988). As a consequence of adaptation to resist amoeba digestion, various bacteria either grow pathogenically, become benign, or
TABLE 2-1 Known Hosts of Legionella
| Organism | Eukaryote | Reference |
|---|---|---|
| Acanthamoeba polyphaga | Amoebae | Rowbotham, 1980 |
| Acanthamoeba castellanii | Amoebae | Rowbotham, 1980 |
| Naegleria gruberi | Amoebae | Rowbotham, 1980 |
| Naegleria jadini | Amoebae | Rowbotham, 1980 |
| Naegleria lovaniensis | Amoebae | Tyndall and Domingue, 1982 |
| Acanthamoeba royreba | Amoebae | Tyndall and Domingue, 1982 |
| Acanthamoeba palestinensis | Amoebae | Anand et al., 1983 |
| Tetrahymena pyriformis | Ciliate | Fields et al., 1984 |
| Naegleria fowleri | Amoebae | Newsome et al., 1985 |
| Vahlkampfia jugosa | Amoebae | Rowbotham, 1986 |
| Echinamoeba exudans | Amoebae | Fields et al., 1989 |
| Hartmanella cantabrigiensis | Amoebae | Fields et al., 1989 |
| Vermamoeba [Hartmanella] vermiformis | Amoebae | Fields et al., 1989 |
| Hartmanella sp. | Amoebae | Fields et al., 1989 |
| Acanthamoeba culbertsoni | Amoebae | Fields et al., 1989 |
| Tetrahymena thermophile | Ciliate | Kikuhara et al., 1994 |
| Dictyostelium disocideum | Amoebae | Hägele et al., 2000 |
| Balamuthia mandrillaris | Amoebae | Shadrach et al., 2005 |
| Caenorhabditis elegans | Nematode | Rasch et al., 2016 |
| Willaertia magna | Amoebae | Dey et al., 2009 |
| Diphylleia rotans1 | Flagellate | Valster et al., 2010 |
| Echinamoeba thermarum1 | Amoebae | Valster et al., 2010 |
| Neoparamoeba spp.1 | Amoebae | Valster et al., 2010 |
| Acanthamoeba griffini, Acanthamoeba jacobsi, Naegleria australiensis, Naegleria philippinensis, Naegleria italica | Amoebae | Hsu et al., 2011 |
| Stylonychia bifaria1 | Ciliate | Rasch et al., 2016 |
| Stylonychia mytilus1 | Ciliate | Rasch et al., 2016 |
| Ciliophrya spp.1 | Ciliate | Rasch et al., 2016 |
1 Candidate host; in vitro studies are needed to confirm their role as environmental host.
form a stable symbiotic relationship with protozoa (Schmitz-Esser et al., 2010; Shu et al., 2018) as discussed further below.
Effects of the Protozoan Host on Virulence and Stress
Over millennia, free-living protozoa have contributed to the development of a repertoire of virulence and stress-response genes in Legionella and other amoeba-resisting bacteria (Guimaraes et al., 2016; Koubar et al., 2011; Trigui et al., 2015). Considerable horizontal
gene transfer between amoeba and their internalized bacteria (Guimaraes et al., 2016) also contributes to ongoing changes in traits within legionellae. The host and the conditions for culturing legionellae may also affect its infectivity, such as if cells are VBNC-like, in the replicative or the transmissive stage of Legionella’s life cycle (Fonseca and Swanson, 2014), or as short rods versus filamentous morphologies (Garduño et al., 2002; Vandenesch et al., 1990). Adaptation of L. pneumophila’s lipid A cell surface is one of a series of factors that affects its ability to infect host amoebae (Albers et al., 2007). As discussed previously, not describing and controlling the cell form(s) used in infection studies could well be contributing to the differing views reported in the literature.
The increase in infectivity following L. pneumophila growth in amoebae may depend on the protozoan host and the animal model used, although the mechanism of this increased virulence is unknown. While no difference was seen in the infectivity of guinea pigs via aerosols of L. pneumophila cells grown in co-culture with an Acanthamoeba spp. compared to pure culture cells from agar plates (Vandenesch et al., 1990), Acanthamoeba castellanii- and Vermamoeba vermiformis-associated L. pneumophila were described as more pathogenic in macrophages and in a mouse model than an equal number of non-amoeba-associated L. pneumophila (Brieland et al., 1997; Cirillo et al., 1994). For example, L. pneumophila grown in Acanthamoeba are some 100-fold more infectious in epithelial cells and ten times more infectious in macrophages or other cell lines than agar-grown L. pneumophila (Cirillo et al., 1994; Garduño et al., 2002). Importantly, after intracellular replication in free-living protozoa, higher resistance has been documented for L. pneumophila stressed by heat, oxidants, acids, osmotic shock (Kwaik et al., 1997), biocides (Barker et al., 1992; Berk et al., 1998), and antibiotics (Barker et al., 1995; Garduño et al., 2002). Other cellular differences between co-cultured L. pneumophila and L. pneumophila grown alone on agar include differences in cellular fatty acid composition (Barker et al., 1993; Vandenesch et al., 1990). However, a limitation of each of these comparative studies is that it is unclear whether the agar-grown cell samples contained exponential-phase “replicative” bacteria that are readily degraded in lysosomes, post-exponential-phase “transmissive” bacteria that evade lysosomes, or a mixture of the two.
Role of Temperature
Legionellae have been observed in environments ranging from 0°C to 45°C, indicating that legionellae are psychrophilic to mesophilic (Wullings and van der Kooij, 2006). However, most of the described species of the genus Legionella are mesophilic and grow between 25°C and 43°C under laboratory conditions (Garrity et al., 2005). Under environmental conditions, growth of L. pneumophila has been observed between 25°C and 45°C (Buse et al., 2017; Tison et al., 1980; van der Kooij et al., 2016; Wadowsky et al., 1985; Yee and Wadowsky, 1982). Since the growth of L. pneumophila in these environments largely depends on amoebae, studies have also focused on L. pneumophila growth in protozoan hosts at different temperatures. L. pneumophila proliferate in Acanthamoeba palestinensis and A. castellanni at 25°C and 35°C, respectively, but are digested by the amoeba at 15°C and 20°C (Anand et al., 1983; Ohno et al., 2008), which is consistent with generally low levels of Legionella observed in natural environments at these lower temperatures.
The optimal temperature range for L. pneumophila to express several factors that promote infection and transmission is between 25°C and 30°C, not 37°C or higher. These factors include flagellar-based motility (Ott et al., 1991); PilD, a critical component of Type II secretion and Type IV pili, two machines that equip L. pneumophila to move across
surfaces by sliding motility (Stewart et al., 2009); and LvhB2, a virulence factor exported by Type II secretion that enhances L. pneumophila infection of host cells at 25°C, but not 37°C (Ridenour et al., 2003). A functional Type II secretion system also increases L. pneumophila survival in water at temperatures of 17°C and below and bacterial replication in amoebae at 22°C to 25°C (Söderberg et al., 2004).
Studies differ in the maximum temperature observed for L. pneumophila growth. Some researchers observed growth up to 45°C (Kusnetsov et al., 1996; Tison et al., 1980), whereas others did not observe growth at 42°C or higher (Ohno et al., 2003; van der Kooij et al., 2016). Van der Kooij and colleagues (2016) observed that L. pneumophila serogroup 1 strains of different sequence types had different optimum and maximum growth temperatures when grown under drinking water conditions in a biofilm monitor. In addition, using the same biofilm monitor, L. pneumophila was not capable of growth in the naturally formed biofilm with temperatures greater than 41°C. A 2-log lower V. vermiformis host count at 42°C compared to 38°C indicated that the absence of a thermotolerant host at 42°C prevents proliferation of L. pneumophila in this system (van der Kooij et al., 2016). Indeed, different L. pneumophila strains have different optimal temperatures when grown in protozoan hosts (Buse and Ashbolt, 2011), and the protozoan community composition also varies with temperature in drinking water systems (Valster, 2011). Thus, both the strain of L. pneumophila and host protozoan diversity affect the temperature range for growth of L. pneumophila.
At temperatures greater than 50°C, the number of cultivable L. pneumophila declines. The time required to reduce the concentration of viable bacteria by 90 percent is referred to as the decimal reduction time. Decimal reduction times between 100 and 1,000 minutes were observed for Legionella at 50°C (van der Kooij, 2014). The lowest reduction time (100 minutes) was observed for L. pneumophila precultured with natural microbiota. Moreover, these decimal reduction times decreased to around two minutes at 60°C for L. pneumophila pure cultures grown under natural conditions (Dennis et al., 1984; Schulze-Robbecke et al., 1987; van der Kooij, 2014). No cultivable L. pneumophila were observed after heat shock treatment for 10 minutes at 70°C with pure cultures of different L. pneumophila strains (Allegra et al., 2008). In contrast, metrics of viability (membrane-intact cells and adenosine triphosphate or ATP present) could still be measured for most strains, and, when the suspension was subsequently incubated with a protozoan host, L. pneumophila was cultivated. Membrane integrity can be a poor indicator for viability in L. pneumophila (Hammes et al., 2011, Wullings et al., 2016), and free ATP can still be present after cell death (Nescerecka et al., 2016). Nevertheless, the capacity to culture bacteria from a protozoan host demonstrates that at least some L. pneumophila cells remained viable after heat-shock treatment for 10 minutes. Unfortunately, a decimal reduction time was not calculated. An earlier study of several L. pneumophila strains showed that the decimal reduction time at 70°C varied between 1.1 and 2.6 minutes (Stout et al., 1986), meaning that 10 minutes of exposure to 70°C would be expected to result in a 3.8- to 9.1-log reduction. Therefore, the Allegra et al. (2008) report that L. pneumophila cells survived 10 minutes of treatment at 70°C is consistent with previous observations.
Inactivation of pure cultures of ten different Legionella species, eight different L. pneumophila serogroups, and one to five different L. pneumophila strains showed that the decimal reduction times at 60°C ranged from 2 to 5 minutes (Stout et al., 1986). This difference in decimal reduction time was observed between species, serogroups, and strains, but was only determined by loss of culturability, not by other measures of viability loss (hence, some cells may be VBNC-like). In addition, Allegra and colleagues (2008) also observed that the different L. pneumophila strains showed different reduction curves for the membrane-intact cells when treated at 70°C (see Figure 2-8). Thus, the reduction times after temperature dis-
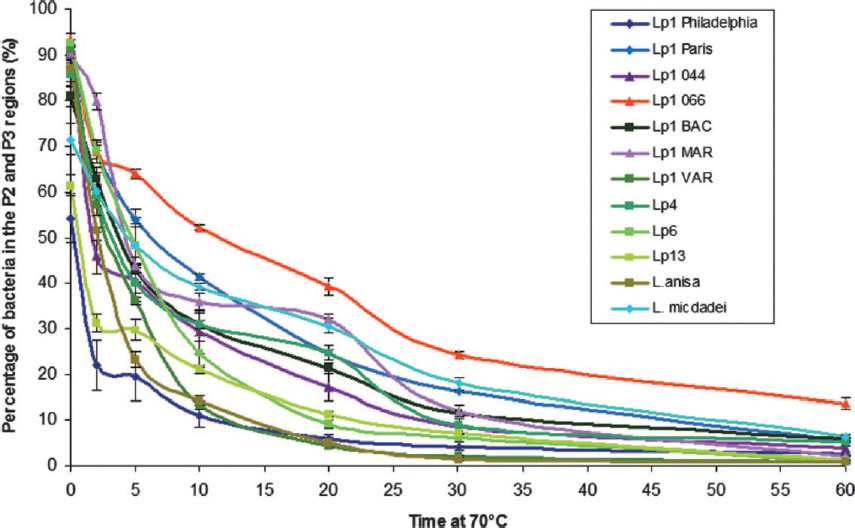
SOURCE: Allegra et al. (2008).
infection can be strain-, serogroup-, and species-dependent. Interestingly, Allegra et al. (2011) showed that repeated thermal shocks actually selected for heat-resistant L. pneumophila strains.
Most recently, Cervero-Aragó and colleagues (2019) investigated two L. pneumophila strains for more than 80 days using a combination of cell-based viability indicators with cells incubated at 55°C, 60°C, and 70°C. Culturability was lost after 3 to 8 hours, 60 minutes, and less than 2 minutes, respectively; whereas, based on viability indicators, a 4-log reduction was achieved only after 150, 8 to 15, and 1 to 4 days, respectively. To investigate cells in a VBNC-like state, Cervero-Aragó et al. (2019) evaluated the infectivity of these heat-shocked L. pneumophila in amoebae and a lung macrophage cell line (THP-1). Infectivity lasted for at least 85 days at 55°C and 60°C and for up to 8 days at 70°C, albeit with reduced efficiency. Cervero-Aragó et al. (2019) concluded that a prolonged thermal regime at or above 60°C at the central parts of warm-water building systems is effective against not only culturable L. pneumophila but also VBNC-like cells.
Since L. pneumophila seems to multiply mainly in protozoan hosts, the effect of high temperatures on possible protozoan hosts for L. pneumophila might also contribute to eradication of L. pneumophila. Most protozoan host species of the genus Acanthamoeba cannot multiply at temperatures above 37°C (de Jonckheere, 1980). However, another protozoan host for L. pneumophila (such as V. vermiformis) continues to multiply above 50°C (Kuchta et al., 1993; Rohr et al., 1998). Cervero-Arago et al. (2013) showed that trophozoites of two Acanthamoeba strains and two V. vermiformis strains, isolated from
drinking water, had a decimal reduction time of 7.2 to 11 minutes at 50°C and less than 0.34 to 1.4 minutes at 60°C. In contrast, the decimal reduction time of cysts of these protozoan species varied between 30 and 76 minutes at 50°C and between 4.7 and 10.7 minutes at 60°C, with Acanthamoeba strains being the most thermotolerant. These reduction times at 60°C are considerably longer than for L. pneumophila, implying that decimal reduction times for L. pneumophila inside cysts are likely to be longer than for planktonic L. pneumophila cells. Indeed, Storey et al. (2004b) showed persistence of L. pneumophila within cysts after 10 minutes at 80°C.
Role of Nutrients
Growth of legionellae in defined media demonstrate that these bacteria require specific nutrients, including certain amino acids and ferric iron (George et al., 1980; Ristroph et al., 1981). Indeed, legionellae are readily recovered in relatively nutrient-rich reclaimed wastewaters (as reviewed by Caicedo et al., 2019). Conversely, free cells of L. pneumophila cannot compete successfully with oligotrophic bacteria in nutrient-poor aquatic environments such as drinking water (van der Kooij, 2014). As described above, L. pneumophila circumvents this problem by multiplying in protozoan hosts, which provide its required nutrients. Since these protozoan hosts graze on bacteria, the concentration of protozoa depends on the concentration of prey bacteria that are present. A clear correlation between biofilm concentration and numbers of cultivable L. pneumophila has been observed in a biofilm monitor fed with unchlorinated drinking water types that differ in assimilable organic carbon (AOC) concentration and inoculated with L. pneumophila (Learbuch et al., 2019; van der Kooij et al., 2017). The biofilm concentration also correlates with the AOC concentration in the drinking water (van der Kooij et al., 2017). This implies that the Legionella numbers in drinking water are indirectly correlated to the AOC concentration of the water. Based on these data, the authors deduced for drinking waters sourced via riverbank infiltration a threshold biofilm concentration of 50 µg ATP/cm2 and a threshold AOC concentration of 1 µg C/L to initiate growth of L. pneumophila in the product drinking water.
Notably, an AOC concentration of less than 1 µg C/L in drinking water is extremely low and cannot be achieved by drinking water plants in the United States (Volk and LeChevallier, 2000). Even in The Netherlands, where microbial growth in drinking water systems is limited by controlling AOC concentrations instead of using a disinfectant residual, the majority of treatment plants produce drinking water with AOC concentrations above 1 µg C/L (Paul van der Wielen, personal communication). Others have not observed a relationship between organic carbon and L. pneumophila numbers in drinking water supplemented with different fulvic acid concentrations (Williams et al., 2015), indicating that other environmental factors (e.g., host protozoan concentration) contribute to L. pneumophila growth as well.
A relationship between several minerals (e.g., iron, zinc, calcium, manganese, magnesium) and Legionella numbers in premise plumbing systems has been observed (van der Kooij, 2014, and references therein), although findings are not consistent among studies. For instance, Vickers and colleagues (1987) observed that calcium and magnesium, but not iron and zinc, correlated with L. pneumophila in hospital water systems, whereas Bargellini and colleagues (2011) observed that iron and zinc correlated with L. pneumophila in hot-water systems of hotels and private homes. The role of ferric iron in growth of L. pneumophila has been studied intensively; it is a prerequisite for growth of L. pneumophila in chemically defined media (Reeves et al., 1981; States et al., 1985; Warren and Miller, 1979). The addition of the iron chelator lactoferrin reduces viability of L. pneumophila (Goldoni et al.,
2000). In addition, ferric/ferrous ions also promote L. pneumophila virulence (Allard et al., 2009). Iron overload has also been associated with increasing Legionella infectivity rates of amoebae and macrophages as indicators of increased virulence (Buracco et al., 2018; James et al., 1999). On the other hand, excessive iron inhibits Legionella biofilm formation (Hindré et al., 2008). The effect of other mineral elements on the growth of L. pneumophila in premise plumbing is still not well understood (Borella et al., 2005).
Low nutrient availability in artificial media inhibits L. pneumophila growth, generating VBNC-like cells. As discussed previously, whether Legionella forms VBNC cells is controversial. VBNC-like L. pneumophila cells persisting in drinking water for more than 100 days that can be resuscitated by amoeba co-culture have been described (Steinert et al., 1997). It remains difficult, however, to make definitive conclusions because the lack of growth in the absence of a protozoan host was not proven (only that L. pneumophila was below a certain detection limit). Still, others have made similar observations in which resuscitation through a protozoan host recovered the culturability of L. pneumophila on agar media (Alleron et al., 2008; Ducret et al., 2014; Garcia et al., 2007; Hwang et al., 2006). Furthermore, it was observed that detection of L. pneumophila from environmental samples is improved when a co-culture method with a protozoan host is used (Garcia et al., 2007; La Scola et al., 2001; Sanden et al., 1992; Schalk et al., 2012). This is strong evidence that not all viable L. pneumophila strains in the environment will be cultivated when using artificial media.
Legionella in the Natural Environment
Aquatic Environment
Soon after L. pneumophila was first isolated and described, the organism was observed at low numbers (approximately 1 percent of total bacterial community) in water from different lakes and rivers in the United States using DFA techniques and isolated by infecting guinea pigs (Fliermans et al., 1979, 1981). The frequency of L. pneumophila isolation correlated with water temperature, and more strains could be isolated from surface waters in summer (Fliermans et al., 1979). Around the world, legionellae were identified among the natural microbiota in surface waters (e.g., Carvalho et al., 2007, 2008; Ortiz-Roque and Hazen, 1987; Parthuisot et al., 2010) or groundwater (Costa et al., 2005; Riffard et al., 2001). These studies show in general that legionellae numbers are low (less than 103 CFU/L). Detected only sporadically, L. pneumophila is not among the dominant Legionella species in surface waters and groundwater that have water temperatures below 25°C. In contrast, L. pneumophila predominated among the legionellae isolated from natural hydrothermal vents and hot springs where water temperatures were above 27°C, reaching numbers up to 106 CFU/L (Marrão et al., 1993; Verissimo et al., 1991). In an acidic hot spring with a temperature of 30°C to 47°C and a pH of 2.7, 16S rRNA gene sequences of L. pneumophila were not detected, indicating that the low pH selected for other Legionella species (Sheehan et al., 2005). L. pneumophila has been identified in natural water from thermal springs used in spas, a suspected source for Legionnaires’ disease (van Heijnsbergen et al., 2015). It should be noted that the protozoan hosts for L. pneumophila also reside in natural aquatic environments (reviewed in Plutzer and Karanis, 2016).
Soil Environment
Legionellae also inhabit soil. For example, a large number of legionellae isolates (n = 114) belonging to 12 different Legionella species were obtained from soil samples
in Thailand (Travis et al., 2012). Moreover, L. longbeachae, another causative agent of Legionnaires’ disease, is mostly isolated from potting mixes and gardening soil (Whiley and Bentham, 2011 and references therein), although it resides in aquatic environments as well (Joly et al., 1984; Marrie et al., 1994). In contrast to L. pneumophila, L. longbeachae contains genes that encode 12 cellulolytic enzymes, which together may fully degrade cellulose. Accordingly, L. longbeachae is equipped to metabolically degrade plant material (Cazalet et al., 2010), supporting the notion of its natural habitat being associated with degrading plant matter.
Morris and colleagues (1979) isolated L. pneumophila from mud and sand sediments sampled from the bottom of a stream impacted by thermally polluted water. Furthermore, L. pneumophila was detected in these soil samples by fluorescent antibody staining, although the microbes could not be cultivated on agar media or after guinea pig inoculation. Others have detected L. pneumophila in soil, in one case throughout the year in the same garden soil (Travis et al., 2012; van Heijnsbergen et al., 2016). In Australia, the same unusual L. pneumophila strain type was isolated from a Legionnaires’ disease patient and from soil in the patient’s work area, implying a direct link between soil and disease (Wallis and Robinson, 2005). Likewise, natural soil has been identified as a possible source for various cases of Legionnaires’ disease (van Heijnsbergen et al., 2015). Although different Legionella species have been isolated from soil, and soil or dust contamination of engineered systems has been associated with outbreaks (van Heijnsbergen et al., 2016), the ecology of legionellae in soil has not been studied intensively. Free-living protozoa known to be hosts for L. pneumophila (e.g., V. vermiformis and Acanthamoeba spp.) are also common in soil samples (Denet et al., 2017; Tyml et al., 2016). Hence, L. pneumophila in the soil environment likely replicate within protozoan hosts, although proof for this phenomenon is still lacking.
Legionella in Engineered Environments
The growth of pathogenic legionellae to problematically high densities (Hamilton et al., 2019) seems to be favored in various engineered environments that support free-living protozoa associated with biofilms and that generate aerosols (Buse et al., 2012). The engineered environments that support Legionella growth include drinking water treatment plants, plumbing within buildings (i.e., premise plumbing), cooling towers, wastewater treatment plants that receive warm industrial effluents (Loenenbach et al., 2018), and a myriad of devices that operate with warm water including hot tubs. As shown in Figure 2-9, these compartments are a potential source of Legionnaires’ disease if contaminated aerosols are generated that can be inhaled or aspirated by people in their vicinity.
Drinking Water Distribution Systems and Premise Plumbing
Various species of legionellae have been detected in drinking water (Buse et al., 2012; Donohue et al., 2014; Hull et al., 2017). Legionella can enter drinking water systems by many possible routes, starting with source water that is contaminated by industrial, thermally elevated wastewater. For example, Legionella can grow within free-living protozoa associated with sand filters used in drinking water treatment and within amoebae that pass through drinking water treatment into distribution (Loret and Greub, 2010; Lu et al., 2016). Lu et al. (2015) provided evidence for atmospheric (soil/dust) ingress of Legionella through air vents into drinking water reservoirs and growth within reservoir sediments. Legionella may also enter drinking water distribution systems during construction (Francois Watkins
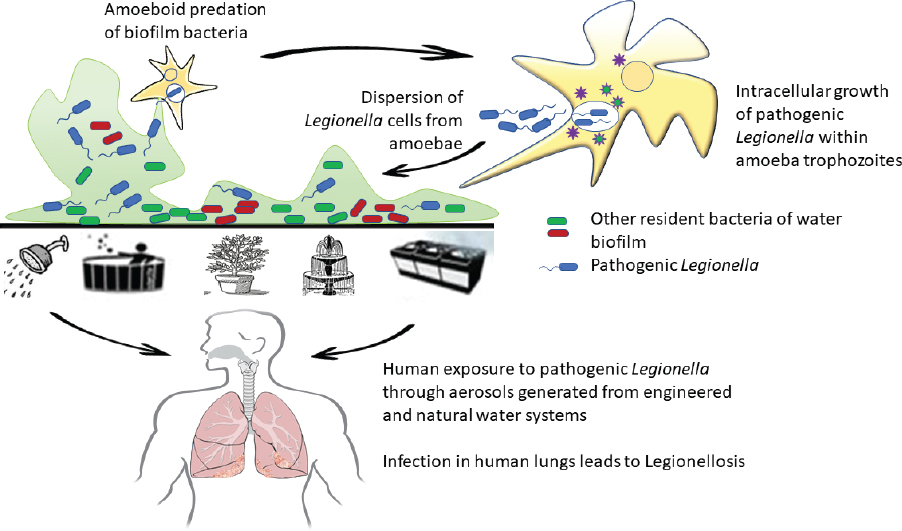
SOURCE: Courtesy of Mohamed Shaheen.
et al., 2016; Knox et al., 2016; hypothesized in Mermel et al., 1995; Stout et al., 2000) or main breaks (as suggested in Rhoads et al., 2017a).
Drinking water distribution systems containing free-living protozoa harboring pathogenic legionellae can also “seed” premise plumbing; subsequent growth in dead-ends and other stagnant regions may lead to sporadic cases and outbreaks of Legionnaires’ disease (Beer et al., 2015; Hamilton and Haas, 2016; Schoen and Ashbolt, 2011). In particular, the last few distal meters of piped water in premise plumbing are likely zones where problematic concentrations of L. pneumophila develop (i.e., 105 to 106 CFU/L; Hamilton et al., 2019; Schoen and Ashbolt, 2011). L. pneumophila has been isolated from hot/cold mixing valves, tap aerators, plastic shower hosing, and shower heads (Collins et al., 2017; Proctor et al., 2018); household “cold”-water storage tanks (Peter and Routledge, 2018); and stagnant hot-water lines within an optimal temperature window for intracellular growth (i.e., 28°C to 45°C) (Proctor et al., 2017). The many reasons for this proliferation are discussed below.
Stagnation. Stagnant areas in premise plumbing support more cultivable legionellae (including L. pneumophila) than other parts of premise plumbing (Fisher-Hoch et al., 1982; Tobin et al., 1981). Ciesielski et al. (1984) showed that hot-water tanks with stagnant water support higher L. pneumophila numbers (105 to 106 CFU/L) compared to hot-water tanks in which water was continuously replaced (less than 104 CFU/L). Compared to non-stagnant water, stagnant water has lower or no disinfectant residual (Fisher-Hoch et al., 1982; Wang et al., 2012), lower water temperatures (Patterson et al., 1994), higher concentrations of organics (LeChevallier et al., 1996; Wang et al., 2012), lower dissolved oxygen concentrations (Wang et al., 2012), higher biomass concentrations (Lautenschlager et al., 2010), altered microbial community composition (Dai et al., 2018a; Lautenschlager et al., 2010), and higher numbers of protozoan hosts (Wang et al., 2015b)—factors that all influence L. pneumophila growth.
Corrosion. The impact of corrosion products on Legionella proliferation is multifaceted (Brazeau and Edwards, 2013). By consuming residual disinfectant, these compounds create a more favourable environment for Legionella growth. Increased bioavailability of various metal corrosion products, such as iron, may also upregulate virulence in legionellae, stimulate general biofilm growth (Buse et al., 2012), and contribute to Legionella growth in hot-water heaters (Dai et al., 2018b; Ji et al., 2017; Proctor et al., 2017). Iron released during the recent massive corrosion event in Flint, Michigan, contributed to loss of chlorine residual, and, as a required nutrient (Reeves et al., 1981; States et al., 1985; Warren and Miller, 1979), this metal was also hypothesized to stimulate Legionella growth (Rhoads et al., 2017a). Van der Lugt et al. (2017) also recently reported that iron rust in stainless-steel shower heads resulted in increased Legionella anisa plate counts. Corrosion products can also promote heterotrophic biofilm growth by producing electron donors, such as hydrogen, and by stimulating autotrophic metabolism and fixation of organic carbon in the system (Rhoads et al., 2017b).
Pipe Materials. Pipe material may influence growth of L. pneumophila. For example, rubber material in a model pipe system enhanced growth of L. pneumophila, except when a biocide (thiuram) was present in the rubber material (Niedeveld et al., 1986). Plastic pipe materials can also enhance growth of L. pneumophila, especially those used in premise plumbing, such as soft PVC (PVC-P), polyethylene, polypropylene, or polybutylene materials (Rogers et al., 1994a,b; van der Kooij et al., 2002). The biofilm concentration on each type of pipe material correlates with L. pneumophila load (Learbuch et al., 2019). Therefore, pipe materials most likely affect L. pneumophila growth indirectly: higher biofilm concentrations support more protozoan hosts, which generate higher counts of L. pneumophila (van der Kooij et al., 2017). European standardized laboratory tests have demonstrated that, compared to an inert material such as glass, rubber (natural and synthetic), soft PVC (i.e., PVC-P), polyethylene, polypropylene, and polybutylene significantly enhance microbial growth (Hambsch et al., 2014) because of the growth-promoting organic compounds that these materials release. In contrast, stainless steel, PVC-C, and PVC-U did not enhance growth of L. pneumophila in these laboratory tests. A field study of several buildings demonstrated that the highest cultivable legionellae numbers were present in the biofilm on rubber components of taps (van Hoof et al., 2014), consistent with various laboratory test results.
In premise plumbing, the impact of copper pipes on legionellae is not consistent among studies, possibly due to differences in biofilm microbiota and the physiological status of cells. Several laboratory studies report that copper inhibits growth of L. pneumophila (e.g., Learbuch et al., 2019; Rogers et al., 1994b; Schoenen et al., 1988). In addition, Danish buildings with copper premise plumbing showed lower cultivable legionellae counts than buildings with iron pipes (Pringler et al., 2002). In contrast, others observed enhanced growth of L. pneumophila on copper compared to PVC-U or PVC-C (Buse et al., 2014a,b; Gião et al., 2015; van der Kooij et al., 2002). Likewise, by comparing bacteria growing in tubing downstream of biofilm reactors with copper versus PVC-U coupons, Lu et al. (2014) also noted that injected L. pneumophila actually survive better downstream of copper. A companion paper by the same group (Buse et al., 2014a) further indicated that the copper coupons were colonized by and released a greater number of L. pneumophila when co-inoculated with Acanthamoebae polyphaga and measured by qPCR, but L. pneumophila were only cultivable from PVC-U coupons.
There are several possible explanations for the apparent enigma of net effects of copper plumbing on Legionella. First, van der Kooij and colleagues (2005) observed that new unused copper material initially inhibited growth of L. pneumophila due to the release
of copper ions, but when the copper material was corroded, release of copper ions was reduced and inhibition of L. pneumophila no longer occurred. Interactions of the copper pipe with the local water chemistry is also important to consider. Proctor et al. (2017) noted that benefits of copper pipe for depressing L. pneumophila levels were only apparent at or below 41°C. Above 53°C, L. pneumophila were no longer detectable, and thus pipe material did not matter. Buse et al. (2017) noted that a higher pH (greater than 8.2), which limits dissolution, can also limit antimicrobial activity of copper pipe. Build-up of corrosion byproducts over time also limits the antimicrobial activity of copper toward Legionella (van der Kooij et al., 2005). In addition to having stronger antimicrobial properties than solid Cu, free Cu2+ in solution can induce other reactions, such as corrosion and associated hydrogen production, which could indirectly impact Legionella (Proctor et al., 2017; Rhoads et al., 2017b).
Copper might also induce a VBNC-like state for L. pneumophila, as has been suggested for Pseudomonas aeruginosa (Flemming et al., 2014). Induction of a VBNC-like state through copper exposure decreased the number of L. pneumophila detected by cultivation (Learbuch et al., 2019; Rogers et al., 1994a; Schoenen et al., 1988; van der Kooij et al., 2002), but not the number quantified by DNA-based methods (Buse et al., 2014a,b; Gião et al., 2015). Consistent with this hypothesis, after batch incubations with copper ions, Proctor et al. (2017) reported sharper decreases in L. pneumophila numbers by plate counts versus qPCR. Also, copper (and other) materials influence the microbial composition of premise plumbing biofilms (Buse et al., 2014a; Proctor et al, 2018), with copper resulting in less biofilm growth than various hard and soft plastics (Proctor et al., 2018; van der Kooij et al., 2017). Interestingly, while less biofilm may accumulate on copper materials than on plastics, the types of bacteria and amoeba present could be more supportive of L. pneumophila growth than those on plastics (Buse et al., 2014a,b; Gião et al., 2015). In particular, V. vermiformis is the L. pneumophila host most often associated with warm- and hot-water (largely copper-pipe) systems (Buse et al., 2017; Ji et al., 2017). Hence, along with biofilm concentration, the species composition of the biofilm is important for growth of amoebae that favor L. pneumophila replication. Overall, L. pneumophila growth appears enhanced in biofilms dominated with a-Proteobacteria, key prey for protozoan hosts (van der Kooij et al., 2018).
***
Once within the complex plumbing of a large building, L. pneumophila may persist given the right combinations of temperature, stagnation, and subsequent loss of residual disinfectant, often exacerbated by the presence of iron oxides/hydroxioxides/humics (Butterfield et al., 2002) and other pipe corrosion products (Rhoads et al., 2017b). L. pneumophila strains have remained detectable in simulated building water systems for a long time (i.e., up to 2.4 years) (Paszko-Kolva et al., 1992; Skaliy and McEachern, 1979; Wadowsky and Yee, 1985), with the one apparent clone in buildings causing outbreaks over decades (Garcia-Nunez et al., 2008). This prolonged survival in water has been attributed to the organism’s ability to produce and store poly-3-hydrobutyrate, a carbon/energy source when nutrients are scarce (James et al., 1999; Mauchline et al., 1992). Recently, Shaheen et al. (2019) showed that viable cells of a L. pneumophila serogroup 1 strain remained in a dormant-like state associated with amoebae for more than two years in drinking water. Such persistence may be associated with the expression of a Type II secretion system, a transport mechanism that promotes bacterial growth at ambient drinking water temperatures (less than 25°C) (Söderberg et al., 2004). In cold water, Legionella persist within trophozoites and acquire
nutrients sufficient for maintenance from the host, but the population does not expand. Cold-water systems are unlikely to result in cases of legionellosis, except in warm climates where the water temperature exceeds 25°C for extended periods.
Cooling Towers and Industry Wastewater Treatment Works
Cooling towers provide a favorable environment for proliferation of Legionella due to not only their warm water temperatures but also the large surface area available for biofilm colonization. In addition, cooling towers can broadly disperse aerosols, from hundreds of meters to kilometres away (Nguyen et al., 2006). Cooling towers operate within the temperature window that supports L. pneumophila growth within amoebae (Berk et al., 2006; Canals et al., 2015; Critchley and Bentham, 2007; Hammer, 2018; Llewellyn et al., 2017; Nguyen et al., 2006; Scheikl et al., 2016), making control reliant primarily upon the use of biocides. It is also critical to position building air intakes well away from cooling tower-generated aerosols, while accounting for thermal inversion and other atmospheric phenomena (Engelhart et al., 2008).
Extensive outbreaks of legionellosis have also tracked to activated sludge treatment of pulp and paper mill and brewery effluents, which are typically not disinfected and are within 30°C to 40°C (Maisa et al., 2015; Nygård et al., 2008). On the other hand, conventional municipal wastewater treated by the activated sludge process typically has low levels of legionellae, probably in part due to the temperatures being below 30°C (Caicedo et al., 2018).
Hot Tubs
Public and private hot tubs are typically maintained at 38°C to 44°C, a favorable temperature range for L. pneumophila. A common feature linked to Legionella growth in hot tubs is biofilm build-up on internal plumbing surfaces that periodically contain stagnant warm water for extended periods (hours to days) (Costa et al., 2010). Sudden mobilization with entrained air and/or water circulation can aerosolize Legionella cells (likely associated with biofilm and amoebae) and lead to human infections (Fallon and Rowbotham, 1990; Leoni et al., 2018). Because hot tubs are inherently warm, there is a heavy reliance on biocides to suppress microbial growth (e.g., Qin et al., 2013). However, biocides can by no means eradicate all resident microbes, and these chemicals can also shape the microbial communities inhabiting the hot tubs. Hot tubs are Legionella infection risks not only for their users but also for attendants and patrons who pass down-wind of aerosols (Costa et al., 2010; Hamilton et al., 2018).
Other Devices
As with any device exposed to air and moisture, humidifiers and nebulizers can support biofilm growth and associated legionellae and deliver aerosols to susceptible individuals, particularly in healthcare settings (Kyritsi et al., 2018; Mastro et al., 1991; Yiallouros et al., 2013). For example, supermarket vegetable misters have been sources of Legionnaires’ disease (Barrabeig et al., 2010). Other devices that generate aerosols from stored/stagnating warm water sources and have been associated with cases of Legionnaires’ disease include car windshield wiper fluids and sprayers (Wallensten et al., 2010), car wash facilities (Baldovin et al., 2018), and decorative fountains (Decker and Palmore, 2013).
Plant Growth Media and Bagged Compost
The major non-water medium associated with cases of Legionnaires’ disease is garden soil and compost materials, which most often promote the growth and detection of L. longbeachae (Currie et al., 2014; Whiley and Bentham, 2011). However, L. longbeachae is probably under-reported even more than non-serogroup 1 L. pneumophila and may also occur within a range of environments discussed above such as cooling towers (Bacigalupe et al., 2017; Thornley et al., 2017).
EXPOSURE PATHWAYS
As pathogenic legionellae grow to high concentrations in free-living protozoa associated with biofilms, infectious Legionella cells may be released in aerosols in various forms: as free cells, cells within biofilm fragments, or cells associated with free-living protozoan trophozoites, cysts, or expelled vesicles (membrane-bound structures containing undigested materials) (see Figure 2-7; Lau and Ashbolt, 2009; Shaheen and Ashbolt, 2017). While Legionella infections are the most frequently identified etiologic agents from environmental aerosol exposures, other amoeba-resisting bacterial pathogens may also cause infections but go undetected because of inadequate methodologies used to identify agents (Lamonth and Greub, 2010; Lienard et al., 2017).
Common exposure pathways include breathing in aerosols in the 2- to 10-µm size range generated by showers, hot tubs, humidifiers, spray misters, cooling towers, car washers, windshield wiper spray, aeration basins used in wastewater treatment, and water (decorative) features and fountains. Aspiration of drinking water is another exposure pathway. A particular feature of L. longbeachae infections is the association with aerosols generated from wood and bark residuals and poorly matured compost sold as potting media in sealed plastic bags (Steel et al., 1990), since L. longbeachae and its supporting free-living protozoa hosts thrive in the biofilm environment associated with warm, partly decaying wood residuals.
Most reports of Legionnaires’ disease from building exposures involve hot water mixed with cold water used for showering or held in hot tubs. Cold tap water that has warmed, such as in humidifiers, or aspirated stagnant drinking water can also be problematic. Features that increase the likelihood of significant exposures include the use of hot- or cold-water storage tanks in buildings, aerators on taps, water conserving (finer misting) showerheads, and high-rate aeration of hot tubs.
Notably, most cysts of free-living protozoa are too large to reach the lower respiratory tract (i.e., they are greater than 10 µm). Larger biofilm fragments are also less likely to reach the sites for lung macrophage infection. In contrast, amoebae are known to expel cells and food in membrane-bound vesicles, for which a few hundred infectious L. pneumophila cells have been estimated per vesicle (Shaheen and Ashbolt, 2018), making Legionella-encapsulated vesicles a likely but rarely studied environmental form of exposure. Also, infected trophozoites may rupture within the airways, releasing several hundred free bacterial cells (Buse and Ashbolt, 2012) within the respiratory size. In support of amoeba-associated infectious L. pneumophila cells is the report of Acanthamoeba antibodies in people with legionellosis (C. Chappell, personal communication), but this association may be accidental, given the high likelihood for people to have antibodies to this genus of environmental amoebae (Chappell et al., 2001). Unfortuntely, neither the biochemical nor genetic attributes that promote L. pneumophila survival in aerosols have been identified, despite the large number of cases and isolates obtained from outbreaks (e.g., Bennett et al., 2014).
Physicochemical conditions of the aerosols may impact domestic, commercial, and industrial human exposures. For example, lung deposition may be more likely for bacteria in an isotonic solution (close to 0.9 percent salinity or 1,100 milliSiemens [mS]/cm2) (Haddrell et al., 2014). This level is typical of hot-tub aerosols, which are around 1,200 mS/cm2 compared to 4,000 mS/cm2 in cooling towers and less than 200 mS/cm2 in tap water. Hence, aerosols from hot tubs and perhaps cooling towers may be more likely to reach deep into the lungs (Richard Bentham, Flinders University, Adelaide, personal communication, May 14, 2018). The likelihood of aerosol entry into the lungs is further enhanced in hot tubs because people sitting in or near the units breathe in close proximity to the water surface (Moore et al., 2015). If atmospheric conditions such as relative humidity and wind direction are favorable, then Legionella-containing aerosols may infect people some tens of kilometers downwind (Nygård et al., 2008).
From epidemiology studies of Legionnaires’ disease in the United States (reviewed by Garrison et al., 2016), the common known pathways for exposure appear to be from potable water within buildings and from cooling tower aerosols, followed by hot tubs, fountains, and other devices. Among the building categories, hotels and resorts, long-term care facilities such as nursing homes, and hospitals are the most likely to be associated with an outbreak of Legionnaires’ disease. However, given that most cases of Legionnaires’ disease are sporadic (see Chapter 3) and the sources of such exposures are not identified, the relative disease burden from cooling towers, from homes and larger buildings, or from other engineered water features has yet to be determined.
CONCLUSIONS AND RECOMMENDATIONS
L. pneumophila is now the leading cause of reportable drinking water-associated disease outbreaks in the United States. To reduce this significant healthcare burden, a deeper understanding of Legionella ecology, the genetic traits that equip Legionella strains to colonize engineered water systems, and how Legionella survive in aerosols and thrive in the human lung is required. These and other important topics for future research are discussed in greater detail below.
There is a need to better understand the mechanistic pathways for the development of Pontiac fever, and what roles the pathogen, endotoxins, Legionella-harboring amoebae, or other exposures play in disease pathogenesis. Additional studies aimed at understanding the differences in Legionella species characteristics associated with Pontiac fever are also needed. Because Pontiac fever is associated with less mortality, focused studies examining this clinical entity have been limited to date. Pontiac fever reporting occurs primarily through outbreak investigations, which limits assessments of true incidence, population risk, and an understanding of the relationship between certain Legionella species and serotypes and disease manifestations. There is a need to develop improved diagnostic tools for Pontiac fever (including molecular methods) that would enhance overall Legionella epidemiology and outbreak investigation and detection.
There is a need to better characterize legionellosis among neonates, young children, and adolescents, who may have varied epidemiologic risk factors for exposure to Legionella and differing risk for disease manifestations. There have been limited studies of Legionnaires’ disease among children. The majority of current guidelines and recommendations focus on adult disease or target high-risk patient populations (e.g., immunocompromised hosts).
Studies of community-acquired pneumonia, fever, and non-respiratory viral influenza-like illness among pediatric populations that assess Legionella are needed.
Studies that further assess the contribution of aspiration of potable water as a mechanism for clinical legionellosis both in community outbreaks and among sporadic cases are needed. Inhaled respiratory droplets are thought to be the primary mode of exposure to Legionella, but other methods of exposure through water-based contact are poorly characterized. Aspiration, including microaspiration and silent aspiration, is thought to be potentially linked to legionellosis. Links between feeding tubes and among conditions that increase risk for aspiration, together with studies demonstrating potential short-term oral colonization, suggest potential risks from this pathway.
The capacity of L. pneumophila to resist detergents, heat, chemical disinfectants, and antibiotics, as well as predatory amoebae and white blood cells, depends on its growth phase. The resilience and infection potential differs by orders of magnitude for replicative, stationary or transmissive phase, and the mature infectious form of L. pneumophila. Therefore, protocols should be developed to generate, identify, enumerate, and report distinct Legionella cell types. These classifications are fundamental to not only experimental reproducibility but also the development and accurate assessment of strategies to eliminate virulent legionellae from patients and engineered water systems.
Whether L. pneumophila persistence within built water systems is promoted by the bacterium’s differentiation into an apparent viable-but-non-culturable state that is both resilient and reversible remains an urgent question with implications for public health. To date, studies of VBNC-like L. pneumophila are largely descriptive; for example, quantification of resuscitation from the VBNC-like form is lacking. Needed are protocols to generate and isolate pure populations of VBNC-like cells for physiological, biochemical, genetic, molecular, and infection studies. Molecular or biochemical markers that distinguish individual VBNC-like, MIF, replicating, and stationary phase L. pneumophila cells would also accelerate this research and enable experimentalists to distinguish true resuscitation of VBNC-like cells from regrowth of a minor population of cells.
Ecological studies have almost exclusively focused on the impact of environmental conditions on growth, survival, and inactivation of L. pneumophila. To clarify whether the ecological principles observed for L. pneumophila apply to other pathogenic Legionella species, research on the ecology of L. longbeachae, L. micdadei, L. dumoffi, and other pathogenic Legionella species is warranted. In addition, studies on the ecology of L. pneumophila have focused primarily on building water systems, leaving the ecological conditions responsible for L. pneumophila growth in other environments (e.g., cooling towers, wastewater treatment plants, hot springs, soils) largely unexplored. New research in these two areas could result in improved control measures for other pathogenic Legionella species and for L. pneumophila in environments other than premise plumbing.
The acceleration of genomics in the clinical, environmental, and laboratory sciences has expanded awareness of the extraordinary diversity within the Legionella genus. Yet, most knowledge of L. pneumophila pathogenesis comes from a relatively small number of domesticated laboratory strains. A wealth of genomic data can now inform research to identify specific genetic markers of resilience and virulence among the environmental and clinical
Legionella and accelerate the rational design of risk assessment tools and environmental remediation methods.
Clarifying the diversity of free-living protozoa that support or diminish the intracellular growth of legionellae in water and soil environments is fundamental to understanding and controlling legionellosis. Whether legionellae persist within free-living protozoa versus growing to high numbers appears to be influenced by many poorly understood factors, including temperature, species of bacterial prey available, presence of host symbionts, and host cell form. Direct (locational) observations and metagenomic studies of microbial diversity are required to provide a stronger ecological foundation to identify the protozoa that control the growth of pathogenic Legionella in various environments. Microcosm studies could investigate how nutrients and biocides affect the life stages of the host protozoa (e.g., by triggering encystation), identify the key host species, and elucidate the role of other free-living protozoa that might feed on the primary hosts of legionellae.
Despite clear evidence that people acquire Legionnaries’ disease by inhaling contaminated water or soil, neither the biochemical nor genetic attributes that equip Legionella bacteria to survive in aerosols have been identified. Knowledge of the antigens or genetic loci that confer resilience of airborne legionellae and other factors like packaging (within biofilm, vesicles, trophozoites) would guide risk assessment of microbes that colonize engineered water systems.
REFERENCES
Abdelhady, H., and R. A. Garduño. 2013. The progeny of Legionella pneumophila in human macrophages shows unique developmental traits. FEMS Microbiology Letters 349: 99-107.
Abu Kwaik, Y., L.-Y. Gao, B. J. Stone, C. Venkataraman, and O. S. Harb. 1998. Invasion of protozoa by Legionella pneumophila and its role in bacterial ecology and pathogenesis. Applied and Environmental Microbiology 64:3127-3133.
Al-Bana, B. H., M. T. Haddad, and R. A. Garduño. 2014. Stationary phase and mature infectious forms of Legionella pneumophila produce distinct viable but non-culturable cells. Environmental Microbiology 16(2):382-395.
Albers, U., A. Tiaden, T. Spirig, D. Al Alam, S. M. Goyert, S. C. Gangloff, and H. Hilbi. 2007. Expression of Legionella pneumophila paralogous lipid A biosynthesis genes under different growth conditions. Microbiology 153(11):3817-3829.
Alexander, N. T., B. S. Fields, and L. A. Hicks. 2008. Epidemiology of reported pediatric Legionnaires’ disease in the United States, 1980–2004. Presented at 48th Interscience Conference on Antimicrobial Agents and Chemotherapy, Washington, DC. Abstract #G1–1694.
Allard, K. A., J. Dao, P. Sanjeevaiah, K. McCoy-Simandle, C. H. Chatfield, D. S. Crumrine, D. Castignetti, and N. P. Cianciotto. 2009. Purification of legiobactin and importance of this siderophore in lung infection by Legionella pneumophila. Infection and Immunology 77(7):2887-2895.
Allegra, S., F. Berger, P. Berthelot, F. Grattard, B. Pozzetto, and S. Riffard. 2008. Use of flow cytometry to monitor Legionella viability. Applied and Environmental Microbiology 74(24):7813-7816.
Allegra, S., F. Grattard, F. Girardot, S. Riffard, B. Pozzetto, and P. Berthelot. 2011. Longitudinal evaluation of the efficacy of heat treatment procedures against Legionella spp. in hospital water systems by using a flow cytometric assay. Applied and Environmental Microbiology 77(4):1268-1275.
Alleron, L., N. C. Merlet, C. Lacombe, and J. Frère. 2008. Long-term survival of Legionella pneumophila in the viable but nonculturable state after monochloramine treatment. Current Microbiology 57:497-502.
Almahmoud, I., E. Kay, D. Schneider, and M. Maurin. 2009. Mutational paths towards increased fluoroquinolone resistance in Legionella pneumophila. Journal of Antimicrobial Chemotherapy 64(2):284-293.
Amaro, F., W. Wang, J. A. Gilbert, O. R. Anderson, and H. A. Shuman. 2015. Diverse protist grazers select for virulence-related traits in Legionella. International Society for Microbial Ecology Journal 9(7):1607-1618.
Amemura-Maekawa, J., K. Kikukawa, J. H. Helbig, S. Kaneko, A. Suzuki-Hashimoto, K. Furuhata, B. Chang, M. Murai, M. Ichinose, M. Ohnishi, F. Kura, and the Working Group for Legionella in Japan. 2012. Distribution of monoclonal antibody subgroups and sequence-based types among Legionella pneumophila serogroup 1 isolates derived from cooling tower water, bathwater, and soil in Japan. Applied and Environmental Microbiology 78(12):4263-4270.
Ampel, N., and E. Wing. 1990. Legionella infection in transplant patients. Seminars in Respiratory Infections 5(1):30-37.
Anand, C. M., A. R. Skinner, A. Malic, and J. B. Kurtz. 1983. Interaction of L. pneumophila and a free living amoeba (Acanthamoeba palestinensis). Journal of Hygiene 91:167-178.
Appelt, S., and K. Heuner. 2017. The flagellar regulon of Legionella: A review. Frontiers in Cell Infection and Microbiology 7:454.
Arcavi, L., and N. L. Benowitz. 2004. Cigarette smoking and infection. Archives of Internal Medicine 164(20):2206-2216.
ASCE (American Society of Civil Engineers). 2017. Infrastructure report card—Drinking water. https://www.infrastructurereportcard.org/wp-content/uploads/2017/01/Drinking-Water-Final.pdf.
Ashbolt, N. J. 2015. Environmental (saprozoic) pathogens of engineered water systems: Understanding their ecology for risk assessment and management. Pathogens 4(2):390-405.
Avni, T., A. Bieber, H. Green, T. Steinmetz, L. Leibovici, and M. Paul. 2016. Diagnostic accuracy of PCR alone and compared to urinary antigen for the diagnosis of Legionella spp.: Systematic review. Journal of Clinical Microbiology 54(2):401-411.
Ayrapetyan, M., T. C. Williams, and J. D. Oliver. 2014. Interspecific quorum sensing mediates the resuscitation of viable but nonculturable vibrios. Applied and Environmental Microbiology 80(8):2478-2483.
Bacigalupe, R., D. Lindsay, G. Edwards, and J. R. Fitzgerald. 2017. Population genomics of Legionella longbeachae and hidden complexities of infection source attribution. Emerging Infectious Diseases 23(5):750-757.
Baldovin, T., A. Pierobon, C. Bertoncello, E. Destefani, M. Gennari, A. Stano, and V. Baldo. 2018. May car washing represent a risk for Legionella infection? Annali di Igiene 30(1):57-65.
Banderet, F., A. Blaich, E. Soleman, V. Gaia, and M. Osthoff. 2017. Septic arthritis due to Legionella cincinnatiensis: Case report and review of the literature. Infection 45(4):551-555.
Bargellini, A., I. Marchesi, E., E. Righi, A. Ferrari, S. Cencetti, P. Borella, and S. Rovesti. 2011. Parameters predictive of Legionella contamination in hot water systems: Association with trace elements and heterotrophic plate counts. Water Research 45(6):2315-2321.
Bargellini, A., I. Marchesi, P. Marchegiano, L. Richeldi, R. Cagarelli, G. Ferranti, and P. Borella. 2013. A culture-proven case of community-acquired Legionella pneumonia apparently classified as nosocomial: Diagnostic and public health implications. Case Reports in Medicine 2013:303712.
Barker, J., M. R. Brown, P. J. Collier, I. Farrell, and P. Gilbert. 1992. Relationship between Legionella pneumophila and Acanthamoeba polyphaga: Physiological status and susceptibility to chemical inactivation. Applied and Environmental Microbiology 58(8):2420-2425.
Barker, J., P. A. Lambert, and M. R. Brown. 1993. Influence of intra-amoebic and other growth conditions on the surface properties of Legionella pneumophila. Infection and Immunology 61(8):3503-3510.
Barker, J., H. Scaife, and M. R. Brown. 1995. Intraphagocytic growth induces an antibiotic-resistant phenotype of Legionella pneumophila. Antimicrobial Agents and Chemotherapy 39(12):2684-2688.
Barna, Z., M. Kádár, E. Kálmán, A. M. Szax, and M. Vargha. 2016. Prevalence of 18 Legionella in premise plumbing in Hungary. Water Research 90:71-78.
Barrabeig, I., A. Rovira, M. Garcia, J. M. Oliva, A. Vilamala, M. D. Ferrer, M. Sabrià, and A. Domínguez. 2010. Outbreak of Legionnaires’ disease associated with a supermarket mist machine. Epidemiology and Infection 138(12):1823-1828.
Bartram, J. 2007. Legionella and the prevention of legionellosis. J. Bartram, Y. Chartier, J.V. Lee, K. Pond, S. Surman-Lee (eds.). World Health Organization. doi:10.3201/eid1406.080345.
Beauté, J. 2017. On behalf of the European Legionnaires’ Disease Surveillance Network. Legionnaires’ disease in Europe, 2011 to 2015. Eurosurveillance 22(27):30566. doi:10.2807/1560-7917.ES.2017.22.27.30566.
Beauté, J., P. Zucs, and B. de Jong. 2013. On behalf of the European Legionnaires’ Disease Surveillance Network. Legionnaires’ disease in Europe, 2009–2010. Eurosurveillance 18(10):20417.
Beer, K. D., J. W. Gargano, V. A. Roberts, V. R. Hill, L. E. Garrison, P. K. Kutty, E. D. Hilborn, T. J. Wade, K. E. Fullerton, and J. S. Yoder. 2015. Surveillance for waterborne disease outbreaks associated with drinking water—United States, 2011–2012. Morbidity and Mortality Weekly Report 64(31):842-848.
Benin, A. L., R. F. Benson, and R. E. Besser. 2002. Trends in Legionnaires’ disease, 1980–1998: Declining mortality and new patterns of diagnosis. Clinical Infectious Diseases 35(9):1039-1046.
Benitez, A. J., and J. M. Winchell. 2013. Clinical application of a multiplex real-time PCR assay for simultaneous detection of Legionella species, Legionella pneumophila, and Legionella pneumophila serogroup 1. Journal of Clinical Microbiology 51(1):348-351.
Bennett, E., M. Ashton, N. Calvert, J. Chaloner, J. Cheesbrough, J. Egan, I. Farrell, I. Hall, T. G. Harrison, F. C. Naik, S. Partridge, Q. Syed, and R. N. Gent. 2014. Barrow-in-furness: A large community legionellosis outbreak in the UK. Epidemiology and Infection 142(8):1763-1777.
Benz-Lemoine, E. V. Delwail, O. Castel, F. Guilhot, R. Robert, G. Grollier, F. Roblot-Casenave, C. Giraud, and J. Tanzer. 1991. Nosocomial Legionnaires’ disease in a bone marrow transplant unit. Bone Marrow Transplantation 7(1):61-63.
Berk, S. G., R. S. Ting, G. W. Turner, and R. J. Ashburn. 1998. Production of respirable vesicles containing live Legionella pneumophila cells by two Acanthamoeba spp. Applied and Environmental Microbiology 64:279-286.
Berk, S. G., J. H. Gunderson, A. L. Newsome, A. L. Farone, B. J. Hayes, K. S. Redding, N. Uddin, E. L. Williams, R. A. Johnson, M. Farsian, A. Reid, J. Skimmyhorn, and M. B. Farone. 2006. Occurrence of infected amoebae in cooling towers compared with natural aquatic environments: Implications for emerging pathogens. Environmental Science and Technology 40(23):7440-7444.
Berrington, W. R., and T. R. Hawn. 2013. Human susceptibility to Legionnaires’ disease. Pp. 541-551 In: Legionella Methods in Molecular Biology (Methods and Protocols). C. Buchrieser and H. Hilbi (eds.). Totowa, NJ: Humana Press.
Blatt, S. P., M. D. Parkinson, E. Pace, P. Hoffman, D. Dolan, P. Lauderdale, R. A. Zajac, and G. P. Melcher. 1993. Nosocomial Legionnaires’ disease: aspiration as a primary mode of disease acquisition. American Journal of Medicine 95(1):16-22.
Boe, D. M., L. A. Boule, and E. J. Kovacs. 2017. Innate immune responses in the ageing lung. Clinical and Experimental Immunology 187(1):16–25.
Borella, P., E. Guerrieri, I. Marchesi, M. Bondi, and P. Messi. 2005. Water ecology of Legionella and protozoan: Environmental and public health perspectives. Biotechnology Annual Reviews 11:355-380.
Borella, P., A. Bargellini, I. Marchesi, S. Rovesti, G. Stancanelli, S. Scaltriti, M. Moro, M. Montagna, D. Tatò, C. Napoli, M. Triassi, S. Montegrosso, F. Pennino, C. M. Zotti, S. Ditommaso, M. Giacomuzzi. 2008. Prevalence of anti-Legionella antibodies among Italian hospital workers. Journal of Hospital Infection 69(2):148-155.
Boswell, T. C., L. E. Marshall, and G. Kudesia. 1996. False-positive Legionella titres in routine clinical serology testing detected by absorption with Campylobacter: Implications for the serological diagnosis of Legionnaires’ disease. Journal of Infection 32(1):23-26.
Botelho-Nevers, E., F. Grattard, A. Viallon, S. Allegra, S. Jarraud, P. Verhoeven, A. Marcuccilli, F. Lucht, B. Pozzetto, and P. Berthelot. 2016. Prospective evaluation of RT-PCR on sputum versus culture, urinary antigens and serology for Legionnaire’s disease diagnosis. Journal of Infection 73(2):123-128.
Bradley, J. S., C. L. Byington, S. S. Shah, B. Alverson, E. R. Carter, C. Harrison, S. L. Kaplan, S. E. Mace, G. H. McCracken Jr, M. R. Moore, S. D. St Peter, J. A. Stockwell, and J. T. Swanson. 2011. The management of community-acquired pneumonia in infants and children older than 3 months of age. Clinical Practice Guidelines by the Pediatric Infectious Diseases Society and the Infectious Diseases Society of America. Clinical Infectious Diseases 53(7):e25-76.
Braun, R. S., N. Mendis, L. Li, and S. P. Faucher. 2019. Quantification of viable but non-culturable cells of Legionella pneumophila. Pp. 45-53 In: Legionella: Methods in Molecular Biology. C. Buchrieser and H. Hilbi (eds.). NewYork: Humana Press.
Brazeau, R. H., and M. A. Edwards. 2013. Role of hot water system design on factors influential to pathogen regrowth: Temperature, chlorine residual, hydrogen evolution, and sediment. Environmental Engineering and Science 30(10):617-627.
Brieland, J. K., J. C. Fantone, D. G. Remick, M. LeGendre, M. McClain, and N. C. Engleberg. 1997. The role of Legionella pneumophila-infected Hartmanella vermiformis as an infectious particle in a murine model of Legionnaire’s disease. Infection and Immunity 65:5330-5333.
Brüggemann, H., A. Hagman, M. Jules, O. Sismeiro, M. A. Dillies, C. Gouryett, F. Kunst, M. Steinert, K. Heuner, J. Y. Coppée, and C. Buchrieser. 2006. Virulence strategies for infecting phagocytes deduced from the in vivo transcriptional program of Legionella pneumophila. Cellular Microbiology 8:1228-1240.
Bruin, J. P., T. Koshkolda, E. P. F. I. Jzerman, C. Lück, B. M. Diederen, J. W. den Boer, and J. W. Mouton. 2014. Isolation of ciprofloxacin-resistant Legionella pneumophila in a patient with severe pneumonia. Journal of Antimicrobial Chemotherapy 69(10):2869-2871.
Buracco, S., B. Peracino, C. Andreini, E. Bracco, and S. Bozzaro. 2018. Differential effects of iron, zinc, and copper on Dictyostelium discoideum cell growth and resistance to Legionella pneumophila. Frontiers in Cellular and Infection Microbiology 7:536. doi: 10.3389/fcimb.2017.00536.
Burdet, C., R. Lepeule, X. Duval, M. Caseris, C. Rioux, J. C. Lucet, and Y. Yazdanpanah. 2014. Quinolones versus macrolides in the treatment of legionellosis: A systematic review and meta-analysis. Journal of Antimicrobial Chemotheropy 69(9):2354-2360.
Burillo, A., M. L. Pedro-Botet, and E. Bouza. 2017. Microbiology and epidemiology of Legionnaire’s disease. Infectious Disease Clinics of North America 31(1):7-27.
Burnsed, C. J., L. A. Hicks, L. M. K. Smithee, B. S. Fields, K. K. Bradley, N. Pascoe, S. M. Richards, S. Mallonee, L. Littrell, R. F. Benson, M. R. Moore, and the Legionellosis Outbreak Investigation Team. 2007. A large, travel-associated outbreak of legionellosis among hotel guests: Utility of the urine antigen assay in confirming Pontiac fever. Clinical Infection and Disease 44:222-228.
Buse, H. Y., and N. J. Ashbolt. 2011. Differential growth of Legionella pneumophila strains within a range of amoebae at various temperatures associated with in-premise plumbing. Letters in Applied Microbiology 53(2):217-224.
Buse, H. Y., and N. J. Ashbolt. 2012. Counting Legionella cells within single amoeba host cells. Applied and Environmental Microbiology 78(6):2070-2072.
Buse, H. Y., M. E. Schoen, and N. J. Ashbolt. 2012. Legionellae in engineered systems and use of quantitative microbial risk assessment to predict exposure. Water Research 46(4):921-933.
Buse, H. Y., J. Lu, X. Lu, X. Mou, and N. J. Ashbolt. 2014a. Microbial diversities (16S and 18S rRNA gene pyrosequencing) and environmental pathogens within drinking water biofilms grown on the common premise plumbing materials unplasticized polyvinylchloride and copper. FEMS Microbiology Ecology 88:280-295.
Buse, H. Y., J. Lu, I. T. Struewing, and N. J. Ashbolt. 2014b. Preferential colonization and release of Legionella pneumophila from mature drinking water biofilms grown on copper versus unplasticized polyvinylchloride coupons. International Journal of Hygiene and Environmental Health 217(1):219-225.
Buse, H. Y., P. Ji, V. Gomez-Alvarez, A. Pruden, M. A. Edwards, and N. J. Ashbolt. 2017. Effect of temperature and colonization of Legionella pneumophila and Vermamoeba vermiformis on bacterial community composition of copper drinking water biofilms. Microbial Biotechnology 88(2):280-295.
Butterfield, P. W., A. K. Camper, J. A. Biederman, and A. M. Bargmeyer. 2002. Minimizing biofilm in the presence of iron oxides and humic substances. Water Research 36(15):3898-3910.
Byrne, B., and M. S. Swanson. 1998. Expression of Legionella pneumophila virulence traits in response to growth conditions. Infection and Immunology 66:3029-2034.
Byrne, B. G., S. McColm, S. P. McElmurry, P. E. Kilgore, J. Sobeck, R. Sadler, N. G. Love, and M. S. Swanson. 2018. Prevalence of infection-competent serogroup 6 Legionella pneumophila within premise plumbing in southeast Michigan. mBio 9(1):e00016-e00018.
Caicedo, C., K. H. Rosenwinkel, and R. Nogueira. 2018. Temperature-driven growth of Legionella in lab-scale activated sludge systems and interaction with protozoa. International Journal of Hygiene and Environmental Health 221(2):315-322.
Caicedo, C., K. H. Rosenwinkel, M. Exner, W. Verstraete, R. Suchenwirth, P. Hartemann, and R. Nogueira. 2019. Legionella occurrence in municipal and industrial wastewater treatment plants and risks of reclaimed wastewater reuse. Water Research 149:21-34.
Canals, O., A. Serrano-Suarez, H. Salvado, J. Mendez, S Cervero-Arago, V. Ruiz de Porras, J. Dellunde, and R. Araujo. 2015. Effect of chlorine and temperature on free-living protozoa in operational man-made water systems (cooling towers and hot sanitary water systems) in Catalonia. Environmental Science and Pollution Research International 22(9):6610-6618.
Carvalho, F. R. S., R. F. Vazoller, A. S. Foronda, and V. H. Pellizari. 2007. Phylogenetic study of Legionella species in pristine and polluted aquatic samples from a tropical Atlantic forest ecosystem. Current Microbiology 55(4):288-293.
Carvalho, F. R. S., F. R. Nastasi, R. C. Gamba, A. S. Foronda, and V. H. Pellizari. 2008. Occurrence and diversity of Legionellaceae in polar lakes of the Antarctic Peninsula. Current Microbiology 57(4):294-300.
Cassell, K., P. Gacek, J. L. Warren, P. A. Raymond, M. Cartter, and D. M. Weinberger. 2018. Association between sporadic legionellosis and river systems in Connecticut. Journal of Infectious Diseases 217:179-187.
Cassier, P., C. Campese, Y. Le Strat, D. Che, C. Ginevra, J. Etienne, and S. Jarraud. 2015. Epidemiologic characteristics associated with ST23 clones compared to ST1 and ST47 clones of Legionnaires disease cases in France. New Microbes New Infections 3:29-33.
Cazalet, C., S. Jarraud, Y. Ghavi-Helm, F. Kunst, P. Glaser, J. Etienne, and C. Buchrieser. 2008. Multigenome analysis identifies a worldwide distributed epidemic Legionella pneumophila clone that emerged within a highly diverse species. Genome Research 18(3):431-441.
Cazalet, C., L. Gomez-Valero, C. Rusniok, M. Lomma, D. Dervins-Ravault, H. J. Newton, F. M. Sansom, S. Jarraud, N. Zidane, L. Ma, C. Bouchier, J. Etienne, E. L. Hartland, and C. Buchrieser. 2010. Analysis of the Legionella longbeachae genome and transcriptome uncovers unique strategies to cause Legionnaires’ disease. PLoS Genetics 6(2):e1000851.
CDC (Centers for Disease Control and Prevention). 2013. National Center for Chronic Disease Prevention and Health Promotion Division of Population Health. https://www.cdc.gov/aging/pdf/State-Aging-Health-inAmerica-2013.pdf.
CDC. 2017. Legionella (Legionnaire’s disease and Pontiac fever)—Clinical features. https://www.cdc.gov/legionella/clinicians/clinical-features.html.
Cervero-Aragó, S., S. Rodriguez-Martínez, O. Canals, H. Salvado, and R. M. Araujo. 2013. Effect of thermal treatment on free-living amoeba inactivation. Journal of Applied Microbiology 116(3):728-736.
Cervero-Aragó, S., R. Sommer, and R. M. Araujo. 2014. Effect of UV irradiation (253.7 nm) on free Legionella and Legionella associated with its amoebae hosts. Water Research 67:299-309.
Cervero-Aragó, S., B. Schrammel, E. Dietersdorfer, R. Sommer, C. Lück, J. Walochnik, and A. Kirschner. 2019. Viability and infectivity of viable but nonculturable Legionella pneumophila strains induced at high temperatures. Water Research 158:268-279.
Chappell, C. L., J. A. Wright, M. Coletta, and A. L. Newsome. 2001. Standardized method of measuring acanthamoeba antibodies in sera from healthy human subjects. Clinical and Diagnostic Laboratory Immunology 8(4):724-730.
Che, D., C. Campese, P. Santa-Olalla, G. Jacquier, D. Bitar, P. Bernillon, and J. C. Desenclos. 2008. Sporadic community-acquired Legionnaires’ disease in France: A 2-year national matched case-control study. Epidemiology and Infection 136(12):1684-1690.
Chen, D. J., G. W. Procop, S. Vogel, B. Yen-Lieberman, and S. S. Richter. 2015. Utility of PCR, culture, and antigen detection methods for diagnosis of legionellosis. Journal of Clinical Microbiology 53(11):3474-3477.
Chen, N. T., M. J. Chen, C. Y. Guo, K. T. Chen, and H. J. Su. 2014. Precipitation increases the occurrence of sporadic Legionnaires’ disease in Taiwan. PLoS One 9:e114337.
Chidiac, C., D. Che, S. Pires-Cronenberger, S. Jarraud, C. Campese, A. Bissery, P. Weinbreck, C. Brun-Buisson, J. P. Sollet, R. Ecochard, J. C. Desenclos, J. Etienne, P. Vanhems, and the French Legionnaires’ Disease Study Group. 2012. Factors associated with hospital mortality in community-acquired legionellosis in France. European Respiratory Journal 39(4):963-970.
Cianciotto, N. P. 2015. An update on iron acquisition by Legionella pneumophila: New pathways for siderophore uptake and ferric iron reduction. Future Microbiology 10:841-851.
Ciesielski, C. A., M. J. Blaser, and W. L. Wang. 1984. Role of stagnation and obstruction of water flow in isolation of Legionella pneumophila from hospital plumbing. Applied and Environmental Microbiology 48(5):984-987.
Ciesielski, C. A., M. J. Blaser, and W. L. Wang. 1986. Serogroup specificity of Legionella pneumophila is related to lipopolysaccharide characteristics. Infection and Immunity 51:397-404.
Cirillo, J. D., S. Falkow, and L. S. Tompkins. 1994. Growth of Legionella pneumophila in Acanthamoeba castellanii enhances invasion. Infection and Immunity 62:3254-3261.
Collier, S., L. Stockman, L. Hicks, L. Garrison, F. Zhou, and M. Beach. 2012. Direct healthcare costs of selected diseases primarily or partially transmitted by water. Epidemiology and Infection 140:2003-2013.
Collins, S., D. Stevenson, A. Bennett, and J. Walker. 2017. Occurrence of Legionella in UK household showers. International Journal of Hygiene and Environmental Health 220(2 Pt B):401-406.
Coniglio, M. A., S. Pignato, and G. Giammanco. 2009. Prevalence of antibodies against Legionella spp. in HIV-infected subjects and blood donors. Journal of Infection 59(6):423-425.
Conlan, J. W., and L. A. E. Ashworth. 1986. The relationship between the serogroup antigen and lipopolysaccharide of Legionella pneumophila. Journal of Hygiene 96:39-48.
Correia, A. M., J. Goncalves, J. P. Gomes, J. S. Ferreira, V. Borges, A. Nunes, B. Gomes, R. Capucho, D. M. Antunes, S. Almeida, A. Mendes, M. Guerreiro, D. A. Sampaio, L. Vieira, J. Machado, M. J. Simoes, and P. Goncalves. 2016. Probable person-to-person transmission of Legionnaires’ disease. New England Journal of Medicine 374(5):497-498.
Coscollá, M., C. Fernández, J. Colomina, L. Sánchez-Busó, and F. González-Candelas. 2014. Mixed infection by Legionella pneumophila in outbreak patients. International Journal of Medical Microbiology 304(3-4):307-313.
Costa, J., I. Tiago, M.S. da Costa, and A. Verissimo. 2005. Presence and persistence of Legionella spp. in groundwater. Applied and Environmental Microbiology 71(2):663-671.
Costa, J., M. S. da Costa, and A. Verissimo. 2010. Colonization of a therapeutic spa with Legionella spp.: a public health issue. Research in Microbiology 161(1):18-25.
Cramp, G. J., D. Harte, N. M. Douglas, F. Graham, M. Schousboe, and K. Sykes. 2010. An outbreak of Pontiac fever due to Legionella longbeachae serogroup 2 found in potting mix in a horticultural nursery in New Zealand. Epidemiology and Infection 138(1):15-20.
Critchley, M., and R. Bentham. 2007. Legionella and protozoa in cooling towers: Implications for public health and chemical control. Environmental Health 7(2):36-44.
Cross, K. E., J. W. Mercante, A. J. Benitez, E. W. Brown, M. H. Diaz, and J. M. Winchell. 2016. Simultaneous detection of Legionella species and L. anisa, L. bozemanii, L. longbeachae, and L. micdadei using conserved primers and multiple probes in a multiplex real-time PCR assay. Diagnostic Microbiology and Infectious Disease 85(3):295-301.
Cunha, B. A., A. Burillo, and E. Bouza. 2016. Legionnaires’ disease. Lancet 387(10016):376-385.
Cunha, B. A. 1998. Clinical features of Legionnaires’ disease. Seminars in Respiratory Infections 13(2):116-127.
Currie, S. L., T. K. Beattie, C. W. Knapp, and D. S. Lindsay. 2014. Legionella spp. in UK composts—A potential public health issue? Clinical Microbiology and Infection 20(4):O224-9.
Dai, D., W. J. Rhoads, M. A. Edwards, and A. Pruden. 2018a. Shotgun metagenomics reveals taxonomic and functional shifts in hot water microbiome due to temperature setting and stagnation. Frontiers in Microbiology 9:2695.
Dai, D., C. R. Proctor, K. Williams, M. A. Edwards, and A. Pruden. 2018b. Mediation of effects of biofiltration on bacterial regrowth, Legionella pneumophila, and the microbial community structure under hot water plumbing conditions. Environmental Science: Water Research and Technology 4(2):183-194.
Dalebroux, Z. D., R. L. Edwards, and M. S. Swanson. 2009. SpoT governs Legionella pneumophila differentiation in host macrophages. Molecular Microbiology 71:640-658.
David, S., L. Sánchez-Busó, S. R. Harris, P. Marttinen, C. Rusniok, C. Buchrieser, T. G. Harrison, and J. Parkhill. 2017. Dynamics and impact of homologous recombination on the evolution of Legionella pneumophila. PLoS Genetics 13(6):e1006855.
De Jonckheere, J. F. 1980. Growth characteristics, cytopathic effect in cell culture, and virulence in mice of 36 type strains belonging to 19 different Acanthamoeba spp. Applied and Environmental Microbiology 39(4):681-685.
Decker, B. K., and T. N. Palmore. 2013. The role of water in healthcare-associated infections. Current Opinions in Infectious Disease 26(4):345-351.
Declerck, P., L. Vanysacker, A. Hulsmans, N. Lambert, S. Liers, and F. Ollevier. 2010. Evaluation of power ultrasound for disinfection of both Legionella pneumophila and its environmental host Acanthamoeba castellanii. Water Research 44(3):703-710.
del Castillo, M., A. Lucca, A. Plodkowski, Y. T. Huang, J. Kaplan, J. K. Gilhuley, N. E. Babady, S. Seo, and M. Kamboj. 2016. Atypical presentation of Legionella pneumonia among patients with underlying cancer: A fifteen-year review. Journal of Infection 72(1):45-51.
Denet, E., B. Coupat-Goutaland, S. Nazaret, M. Pélandakis, and S. Favre-Bonté. 2017. Diversity of free-living amoebae in soils and their associated human opportunistic bacteria. Parasitology Research 116:3151-3162.
Dennis, P. J., D. Green, and B. P. C. Jones. 1984. A note on the temperature tolerance of Legionella. Journal of Applied Bacteriology 56:349-350.
Descours, G., P. Cassier, F. Forey, C. Ginevra, J. Etienne, G. Lina, and S. Jarraud. 2014. Evaluation of BMPA, MWY, GVPC and BCYE media for the isolation of Legionella species from respiratory samples. Journal of Microbiological Methods 98:119-121.
Descours, G., C. Ginevra, N. Jacotin, F. Forey, J. Chastang, E. Kay, J. Etienne, G. Lina, P. Doublet, and S. Jarraud. 2017. Ribosomal mutations conferring macrolide resistance in Legionella pneumophila. Antimicrobial Agents and Chemotherapy 61(3):e02188-16.
Dey, R., J. Bodennec, M. O. Mameri, and P. Pernin. 2009. Free-living freshwater amoebae differ in their susceptibility to the pathogenic bacterium Legionella pneumophila. FEMS Microbiology Letters 290(1):10-17.
Dietersdorfer, E., A. Kirschner, B. Schrammel, A. Ohradanova-Repic, H. Stockinger, R. Sommer, J. Walochnik, and S. Cervero-Arago. 2018. Starved viable but non-culturable (VBNC) Legionella strains can infect and replicate in amoebae and human macrophages. Water Research 141:428-438.
Dilger, T., H. Melzl, and A. Gessner. 2018. Legionella contamination in warm water systems: A species-level survey. International Journal of Hygiene and Environmental Health 221:199-210.
Doleans, A., H. Aurell, M. Reyrolle, G. Lina, J. Freney, F. Vandenesch, J. Etienne, and S. Jarraud. 2004. Clinical and environmental distributions of Legionella strains in France are different. Journal of Clinical Microbiology 42(1):458-460.
Donohue, M. J., K. O’Connell, S. J. Vesper, J. H. Mistry, D. King, M. Kostich, and S. Pfaller. 2014. Widespread molecular detection of Legionella pneumophila serogroup 1 in cold water taps across the United States. Environmental Science and Technology 48(6):3145-3152.
Dooling, K. L., K.-A. Toews, L. A. Hicks, L. E. Garrison, B. Bachaus, S. Zansky, L. R. Carpenter, B. Schaffner, E. Parker, S. Petit, A. Thomas, S. Thomas, R. Mansmann, C. Morin, B. White, and G. E. Langley. 2015. Active bacterial core surveillance for Legionellosis—United States, 2011–2013. Morbidity and Mortality Weekly Report 64(42):1190-1193.
Dournon, E., A. Bure, N. Desplaces, and M. Carette. 1982. Legionnaires disease related to gastric lavage with tap water [letter]. Lancet 1:797-798.
Dowling, J. N., D. A. McDevitt, and A. W. Pasculle. 1985. Isolation and preliminary characterization of erythromycin-resistant variants of Legionella micdadei and Legionella pneumophila. Antimicrobial Agents and Chemotherapy 27(2):272-274.
Dowling, J. N., W. Pasculle, F. N. Frola, M. K. Zaphyr, and B. B. Yee. 1984. Infections caused by Legionella micdadei and Legionella pneumophila among renal transplant recipients. Journal of Infectious Disease 149(5):703-713.
Ducret, A., M. Chabalier, and S. Dukan. 2014. Characterization and resuscitation of “non-culturable” cells of Legionella pneumophila. BMC Microbiology 14:3.
ECDC (European Centre for Disease Prevention and Control). 2013. Legionnaires’ disease in Europe, 2011. Stockholm, Sweden: ECDC. http://dx.doi.org/10.2900/78974.
ECDC. 2019. Legionnaires’ disease. In: ECDC. Annual epidemiological report for 2017. Stockholm, Sweden: ECDC.
Edagawa, A., A. Kimura, H. Doi, H. Tanaka, K. Tomioka, K. Sakabe, C. Nakajima, and Y. Suzuki. 2008. Detection of culturable and nonculturable Legionella species from hot water systems of public buildings in Japan. Journal of Applied Microbiology 105:2104-2114.
Edelstein, P. H. 2007. Urine antigen tests positive for Pontiac fever: Implications for diagnosis and pathogenesis. Clinical Infectious Diseases 44(5):229-231.
El-Ebiary, M., X. Sarmiento, and A. Torres. 1997. Prognostic factors of severe Legionella pneumonia requiring admission to ICU. American Journal of Respiratory and Critical Care Medicine 156(5):1467-1472.
Engelhart, S., S. Pleischl, C. Lück, G. Marklein, E. Fischnaller, S. Martin, A. Simon, and M. Exner. 2008. Hospital-acquired legionellosis originating from a cooling tower during a period of thermal inversion. International Journal of Hygiene and Environmental Health 211(3-4):235-240.
Epalle, T., F. Girardot, S. Allegra, C. Maurice-Blanc, O. Garraud, and S. Riffard. 2015. Viable but not culturable forms of Legionella pneumophila generated after heat shock treatment are infectious for macrophage-like and alveolar epithelial cells after resuscitation on Acanthamoeba polyphaga. Microbial Ecology 69(1):215-24.
Euser, S. M., M. Pelgrim, and J. W. den Boer. 2010. Legionnaires’ disease and Pontiac fever after using a private outdoor whirlpool spa. Scandinavian Journal of Infectious Diseases 42:910-916.
Eylert, E., V. Herrmann, M. Jules, N. Gillmaier, M. Lautner, C. Buchrieser, W. Eisenreich, and K. Heuner. 2010. Journal of Biological Chemistry 285(29):22232-22243.
Fallon, R. J., and T. J. Rowbotham. 1990. Microbiological investigations into an outbreak of Pontiac fever due to Legionella micdadei associated with use of a whirlpool. Journal of Clinical Pathology 43(6):479-483.
Farnham, A., L. Alleyne, D. Cimini, and S. Balter. 2014. Legionnaires’ disease incidence and risk factors, New York, New York, USA, 2002–2011. Emerging Infectious Diseases 20(11):1795-1802.
Faucher, S. P., C. A. Mueller, and H. A. Shuman. 2011. Legionella pneumophila transcriptome during intracellular multiplication in human macrophages. Frontiers in Microbiology 2:60.
Faulkner, G., and R. A. Garduño. 2002. Ultrastructural analysis of differentiation in Legionella pneumophila. Journal of Bacteriology 184(24):7025-7041.
Faulkner, G., S. G. Berk, E. Garduño, M. A. Ortiz-Jiménez, and R. A. Garduño. 2008. Passage through Tetrahymena tropicalis triggers a rapid morphological differentiation in Legionella pneumophila. Journal of Bacteriology 190(23):7728-7738.
Fenstersheib, M. D., M. Miller, C. Diggins, S. Liska, L. Detwiler, S. B. Werner, D. Lindquist, W. L. Thacker, and R. F. Benson. 1990. Outbreak of Pontiac fever due to Legionella anisa. Lancet 336(8706):35-37.
Fernandez-Moreira, E., J. H. Helbig, and M. S. Swanson. 2006. Membrane vesicles shed by Legionella pneumophila inhibit fusion of phagosomes with lysosomes. Infection and Immunology 74:3285-3295.
Fernández-Sabé, N., B. Rosón, J. Carratalà, J. Dorca, F. Manresa, and F. Gudiol. 2003. Clinical diagnosis of Legionella pneumonia revisited: Evaluation of the Community-Based Pneumonia Incidence Study Group scoring system. Clinical Infectious Diseases 37(4):483-489.
Fields, B. S., E. B. Shotts, J. C. Feeley, G. W. Gorman, and W. T. Martin. 1984. Proliferation of Legionella pneumophila as an intracellular parasite of the ciliated protozoan Tetrahymena pyriformis. Applied and Environmental Microbiology 47:467-471.
Fields, B. S., G. N. Sanden, J.M. Barbaree, W. E. Morrill, R. M. Wadowsky, E. H. White, and J. C. Feeley. 1989. Intracellular multiplication of Legionella pneumophila in amoebae isolated from hospital hot water tanks. Current Microbiology 18(2):131-137.
Fields, B. S., T. Haupt, J. P. Davis, M. J. Arduino, P. H. Miller, and J. C. Butlet. 2001. Pontiac fever due to Legionella micdadei from a whirlpool spa: Possible role of bacterial endotoxin. Journal of Infectious Diseases 184(10):1289-1292.
Fisher-Hoch, S. P., M. G. Smith, and J. S. Colbourne. 1982. Legionella pneumophila in hospital hot water cylinders. Lancet 319(8280):1073.
Fisman, D. N., S. Lim, G. A. Wellenius., C. Johnson, P. Britz, M. Gaskins, J. Maher, M. A. Mittleman, C. V. Spain, C. N. Haas, and C. Newbern. 2005. It’s not the heat, it’s the humidity: Wet weather increases legionellosis risk in the greater Philadelphia metropolitan area. Journal of Infectious Diseases 192:2066-2073.
Fitzgeorge, R. B., A. S. Featherstone, and A. Baskerville. 1990. Efficacy of azithromycin in the treatment of guinea pigs infected with Legionella pneumophila by aerosol. Journal of Antimicrobial Chemotherapy 25(Suppl A):101-108.
Fiumefreddo, R., R. Zaborsky, J. Haeuptle, M. Christ-Crain, A. Trampuz, I. Steffen, R. Frei, B. Muller, and P. Schuetz. 2009. Clinical predictors for Legionella in patients presenting with community-acquired pneumonia to the emergency department. BMC Pulmonary Medicine 9(4):1-9.
Flemming, H. C., B. Bendinger, M. Exner, J. Gebel, T. Kistemann, G. Schaule, U. Szewzyk, and J. Wingender. 2014. The last meters before the tap: Where drinking water quality is at risk. In: Microbial growth in drinking-water supplies. Problems, causes, control and research needs. D. van der Kooij and P. W. J. J. van der Wielen (eds.). London, UK: IWA Publishing.
Flemming, H. C., J. Wingender, U. Szewzyk, P. Steinberg, S. A. Rice, and S. Kjelleberg. 2016. Biofilms: An emergent form of bacterial life. Nature Reviews in Microbiology 14(9):563-575.
Fliermans, C. B., W. B. Cherry, L. H. Orrison, S. J. Smith, D. L. Tison, and D. H. Pope. 1981. Ecological distribution of Legionella pneumophila. Applied and Environmental Microbiology 41(1):9-16.
Fliermans, C. B., W. B. Cherry, L. H. Orrison, and L. Thacker. 1979. Isolation of Legionella pneumophila from nonepidemic-related aquatic habitats. Applied and Environmental Microbiology 37(6):1239-1242.
Fonseca, M. V., and M. S. Swanson. 2014. Nutrient salvaging and metabolism by the intracellular pathogen Legionella pneumophila. Frontiers in Cellular and Infection Microbiology 4:12.
Francois Watkins, L. K., K. E. Toews, A. M. Harris, S. Davidson, S. Ayers-Millsap, C. E. Lucas, B. C. Hubbard, N. A. Kozak-Muiznieks, E. Khan, E., and P. K. Kutty. 2016. Lessons from an outbreak of Legionnaires’ disease on a hematology-oncology unit. Infection Control Hospital Epidemiology 38(3):306-313.
Franzin, L., C. Scolfaro, D. Cabodi, M. Valera, and P. A. Tovo. 2001. Legionella pneumophila pneumonia in a newborn after water birth: a new mode of transmission. Clinical Infectious Diseases 33(9):e103-e104.
Gaia, V., N. K. Fry, B. Afshar, P. C. Lück, H. Meugnier, J. Etienne, R. Peduzzi, and T. G. Harrison. 2005. Consensus sequence-based scheme for epidemiological typing of clinical and environmental isolates of Legionella pneumophila. Journal of Clinical Microbiology 43(5):2047-2052.
Gao, Z., Y. Kang, J. Yu, and L. Ren. 2014. Human pharyngeal microbiome may play a protective role in respiratory tract infections. Genomics, Proteomics, Bioinformatics 12(3):144-150.
García, M. T., S. Jones, C. Pelaz, R. D. Millar, and Y. Abu Kwaik. 2007. Acanthamoeba polyphaga resuscitates viable non-culturable Legionella pneumophila after disinfection. Environmental Microbiology 9:1267-77.
Garcia-Nunez, M., N. Sopena, S. Ragull, M. L. Pedro-Botet, J. Morera, and M. Sabria. 2008. Persistence of Legionella in hospital water supplies and nosocomial Legionnaires’ disease. FEMS Immunology Medical Microbiology 52(2):202-206.
Garcia-Vidal, C., M. Labori, D. Viasus, A. Simonetti, D. Garcia-Somoza, J. Dorca, F. Gudiol, and J. Carratalà. 2013. Rainfall is a risk factor for sporadic cases of Legionella pneumophila pneumonia. PLoS One 8(4):e61036.
Garcia-Vidal, C., I. Sanchez-Rodriguez, A. F. Simonetti, J. Burgos, D. Viasus, M. T. Martin, V. Falco, and J. Carratalà. 2017. Levofloxacin versus azithromycin for treating Legionella pneumonia: a propensity score analysis. Clinical Microbiology and Infection 23(9):653-658.
Garduño, R. A., E. Garduño, M. Hiltz, and P. S. Hoffman. 2002. Intracellular growth of Legionella pneumophila gives rise to a differentiated form dissimilar to stationary-phase forms. Infection and Immunity 70:6273-6283.
Gargano, J. W., E. A. Adam, S. A. Collier, K. E. Fullerton, S. J. Feinman, and M. J. Beach. 2017. Mortality from selected diseases that can be transmitted by water—United States, 2003–2009. Journal of Water Health 15(3):438-450.
Garrison, K., M. S. Shaw, J. T. McCollum, C. Dexter, P. M. Snippes Vagnone, J. H. Thompson, G. Giambrone, B. White, S. Thomas, L. R. Carpenter, M. Nichols, E. Parker, S. Petit, L. A. Hicks, and G. E. Langley. 2014. On-site availability of Legionella testing in acute care hospitals, United States. Infection Control and Hospital Epidemiology 35(7):898-900.
Garrison, L. E. J. M. Kunz, L. A. Cooley, M. R. Moore, C. Lucas, S. Schrag, J. Sarisky, and C. G. Whitney. 2016. Vital signs: Deficiencies in environmental control identified in outbreaks of Legionnaires’ disease—North America, 2000–2014. Morbidity and Mortality Weekly Report. 65(22):576-584.
Garrity, G. M., J. A. Bell, and T. Lilburn. 2005. Legionellales ord. nov. Pp. 210-247 In: Bergey’s Manual® of Systematic Bacteriology: Volume two the proteobacteria part b the gammaproteobacteria. D. J. Brenner, N. R. Krieg, J. T. Staley, G. M. Garrity, D. R. Boone, P. De Vos, M. Goodfellow, F. A. Rainey, and K.-H. Schleifer (eds.). Boston, MA: Springer.
George, J. R., L. Pine, M. W. Reeves, and W. K. Harrell. 1980. Amino acid requirements of Legionella pneumophila. Journal of Clinical Microbiology 11(3):286-291.
Gião, M. S., S. A. Wilks, and C. W. Keevil. 2015. Influence of copper surfaces on biofilm formation by Legionella pneumophila in potable water. Biometals 28(2):329-339.
Gillmaier, N., E. Schunder, E. Kutzner, H. Tlapák, K. Rydzewski, V. Herrmann, M. Stämmler, P. Lasch, W. Eisenreich, and K. Heuner. 2016. Growth-related metabolism of the carbon storage poly-3-hydroxybutyrate in Legionella pneumophila. Journal of Biological Chemistry 291(12):6471-6482.
Goldoni, P., L. Sinibaldi, P. Valenti, and N. Orsi. 2000. Metal complexes of lactoferrin and their effect on the intracellular multiplication of Legionella pneumophila. Biometals 13(1):15-22.
Gomez-Valero, L., C. Rusniok, M. Rolando, M. Neou, D. Dervins-Ravault, J. Demirtas, Z. Rouy, R. J. Moore, H. Chen, N. K. Petty, S. Jarraud, J. Etienne, M. Steinert, K. Heuner, S. Gribaldo, C. Médigue, G. Glöckner, E. L. Hartland, and C. Buchrieser. 2014. Comparative analyses of Legionella species identifies genetic features of strains causing Legionnaires’ disease. Genome Biology 15(11):505.
Gomez-Valero, L., C. Rusniok, D. Carson, S. Mondino, A. E. Pérez-Cobas, M. Rolando, S. Pasricha, S. Reuter, J. Demirtas, J. Crumbach, S. Descorps-Declere, E. L. Hartland, S. Jarraud, G. Dougan, G. N. Schroeder, G. Frankel, and C. Buchrieser. 2019. More than 18,000 effectors in the Legionella genus genome provide multiple, independent combinations for replication in human cells. Proceedings of the National Academy of Sciences 116(6):2265-2273.
Granseth, G., R. Bhattarai, T. Sylvester, S. Prasai, and E. Livar. 2017. Two cases of Legionnaires’ disease in newborns after water births – Arizona, 2016. Morbidity and Mortality Weekly Report 66(22):590-591.
Greenberg, D., C. C. Chiou, R. Famigilleti, T. C. Lee, and V. L. Yu. 2006. Problem pathogens: Paediatric legionellosis—implications for improved diagnosis. Lancet Infectious Diseases 6(8):529-535.
Griffin, A. T., P. Peyrani, T. Wiemken, and F. Arnold. 2010. Macrolides versus quinolones in Legionella pneumonia: Results from the Community-Acquired Pneumonia Organization international study. International Journal of Tuberculous and Lung Disease 14(4):495-499.
Guimaraes, A. J., K. X. Gomes, J. R. Cortines, J. M. Peralta, and R. H. S. Peralta. 2016. Acanthamoeba spp. as a universal host for pathogenic microorganisms: One bridge from environment to host virulence. Microbiological Research 193:30-38.
Haddrell, A. E., J. F. Davies, R. E. Miles, J. P. Reid, L. A. Dailey, and D. Murnane. 2014. Dynamics of aerosol size during inhalation: Hygroscopic growth of commercial nebulizer formulations. International Journal of Pharmacy 463(1):50-61.
Hägele, S., R. Köhler, H. Merkert, M. Schleicher, J. Hacker, and M. Steinert. 2000. Dictyostelium discoideum: A new host model system for intracellular pathogens of the genus Legionella. Cellular Microbiology 2:165-171.
Haldane, D. J., R. Peppard, and R. K. Sumarah. 1993. Direct immunofluorescence for the diagnosis of legionellosis. Canadian Journal of Infectious Disease 4(2):101-104.
Hambsch, B., J. Ashworth, and D. van der Kooij. 2014. Enhancement of microbial growth by materials in contact with drinking water: Problems and test methods. In: Microbial growth in drinking-water supplies. Problems, causes, control and research needs. D. van der Kooij and P. W. J. J van der Wielen (eds.). London: IWA Publishing.
Hamilton, K. A., and C. N. Haas. 2016. Critical review of mathematical approaches for quantitative microbial risk assessment (QMRA) of Legionella in engineered water systems: Research gaps and a new framework. Environmental Science: Water Research and Technology 2:599-613.
Hamilton, K. A., M. T. Hamilton, W. Johnson, P. Jjemba, Z. Bukhari, M. LeChevallier, C. N. Haas, and P. L. Gurian. 2019. Risk-based critical concentrations of Legionella pneumophila for indoor residential water uses. Environmental Science and Technology https://doi.org/10.1021/acs.est.8b03000.
Hamilton, K. A., A. J. Prussin II, W. Ahmed, and C. N. Haas. 2018. Outbreaks of Legionnaires’ disease and Pontiac fever 2006–2017. Current Environmental Health Reports 5(2):263-271.
Hammer, E. 2018. Temporal and ecological community dynamics of water-cooling tower associated Legionella spp. Masters Theses Clemson University, Clemson, South Carolina. https://tigerprints.clemson.edu/all_theses/2938.
Hammes, F., M. Berney, and T. Egli. 2011. Cultivation-independent assessment of bacterial viability. In: High Resolution Microbial Single Cell Analytics. Advances in Biochemical Engineering/Biotechnology. S. Müller and T. Bley (eds.). Berlin: Springer.
Han, X. Y., A. Ihegword, S. E. Evans, J. Zhang, L. Li, H. Cao, J. J. Tarrand, O. El-Kweifi, and R. Patel. 2015. Microbiological and clinical studies of legionellosis in 33 patients with cancer. Journal of Clinical Microbiology 53(7):2180-2187.
Harada, E., K. Iida, S. Shiota, H. Nakayama, and S. Yoshida. 2010. Glucose metabolism in Legionella pneumophila: Dependence on the Entner-Doudoroff pathway and connection with intracellular bacterial growth. Journal of Bacteriology 192(11):2892-2899.
Harpaz, R., R. M. Dahl, and K. L. Dooling. 2016. Prevalence of immunosuppression among U.S. adults, 2013. Journal of the American Medical Association 316(23):2547-2548.
Harrison, T. G., B. Afshar, N. Doshi, N. K. Fry, and J. V. Lee. 2009. Distribution of Legionella pneumophila serogroups, monoclonal antibody subgroups and DNA sequence types in recent clinical and environmental isolates from England and Wales (2000-2008). European Journal of Clinical Microbiology and Infectious Diseases 28(7):781-791.
Häuslein, I., C. Manske, W. Goebel, W. Eisenreich, and H. Hilbi. 2016. Pathway analysis using 13C-glycerol and other carbon tracers reveals a bipartite metabolism of Legionella pneumophila. Molecular Microbiology 100(2):229-246.
Häuslein, I., T. Sahr, P. Escoll, N. Klausner, W. Eisenreich, and C. Buchrieser. 2017. Legionella pneumophila CsrA regulates a metabolic switch from amino acid to glycerolipid metabolism. Open Biology 7(11). https://doi.org/10.1098/rsob.170149.
Hawn, T. R., A. Verbon, K. D. Lettinga, L. P. Zhao, S. S. Li, R. J. Laws, S. J. Skerrett, B. Beutler, L. Schroeder, A. Nachman, A. Ozinsky, K. D. Smith, and A. Aderem. 2003. A common dominant TLR5 stop codon polymorphism abolishes flagellin signaling and is associated with susceptibility to Legionnaires’ disease. Journal of Experimental Medicine 198(10):1563-1572.
Hawn, T. R., A. Verbon, M. Janer, L. P. Zhao, B. Beutler, and A. Aderem. 2005. Toll-like receptor 4 polymorphisms are associated with resistance to Legionnaires’ disease. Proceedings of the National Academy of Sciences 102(7):2487-2489.
Hayden, R. T., J. R. Uhl, X. Qian, M. K. Hopkins, M. C. Aubry, A. H. Limper, R. V. Lloyd, F. R. Cockerill. 2001. Direct detection of Legionella species from bronchoalveolar lavage and open lung biopsy specimens: comparison of LightCycler PCR, in situ hybridization, direct fluorescence antigen detection, and culture. Journal of Clinical Microbiology 39(7):2618-2626.
Hazel, W., W. L. Thacker, R. F. Benson, S. S. Polt, E. Brookings, W. R. Mayberry, D. J. Brenner, R. G. Gilley, and J. K. Kirklin. 1987. Legionella birminghamensis sp. nov. isolated from a cardiac transplant recipient. Journal of Clinical Microbiology 25(11):2120-2122.
Heath, C. H., D. I. Grove, and D. F. Looke. 1996. Delay in appropriate therapy of Legionella pneumonia associated with increased mortality. European Journal of Clinical Microbiology and Infectious Diseases 15(4):286-290.
Hellinga, J. R., R. A. Garduno, J. D. Kormish, J. R. Tanner, D. Khan, K. Buchko, C. Jimenez, M. M. Pinette, and A. K. Brassinga. 2015. Identification of vacuoles containing extraintestinal differentiated forms of Legionella pneumophila in colonized Caenorhabditis elegans soil nematodes. Microbiology Open 4(4):660-681.
Heriot, W. J., H. G. Mack, and R. Stawell. 2014. Ocular involvement in a patient with Legionella longbeachae 1 infection. Clinical and Experimental Ophthalmology 42(5):497-499.
Herwaldt, L. A., G. W. Gorman, T. McGrath, S. Toma, B. Brake, A. W. Hightower, J. Jones, A. L. Reingold, P. A. Boxer, P. W. Tang, C. W. Moss, H. Wilkinson, D. J. Brenner, A. G. Steigerwalt, and C. V. Broome. 1984. A new Legionella species, Legionella feeleii species nova, causes Pontiac fever in an automobile plant. Annals of Internal Medicine 100(3):333-338.
Hicks, L. A., C. E. Rose, B. S. Fields, M. L. Drees, J. P. Engel, P. R. Jenkins, B. S. Rouse, D. Blythe, A. P. Khalifah, D. R. Feikin, and C. G.Whitney. 2007. Increased rainfall is associated with increased risk for legionellosis. Epidemiology and Infection 135(5):811-817.
Hilbi, H., C. Hoffmann, and C. F. Harrison. 2011. Legionella spp. outdoors: Colonization, communication and persistence. Environmental Microbiology Reports 3(3):286-296.
Hindré, T., H. Brüggemann, C. Buchrieser, and Y. Héchard. 2008. Transcriptional profiling of Legionella pneumophila biofilm cells and the influence of iron on biofilm formation. Microbiology 154(Pt 1):30-41.
Hovel-Miner, G., S. Pampou, S. P. Faucher, M. Clarke, I. Morozova, P. Morozov, J. J. Russo, H. A. Shuman, and S. Kalachikov. 2009. SigmaS controls multiple pathways associated with intracellular multiplication of Legionella pneumophila. Journal of Bacteriology 191(8):2461-73.
Hovel-Miner, G., S. P. Faucher, X. Charpentier, and H. A. Shuman. 2010. ArgR-regulated genes are derepressed in the Legionella-containing vacuole. Journal of Bacteriology 192(17):4504-4516.
Hsu, B. M., C. C. Huang, J. S. Chen, N. H. Chen, and J. T. Huang. 2011. Comparison of potentially pathogenic free-living amoeba hosts by Legionella spp. in substrate-associated biofilms and floating biofilms from spring environments. Water Research 45(16):5171-83.
Htwe, T. H., and N. M. Khardori. 2017. Legionnaires’ disease and immunosuppressive drugs. Infectious Disease Clinics of North America 31(1):29-42.
Hull, N. M., E. P. Holinger, K. A. Ross, C. E. Robertson, J. K. Harris, M. J. Stevens, and N. R. Pace. 2017. Longitudinal and source-to-tap New Orleans, LA, U.S.A. drinking water microbiology. Environmental Science and Technology 51(8):4220-4229.
Huhn, G. D., B. Adam, R. Ruden, L. Hilliard, P. Kirkpatrick, J. Todd, W. Crafts, D. Passaro, and M. S. Dworkin. 2002. Outbreak of travel-related Pontiac fever among hotel guests illustrating the need for better diagnostic tests. Journal of Travel Medicine 12(4):173-179.
Hwang, M. G., H. Katayama, and S. Ohgaki. 2006. Effect of intracellular resuscitation of Legionella pneumophila in Acanthamoeba polyphage cells on the antimicrobial properties of silver and copper. Environmental Science and Technology 40:7434-7439.
ISO (International Organization for Standardization). 1998. Water quality—Detection and enumeration of Legionella. ISO 11731:1998. Geneva, Switzerland: ISO.
Jacobson, K. L., M. H. Miceli, J. J. Tarrand, and D. P. Kontoyiannis. 2008. Legionella pneumonia in cancer patients. Medicine 87(3):152-159.
Jaeger, T. M., P. P. Atkinson, B. A. Adams, A. J. Wright, and R. D. Hurt. 1988. Legionella bozemanii pneumonia in an immunocompromised patient. Mayo Clinic Proceedings 63(1):72-76.
Jain, S., W. H. Self, R. G. Wunderink, S. Fakhran, R. Balk, A. M. Bramley, C. Reed, C. G. Grijalva, E. J. Anderson, D. M. Courtney, J. D. Chappell, and C. Qi, et al., for the CDC EPIC Study Team. 2015. Community-acquired pneumonia requiring hospitalization among U.S. adults. New England Journal of Medicine 373(5):415-427.
James, B. W., W. S. Mauchline, P. J. Dennis, C. W. Keevil, and R. Wait. 1999. Poly-3-hydroxybutyrate in Legionella pneumophila, an energy source for survival in low-nutrient environments. Applied and Environmental Microbiology 65:822-827.
Jaresova, M., I. Hlozanek, I. Striz, K. Petrickova, and Z. Kocmoud. 2006. Legionella detection in oropharyngeal aspirates of transplant patients prior to surgery. European Journal of Clinical Microbiology and Infectious Diseases 25:63-64.
Jespersen, S., O. S. Søgaard, H. C. Schønheyder, M. J. Fine, and L. Ostergaard. 2010. Clinical features and predictors of mortality in admitted patients with community- and hospital-acquired legionellosis: A Danish historical cohort study. BMC Infectious Diseases 10:124. doi:10.1186/1471-2334-10-124.
Ji, P., W. J. Rhoads, M. A. Edwards, and A. Pruden. 2017. Impact of water heater temperature setting and water use frequency on the building plumbing microbiome. ISME Journal 11:1318-1330.
Joly, J. R., M. Boissinot, J. Duchaine, M. Duval, J. Rafrafi, D. Ramsay, and R. Letarte. 1984. Ecological distribution of legionellaceae in the Quebec city area. Canadian Journal of Microbiology 30(1):63-67.
Jonas, D., I. Engels, D. Hartung, J. Beyersmann, U. Frank, and F. D. Daschner. 2003. Development and mechanism of fluoroquinolone resistance in Legionella pneumophila. Journal of Antimicrobial Chemotherapy 51(2):275-280.
Joseph, S. J., D. Cox, B. Wolff, S. S. Morrison, N. A. Kozak-Muiznieks, M. Frace, X. Didelot, S. Castillo-Ramirez, J. Winchell, T. D. Read, and D. Dean. 2016. Dynamics of genome change among Legionella species. Scientific Reports 6:33442.
Karagiannis, I., B. Schimmer, and A. M. de Roda Husman. 2009. Compliance with boil water advice following a water contamination incident in The Netherlands in 2007. Eurosurveillance 14(12): pii=19156.
Kashuba, A. D. M., and C. H. Ballow. 1996. Legionella urinary antigen testing: Potential impact on diagnosis and antibiotic therapy. Diagnostic Microbiology and Infectious Disease 24(3):129-139.
Kebbi-Beghdadi, C., and G. Greub. 2014. Importance of amoebae as a tool to isolate amoeba-resisting microorganisms and for their ecology and evolution: The Chlamydia paradigm. Environmental Microbiology Reports 6(4):309-324.
Khan, M. A., N. Knox, A. Prashar, D. Alexander, M. Abdel-Nour, C. Duncan, P. Tang, H. Amatullah, C. C. Dos Santos, N. Tijet, D. E. Low, C. Pourcel, and G. Van Domselaar. 2013. Comparative genomics reveal that host-innate immune responses influence the clinical prevalence of Legionella pneumophila serogroups. PLoS One 8(6). https://doi.org/10.1371/journal.pone.0067298.
Khodr, A., E. Kay, L. Gomez-Valero, C. Ginevra, P. Doublet, C. Buchrieser, and S. Jarraud. 2016. Molecular epidemiology, phylogeny, and evolution of Legionella. Infection, Genetics, and Evolution 43:108-122.
Kikuchi, R., N. Watabe, T. Konno, N. Mishina, K. Sekizawa, and H. Sasaki. 1994. High incidence of silent aspiration in elderly patients with community-acquired pneumonia. American Journal of Respiratory and Critical Care Medicine 150(1).
Kikuhara, H., M. Ogawa, H. Miyamoto, Y. Nikaido, and S. Yoshida. 1994. Intracellular multiplication of Legionella pneumophila in Tetrahymena thermophila. Journal of UOEH 16:263-275.
Kilborn, J. A., L. A. Manz, M. O’Brien, M. C. Douglas, H. M. Horst, W. Kupin, and E. J. Fisher. 1992. Necrotizing cellulitis caused by Legionella micdadei. American Journal of Medicine 92(1):104-106.
King, C. H., E. B. Shotts, R. E. Wooley, and K. G. Porter. 1988. Survival of coliforms and bacterial pathogens within protozoa during chlorination. Applied and Environmental Microbiology 54:3023-3033.
Kirschner, A. K. T. 2016. Determination of viable legionellae in engineered water systems: Do we find what we are looking for? Water Research 93:276-288.
Knirsch, C. A., K. Jakob, D. Schoonmaker, J. A. Kiehlbauch, S. J. Wong, P. Della-Latta, S. Whittier, M. Layton, and B. Scully. 2000. An outbreak of Legionella micdadei pneumonia in transplant patients: Evaluation, molecular epidemiology, and control. American Journal of Medicine 108(4):290-295.
Knox, N. C., K. A. Weedmark, J. Conly, A. W. Ensminger, F. S. Hosein, S. J. Drews, and the Legionella Outbreak Investigation Team. 2016. Unusual Legionnaires’ outbreak in cool, dry Western Canada: an investigation using genetic epidemiology. Epidemiology and Infection 145(2):254-265.
Kohler, R. B., W. C. Winn, and L. J. Wheat. 1984. Onset and duration of urinary antigen excretion in Legionnaires’ disease. Journal of Clinical Microbiology 20(4):605-607.
Koubar, M., M. H. Rodier, R. A. Garduño, and J. Frere. 2011. Passage through Tetrahymena tropicalis enhances the resistance to stress and the infectivity of Legionella pneumophila. FEMS Microbiology Letters 325(1):10-15.
Kozak, N. A., R. F. Benson, E. Brown, N. T. Alexander, T. H. Taylor, B. G. Shelton, and B. S. Fields. 2009. Distribution of lag-1 alleles and sequence-based types among Legionella pneumophila serogroup 1 clinical and environmental isolates in the United States. Journal of Clinical Microbiology 47(8):2525-2535.
Kozak-Muiznieks, N. A., C. E. Lucas, E. Brown, T. Pondo, T. H. Taylor, Jr., M. Frace, D. Miskowski, and J. M. Winchell. 2014. Prevalence of sequence types among clinical and environmental isolates of Legionella pneumophila serogroup 1 in the United States from 1982 to 2012. Journal of Clinical Microbiology 52(1):201-211.
Kozak-Muiznieks, N. A., S. S. Morrison, S. Sammons, L. A. Rowe, M. Sheth, M. Frace, C. E. Lucas, V. N. Loparev, B. H. Raphael, and J. M. Winchell. 2016. Three genome sequences of Legionella pneumophila subsp. pascullei associated with colonization of a health care facility. Genome Announcements 4(3):e00335-16.
Kruse, E. B., A. Wehner, and H. Wisplinghoff. 2016. Prevalence and distribution of Legionella spp. in potable water systems in Germany, risk factors associated with contamination, and effectiveness of thermal disinfection. American Journal of Infection Control 44(4):470-474.
Kuchta, J. M., J. S. Navratil, M. E. Shepherd, R. M. Wadowsky, J. N. Dowling, S. J. States, and R. B. Yee. 1993. Impact of chlorine and heat on the survival of Hartmannella vermiformis and subsequent growth of Legionella pneumophila. Applied and Environmental Microbiology 59(12):4096-4100.
Kuiper, M. W., B. A. Wullings, A. D. L. Akkermans, R. R. Beumer, and D. van der Kooij. 2004. Intracellular proliferation of Legionella pneumophila in Hartmannella vermiformis in aquatic biofilms grown on plasticized polyvinyl chloride. Applied and Environmental Microbiology 70:6826-6833.
Kusnetsov, J. M., E. Ottoila, and P. J. Martikainen. 1996. Growth, respiration, and survival of Legionella pneumophila at high temperatures. Journal of Applied Bacteriology 81(4):341-347.
Kwaik, Y. A., L.-Y. Gao, O. S. Harb, and B. J. Stone. 1997. Transcriptional regulation of the macrophage induced gene (gspA) of Legionella pneumophila and phenotypic characterization of a null mutant. Molecular Microbiology 24:629-642.
Kyritsi, M. A., V. A. Mouchtouri, A. Katsiafliaka, F. Kolokythopoulou, E. Plakokefalos, V. Nakoulas, G. Rachiotis, and C. Hadjichristodoulou. 2018. Clusters of healthcare-associated Legionnaires’ disease in two hospitals of central Greece. Case Reports in Infectious Diseases 2018, 2570758.
La Scola, B., L. Mezi, P. J. Weiller, and D. Raoult. 2001. Isolation of Legionella anisa using an amoebic coculture procedure. Journal of Clinical Microbiology 39:365-366.
Lamoth, F., and G. Greub. 2010. Amoebal pathogens as emerging causal agents of pneumonia. FEMS Microbiology Reviews 34(3):260-280.
Langley, J. M., S. A. Halperin, F. D. Boucher, B. Smith, and Pediatric Investigators Collaborative Network on Infections in Canada (PICNIC). 2004. Azithromycin is as effective as and better tolerated than erythromycin estolate for the treatment of pertussis. Pediatrics 114(1):e96-e101.
Lanternier, F., F. Tubach, P. Ravaud, D. Salmon, P. Dellamonica, S. Bretagne, M. Couret, B. Bouvard, M. Debandt, I. Gueit, J.-P. Gendre, J. Leone, N. Nicolas, D. Che, X. Mariette, O. Lortholary, and the Research Axed on Tolerance of Biotherapies Group. 2013. Incidence and risk factors of Legionella pneumophila pneumonia during anti-tumor necrosis factor therapy: A prospective French study. Chest 144(3):990-998.
Lanternier, F., F. Ader, B. Pilmis, E. Catherinot, S. Jarraud, and O. Lortholary. 2017. Legionnaires’ disease in compromised hosts. Infectious Disease Clinics of North America 31(1):123-135.
Lau, H. Y., and N. J. Ashbolt. 2009. The role of biofilms and protozoa in Legionella pathogenesis: Implications for drinking water. Journal of Applied Microbiology 107(2):368-378.
Lautenschlager, K., N. Boon, Y. Wang, T. Egli, and F. Hammes. 2010. Overnight stagnation of drinking water in household taps induces microbial growth and changes in community composition. Water Research 44(17):4868-4877.
Learbuch, K. L. G., M. C. Lut, G. Liu, H. Smidt, and P. W. J. J. van der Wielen. 2019. Legionella growth potential of drinking water produced by reverse osmosis. Water Research 157:55-63.
LeChevallier, M. W., N. J. Welch, and D. B. Smith. 1996. Full-scale studies of factors related to coliform regrowth in drinking water. Applied and Environmental Microbiology 62(7):2201-2211.
Lee, A. S., and J. H. Ryu. 2018. Aspiration pneumonia and related syndromes. Mayo Clinic Proceedings 93(6):752-762.
Lee, T. C., R. M. Vickers, V. L. Yu, and M. M. Wagener. 1993. Growth of 28 Legionella species on selective culture media: A comparative study. Journal of Clinical Microbiology 31(10):2764-2768.
Leoni, E., F. Catalani, S. Marini, and L. Dallolio. 2018. Legionellosis associated with recreational waters: A systematic review of cases and outbreaks in swimming pools, spa pools, and similar environments. International Journal of Environmental Research and Public Health 15(1612):doi:10.3390/ijerph15081612.
Levy, I., and L. G. Rubin. 1998. Legionella pneumonia in neonates: A literature review. Journal of Perinatology 18(4):287-290.
Li, L., N. Mendis, H. Trigui, J. D. Oliver, and S. P. Faucher. 2014. The importance of the viable-but-non-culturable state in human bacterial pathogens. Frontiers in Microbiology 5:258.
Lienard, J., A. Croxatto, A. Gervaix, Y. Levi, J. F. Loret, K. M. Posfay-Barbe, and G. Greub. 2017. Prevalence and diversity of Chlamydiales and other amoeba-resisting bacteria in domestic drinking water systems. New Microbes and New Infections 15:107-116.
Llewellyn, A. C., C. E. Lucas, S. E. Roberts, E. W. Brown, B. S. Nayak, B. H. Raphael, and J. M. Winchell. 2017. Distribution of Legionella and bacterial community composition among regionally diverse U.S. cooling towers. PLoS ONE 12(12):e0189937.
Lode, H., B. Kemmerich, H. Schäfer, R. Grothe, R. Hartmann, W. Ehret, G. Ruckdeschel. 1987. Significance of non-pneumophila Legionella species in adult community-acquired and nosocomial pneumonias. Klinische Wochenschrift 65(10):463-468.
Loenenbach, A. D., C. Beulens, S. M. Euser, J. P. G. van Leuken, B. Bom, W. van der Hoek, A. M. de Roda Husman, W. L. M. Ruijs, A. A. Bartels, A. Rietveld, J. W. den Boer, and P. S. Brandsema. 2018. Two community clusters of Legionnaires’ disease directly linked to a biologic wastewater treatment plant, The Netherlands. Emerging Infectious Diseases 24(10):1914-1918.
Loret, J. F., and G. Greub. 2010. Free-living amoebae: Biological by-passes in water treatment. International Journal of Hygiene and Environmental Health 213(3):167-175.
Lu, J., H. Buse, V. Gomez-Alvarez, I. Struewing, J. Santo Domingo, and N. J. Ashbolt. 2014. Impact of drinking water conditions and copper materials on downstream biofilm microbial communities and Legionella pneumophila colonization. Journal of Applied Microbiology 117(3):905-918.
Lu J., I. Struewing, S. Yelton, and N. Ashbolt. 2015. Molecular survey of occurrence and quantity of Legionella spp., Mycobacterium spp., Pseudomonas aeruginosa and amoeba hosts in municipal drinking water storage tank sediments. Journal of Applied Microbiology 119(1):278-288.
Lu, J., I. Struewing, E. Vereen, A. E. Kirby, K. Levy, C. Moe, and N. Ashbolt. 2016. Molecular detection of Legionella spp. and their associations with Mycobacterium spp., Pseudomonas aeruginosa, and amoeba hosts in a drinking water distribution system. Journal of Applied Microbiology 120(2):509-521.
Lucas, C. E., T. H. Taylor, Jr., and B. S. Fields. 2011. Accuracy and precision of Legionella isolation by U.S. laboratories in the ELITE program pilot study. Water Research 45:4428-4436.
MacIntyre, C. R., A. Dyda, C. M. Bui, and A. A. Chughtai. 2018. Rolling epidemic of Legionnaires’ disease outbreaks in small geographic areas. Emerging Microbes and Infections 7(1):36.
Maisa, A., A. Brockmann, F. Renken, C. Lück, S. Pleischl, M. Exner, I. Daniels-Haardt, and A. Jurke. 2015. Epidemiological investigation and case-control study: A Legionnaires’ disease outbreak associated with cooling towers in Warstein, Germany, August–September 2013. Eurosurveillance 20(46). https://doi.org/10.2807/1560-7917.ES.2015.20.46.30064.
Maita, C., M. Matsushita, M. Miyoshi, T. Okubo, S. Nakamura, J. Matsuo, M. Takemura, M. Miyake, H. Nagai, and H. Yamaguchi. 2018. Amoebal endosymbiont Neochlamydia protects host amoebae against Legionella pneumophila infection by preventing Legionella entry. Microbes and Infection 20(4):236-244.
Mandell, L. A., R. G. Wunderink, A. Anzueto, J. G. Bartlett, G. D. Campbel, N. C. Dean, S. F. Dowell, T. M. File, Jr., D. M. Musher, M. S. Niederman, A. Torres, and C. G. Whitney. 2007. Infectious Diseases Society of America/American Thoracic Society consensus guidelines on the management of community-acquired pneumonia in adults. Clinical Infectious Diseases 44:S27-72.
Marrão, G., A. Verissimo, R. G. Bowker, and M. S. da Costa. 1993. Biofilms as major sources of Legionella sp. in hydrothermal areas and their dispersion into streams. FEMS Microbiology Ecology 12:25-33.
Marrie, T. J., D. Haldane, S. MacDonald, K. Clarke, C. Fanning, S. LeFort-Jost, G. Bezanson, and J. Joly. 1991. Control of endemic nosocomial Legionnaires’ disease by using sterile potable water for high risk patients. Epidemiology and Infection 107:591-605.
Marrie, T., P. Green, S. Burbridge, G. Bezanson, S. Neale, P. S. Hoffman, and D. Haldane. 1994. Legionellaceae in the potable water of Nova Scotia hospitals and Halifax residences. Epidemiology and Infection 112(1):143-150.
Marston, B. J., H. B. Lipman, and R. F. Breiman. 1994. Surveillance for Legionnaires’ disease. Risk factors for morbidity and mortality. Archives of Internal Medicine 154:2417-2422.
Martiny, A. C., T. M. Jorgensen, H. J. Albrechtsen, E. Arvin, and S. Molin. 2003. Long-term succession of structure and diversity of a biofilm formed in a model drinking water distribution system. Applied and Environmental Microbiology 69(11):6899-6907.
Mastro, T. D. 1991. Nosocomial Legionnaires’ disease and use of medication nebulizers. Journal of Infectious Diseases 163:667-670.
Mauchline, W. S., R. Araujo, R. Wait, A. B. Dowsett, P. J. Dennis, and C. W. Keevil. 1992. Physiology and morphology of Legionella pneumophila in continuous culture at low oxygen concentration. Microbiology 138:2371-2380.
Mendis, N., P. McBride, J. Saoud, T. Mani, and S. P. Faucher. 2018. The LetA/S two-component system regulates transcriptomic changes that are essential for the culturability of Legionella pneumophila in water. Scientific Reports 8(1):6764.
Mérault, N., C. Rusniok, S. Jarraud, V. Gomez-Valero, C. Cazalet, M. Marin, E. Brachet, P. Aegerter, J. L. Gaillard, J. Etienne, J. L. Herrmann, the DELPH-I Study Group, C. Lawrence, and C. Buchrieser. 2011. Specific real-time PCR for simultaneous detection and identification of Legionella pneumophila serogroup 1 in water and clinical samples. Applied and Environmental Microbiology 77(5):1708-1717.
Mercante, J. W., and J. M. Winchell. 2015. Current and emerging Legionella diagnostics for laboratory and outbreak investigations. Clinical Microbiology Reviews 28(1):95-133.
Mermel, L. A., S. L. Josephson, C. H. Giorgio, J. Dempsey, and S. Parenteau. 1995. Association of Legionnaires’ disease with construction: Contamination of potable water? Infection Control and Hospital Epidemiology 16(2):76-81.
Misch, E. A., A. Verbon, J. M. Prins, S. J. Skerrett, T. R. Hawn. 2013. A TLR6 polymorphism is associated with increased risk of Legionnaires’ disease. Genes and Immunity 14(7):420-426.
Mittal, S., A. P. Singh, M. Gold, A. N. Leung, L. B. Haramati, and D. S. Katz. 2017. Thoracic imaging features of Legionnaires’ disease. Infectious Disease Clinics of North America 31(1):43-54.
Miyashita, N., F. Higa, Y. Aoki, T. Kikuchi, M. Seki, K. Tateda, N. Maki, K. Uchino, K. Ogasawara, H. Kiyota, A. Watanabe. 2017. Clinical presentation of Legionella pneumonia: Evaluation of clinical scoring systems and therapeutic efficacy. Journal of Infection and Chemotherapy 23(11):727-732.
Moore, G., M. Hewitt, D. Stevenson, J. T. Walker, and A. M. Bennett. 2015. Aerosolisation of respirable droplets from a domestic spa pool: The use of MS-2 coliphage and Pseudomonas aeruginosa as markers for Legionella pneumophila. Applied and Environmental Microbiology 81(2):555-561.
Morris, G. K., C. M. Patton, J. C. Feeley, S. E. Johnson, G. Gorman, W. T. Martin, P. Skaliy, G. F. Mallison, B. D. Politi, and D. C. Mackel. 1979. Isolation of the Legionnaires’ disease bacterium from environmental samples. Annals of Internal Medicine 90(4):664-666.
Morris, A., J. M. Beck, P. D. Schloss, T. B. Campbell, K. Crothers, J. L. Curtis, S. C. Flores, A. P. Fontenot, E. Ghedin, L. Huang, K. Jablonski, E. Kleerup, S. V. Lynch, E. Sodergren, H. Twigg, V. B. Young, C. M. Bassis, A. Venkataraman, T. M. Schmidt, G. M. Weinstock, and the Lung HIV Microbiome Project. 2013. Comparison of the respiratory microbiome in healthy nonsmokers and smokers. American Journal of Respiratory and Criticial Care Medicine 187(10):1067-1075.
Muder, R. R., and L. Y. Victor. 2002. Infection due to Legionella species other than L. pneumophila. Clinical Infectious Diseases 35(8):990-998.
Muder, R. R., J. E. Stout, and Y. C. Yee. 1992. Isolation of Legionella pneumophila serogroup 5 from empyema following esophageal perforation: Source of the organism and mode of transmission. Chest 102(5):1601-1603.
Muldoon, R. L., D. L. Jaecker, and H. K. Kiefer. 1981. Legionnaires’ disease in children. Pediatrics 67(3):329-332.
Muñoz, M. J., M. C. Martínez Toldos, G. Yagüe, and M. Segovia. 2009. Evaluation of three immunochromatographic assays for detection of Legionella pneumophila serogroup 1 antigen in urine samples. Revista Española de Quimioterapia 22(4):207-209. http://www.ncbi.nlm.nih.gov/pubmed/20082041.
Murdoch, D. R. 2003. Diagnosis of Legionella infection. Clinical Infectious Diseases 36:64-69.
Murdoch, D. R., R. G. Podmore, T. P. Anderson, K. Barratt, M. J. Maze, K. E. French, S. A. Young, S. T. Chambers, and A. M. Werno. 2013. Impact of routine systematic polymerase chain reaction testing on case finding for Legionnaires’ disease: A pre-post comparison study. Clinical Infectious Diseases 57(9):1275-1281.
Musso, D., and D. Raoult. 1997. Serological cross-reactions between Coxiella burnetii and Legionella micdadei. Clinical and Vaccine Immunology 4(2):208-212.
Mykietiuk, A., J. Carratala, N. Fernandez-Sabe, J. Dorca, R. Verdaguer, F. Manresa, and F. Gudiol. 2005. Clinical outcomes for hospitalized patients with Legionella pneumonia in the antigenuria era: The influence of levofloxacin therapy. Clinical Infectious Diseases 40(6):794-799.
Nadarajah, M., S. Singam, and H. A. Jalil. 1987. Sero-survey for Legionella pneumophila antibodies—Singapore experience. Annals of the Academy of Medicine Singapore 16(4):583-585.
Nahapetian, K., O. Challemel, D. Beurtin, S. Dubrou, P. Gounon, and F. Squinazi. 1991. The intracellular multiplication of Legionella pneumophila in protozoa from hospital plumbing systems. Research in Microbiology 142:677-685.
Nakamura, S., K. Yanagihara, K. Izumikawa, M. Seki, H. Kakeya, Y. Yamamoto, H. Senjyu, A. Saito, and S. Kohno. 2009. The clinical efficacy of fluoroquinolone and macrolide combination therapy compared with single-agent therapy against community-acquired pneumonia caused by Legionella pneumophila. Journal of Infection. 59(3):222-224.
Neil, K., and R. Berkelman. 2008. Increasing incidence of legionellosis in the United States, 1990–2005: Changing epidemiologic trends. Clinical Infectious Diseases 47(5):591-599.
Nescerecka, A., T. Juhna, and F. Hammes. 2016. Behavior and stability of adenosine triphosphate (ATP) during chlorine disinfection. Water Research 101:490-497.
Nevo, O., T. Zusman, M. Rasis, Z. Lifshitz, and G. Segal. 2014. Identification of Legionella pneumophila effectors regulated by the LetAS-RsmYZ-CsrA regulatory cascade, many of which modulate vesicular trafficking. Journal of Bacteriology 196(3):681-692.
Newsome, A. L., R. L. Baker, R. D. Miller, and R. R. Arnold. 1985. Interactions between Naegleria fowleri and Legionella pneumophila. Infection and Immunity 50:449-452.
Newton, H. J., D. K. Y. Ang, I. R. Van Driel, and E. L. Hartland. 2010. Molecular pathogenesis of infections caused by Legionella pneumophila. Clinical Microbiology Reviews 23(2):274-298.
Nguyen, T. M. N., D. Ilef, S. Jarraud, L. Rouil, C. Campese, D. Che, S. Haeghebaert, F. Ganiayre, F. Marcel, J. Etienne, and J. C. Desenclos. 2006. A community-wide outbreak of Legionnaires’ disease linked to industrial cooling towers—How far can contaminated aerosols spread? Journal of Infectious Diseases 193(1):102-111.
Niedeveld C. J., F. M. Pet, and P. L. Meenhorst. 1986. Effect of rubbers and their constituents on proliferation of Legionella pneumophila in naturally contaminated hot water. Lancet 328(8500):180-184.
Nielsen, K., J. M. Bangsborg, and N. Høiby. 2000. Susceptibility of Legionella species to five antibiotics and development of resistance by exposure to erythromycin, ciprofloxacin, and rifampicin. Diagnostic Microbiology and Infectious Disease 36(1):43-48.
Nygård, K., O. Werner-Johansen, S. Rønsen, D. A. Caugant, Ø. Simonsen, A. Kanestrøm, E. Ask, J. Ringstad, R. Ødegård, T. Jensen, T. Krogh, E. A. Høiby, E. Ragnhildstveit, I. S. Aaberge, and P. Aavitsland. 2008. An outbreak of Legionnaires’ disease caused by long-distance spread from an industrial air scrubber in Sarpsborg, Norway. Clinical Infectious Diseases 46(1):61-69.
Ohno, A., N. Kato, K. Yamada, and K. Yamaguchi. 2003. Factors influencing survival of Legionella pneumophila serotype 1 in hot spring water and tap water. Applied and Environmental Microbiology 69(5):2540-2547.
Ohno, A., N. Kato, R. Sakamoto, S. Kimura, and K. Yamaguchi. 2008. Temperature-dependent parasitic relationship between Legionella pneumophila and a free-living amoeba (Acanthamoeba castellanii). Applied and Environmental Microbiology 74:4585-4588.
Okubo, T., M. Matsushita, S. Nakamura, J. Matsuo, H. Nagai, and H. Yamaguchi. 2018. Acanthamoeba S13WT relies on its bacterial endosymbiont to backpack human pathogenic bacteria and resist Legionella infection on solid media. Environmental Microbiology Reports 10(3):344-354.
Oliva, G., T. Sahr, and C. Buchrieser. 2018. The life cycle of L. pneumophila: Cellular differentiation is linked to virulence and metabolism. Frontiers in Cellular and Infection Microbiology 8:3 doi: 10.3389/fcimb.2018.00003.
Ongut, G., A. Yavuz, D. Ogunc, M. Tuncer, F. Ozturk, D. Mutlu, L. Donmez, D. Colak, F. Ersoy, G. Yakupoglu, and M. Gultekin. 2003. Seroprevalence of antibodies to Legionella pneumophila in hemodialysis patients. Transplantation Proceedings 36(1):44-46.
Ortiz-Roque, C. M., and T. C. Hazen. 1987. Abundance and distribution of Legionellaceae in Puerto Rican waters. Applied and Environmental Microbiology 53(9):2231-2236.
Ott, M., P. Messner, J. Heesemann, R. Marre, and J. Hacker. 1991. Temperature-dependent expression of flagella in Legionella. Journal of General Microbiology 137(8):1955-61.
Parthuisot, N., N. J. West, P. Lebaron, and J. Baudart. 2010. High diversity and abundance of Legionella spp. in a pristine river and impact of seasonal and anthropogenic effects. Applied and Environmental Microbiology 76:8201-8210.
Paszko-Kolva, C., M. Shahamat, and R. R. Colwell. 1992. Long-term survival of Legionella pneumophila serogroup 1 under low-nutrient conditions and associated morphological changes. FEMS Microbiology Letters 102:45-55.
Patterson, W. J., D. V. Seal, E. Curran, T. M. Sinclair, and J. C. McLuckie. 1994. Fatal nosocomial Legionnaires’ disease: Relevance of contamination of hospital water supply by temperature-dependent buoyancy-driven flow from spur pipes. Epidemiology and Infection 112(3):513-525.
Pea, F. 2018. Intracellular pharmacokinetics of antibacterials and their clinical implications. Clinical Pharmacokinetics 57(2):177-189.
Pearce, M. M., N. Theodoropoulos, G. A. Noskin, J. P. Flaherty, M. E. Stemper, T. Aspeslet, N. P. Cianciotto, and K. D. Reed. 2011. Native valve endocarditis due to a novel strain of Legionella. Journal of Clinical Microbiology 49(9):3340-3342.
Pedro-Botet, M. L., and V. L. Yu. 2009. Treatment strategies for Legionella infection. Expert Opinion on Pharmacotherapy 10(7):1109-1121.
Pedro-Botet, M. L., M. Sabria-Leal, N. Sopena, J. M. Manterola, J. Morera, R. Blavia, E. Padilla, L. Matas and J. M. Gimeno. 1998. Role of immunosuppression in the evolution of Legionnaires’ disease. Clinical Infectious Diseases 26(1):14-19.
Peter, A., and E. Routledge. 2018. Present-day monitoring underestimates the risk of exposure to pathogenic bacteria from cold water storage tanks. PLoS ONE 13(4):e0195635.
Phin, N., F. Parry-Ford, T. Harrison, H. R. Stagg, N. Zhang, K. Kumar, O. Lortholary, A. Zumla, and I. Abubakar. 2014. Epidemiology and clinical management of Legionnaires’ disease. Lancet Infectious Diseases 14(10):1011-1021.
Piao, Z., C. C. Sze, O. Barysheva, K. Iida, and S. Yoshida. 2006. Temperature-regulated formation of mycelial mat-like biofilms by Legionella pneumophila. Applied and Environmental Microbiology 72:1613-1622.
Pierre, D. M., J. Baron, V. L. Yu, and J. E. Stout. 2017. Diagnostic testing for Legionnaires’ disease. Annals of Clinical Microbiology and Antimicrobials 16(1):1-4.
Plouffe, J. F., T. M. File, R. F. Breiman, B. A. Hackman, S. J. Salstrom, B. J. Marston, B. S. Fields, and the Community Based Pneumonia Incidence Study Group. 1995. Reevaluation of the definition of Legionnaires’ disease: use of the urinary antigen assay. Community-Based Pneumonia Incidence Study Group. Clinical Infectious Diseases 20(5):1286-1291.
Plouffe, J. F., R.F. Breiman, B. S. Fields, M. Herbert, J. Inverso, C. Knirsch, A. Kolokathis, T. J. Marrie, L. Nicolle and D. B. Schwartz. 2003. Azithromycin in the treatment of Legionella pneumonia requiring hospitalization. Clinical Infectious Diseases 37(11):1475-1480.
Plutzer, J., and P. Karanis. 2016. Neglected waterborne parasitic protozoa and their detection in water. Water Research 101:318-332.
Poirier, R., J. Rodrigue, J. Villeneuve, and Y. Lacasse. 2017. Early radiographic and tomographic manifestations of Legionnaires’ disease. Canadian Association of Radiologists Journal 68(3):328-333.
Prasad, B., K. A. Hamilton, and C. N. Haas. 2017. Incorporating time-dose-response into Legionella outbreak models. Risk Analysis 37:291-304.
Prashar, A., S. Bhatia, Z. Tabatabaeiyazdi, C. Duncan, R. A. Garduño, P. Tang, D. E. Low, C. Guyard, and M. R. Terebiznik. 2012. Mechanism of invasion of lung epithelial cells by filamentous Legionella pneumophila. Cellular Microbiology 14:1632-1655.
Prashar, A., S. Bhatia, D. Gigliozzi, T. Martin, C. Duncan, C. Guyard, and M. R. Terebiznik. 2013. Filamentous morphology of bacteria delays the timing of phagosome morphogenesis in macrophages. Journal of Cell Biology 203:1081-1097.
Principe, L., P. Tomao, and P. Visca. 2017. Legionellosis in the occupational setting. Environmental Research 152:485-495.
Pringler, N., P. Brydov, and S. A. Uldum. 2002. Occurrence of Legionella in Danish hot water systems. In: Legionella. N. Cianciotto, Y. Kwaik, P. Edelstein, B. Fields, D. Geary, T. Harrison, C. Joseph, R. Ratcliff, J. Stout, and M. Swanson (eds.). Washington, DC: ASM Press.
Proctor, C. R., D. Dai, M. A. Edwards, and A. Pruden. 2017. Interactive effects of temperature, organic carbon, and pipe material on microbiota composition and Legionella pneumophila in hot water plumbing systems. Microbiome 5(1):130.
Proctor, C. R., M. Reimann, B. Vriens, and F. Hammes. 2018. Biofilms in shower hoses. Water Research 131:274-286.
Qin, X., P. M. Abe, S. J. Weissman, and S. C. Manning. 2002. Extrapulmonary Legionella micdadei infection in a previously healthy child. Pediatric Infectious Disease Journal 21(12):1174-1176.
Qin, T., G. Yan, H. Ren, H. Zhou, H. Wang, Y. Xu, M. Zhao, H. Guan, M. Li, and Z. Shao. 2013. High prevalence, genetic diversity, and intracellular growth ability of Legionella in hot spring environments. PLoS ONE 8(3):e59018.
Qin, T., H. Zhou, H. Ren, H. Guan, M. Li, B. Zhu, and Z. Shao. 2014. Distribution of sequence-based types of Legionella pneumophila serogroup 1 strains isolated from cooling towers, hot springs, and potable water systems in China. Applied and Environmental Microbiology 80(7):2150-2157.
Rasch, J., S. Krüger, D. Fontvieille, C. M. Ünal, R. Michel, A. Labrosse, and M. Steinert. 2016. Legionella-protozoa-nematode interactions in aquatic biofilms and influence of Mip on Caenorhabditis elegans colonization. International Journal of Medical Microbiology 306:443-451.
Ratzow, S., V. Gaia, J. H. Helbig, N. K. Fry, and P. C. Luck. 2007. Addition of neuA, the gene encoding N-acylneuraminate cytidylyl transferase, increases the discriminatory ability of the consensus sequence-based scheme for typing Legionella pneumophila serogroup 1 strains. Journal of Clinical Microbiology 45(6):1965-1968.
Reeves, M. W., L. Pine, S. H. Hutner, J. R. George, W. K. Harrell. 1981. Metal requirements of Legionella pneumophila. Journal of Clinical Microbiology 13:688-695.
Reller, L. B., M. P. Weinstein, and D. R. Murdoch. 2003. Diagnosis of Legionella infection. Clinical Infectious Diseases 36(1):64-69.
Remen, T., L. Mathieu, A. Hautemaniere, M. Deloge-Abarkan, P. Hartemann, and D. Zmirou-Navier. 2011. Pontiac fever among retirement home nurses associated with airborne Legionella. Journal of Hospital Infection 78:269-273.
Rhoads, W. J., E. D. Garner, P. Ji, N. Zhu, J. Parks, D. O. Schwake, A. Pruden, and M. A. Edwards. 2017a. Distribution system operational deficiencies coincide with reported Legionnaires’ disease clusters in Flint, MI. Environmental Science and Technology 51(20):11986-11995.
Rhoads, W. J., A. Pruden, and M. A. Edwards. 2017b. Interactive effects of corrosion, copper, and chloramines on Legionella and mycobacteria in hot water plumbing. Environmental Science and Technology 51(12):7065-7075.
Ricci, M. L., A. Grottola, G. Fregni Serpini, A. Bella, M. C. Rota, F. Frascaro, E. Pegoraro, M. Meacci, A. Fabio, E. Vecchi, A. Girolamo, F. Rumpianesi, M. Pecorari, and M. Scaturro. 2018. Improvement of Legionnaires’ disease diagnosis using real-time PCR assay: A retrospective analysis, Italy, 2010 to 2015. Eurosurveillance 23(50). doi:10.2807/1560-7917.
Ricketts, K., A. Charlett, D. Gelb, C. Lane, J. Lee, and C. Joseph. 2018. Weather patterns and Legionnaires’ disease: A meteorological study. Epidemiology and Infection 137(September 2009):1003-1012.
Ridenour, D. A., S. L. Cirillo, S. Feng, M. M. Samrakandi, and J. D. Cirillo. 2003. Identification of a gene that affects the efficiency of host cell infection by Legionella pneumophila in a temperature-dependent fashion. Infection and Immunity 71(11):6256-6263.
Riffard, S., S. Douglass, T. Brooks, S. Springthorpe, L. G. Filion, S. A. Sattar. 2001. Occurrence of Legionella in groundwater: an ecological study. Water Science and Technology 43(12):99-102.
Ristroph, J. D., K. W. Hedlund, and S. Gowda. 1981. Chemically defined medium for Legionella pneumophila growth. Journal of Clinical Microbiology 13(1):115-119.
Robertson, P., H. Abdelhady, and R. A. Garduño. 2014. The many forms of a pleomorphic bacterial pathogen-the developmental network of Legionella pneumophila. Frontiers in Microbiology 5:670.
Rogers, J., A. B. Dowsett, P. J. Dennis, J. V. Lee, and C. W. Keevil. 1994a. Influence of temperature and plumbing material selection on biofilm formation and growth of Legionella pneumophila in a model potable water system containing complex microbial flora. Applied and Environmental Microbiology 60:1585-1592.
Rogers, J., A. B. Dowsett, P. J. Dennis, J. V. Lee, and C. W. Keevil. 1994b. Influence of plumbing materials on biofilm formation and growth of Legionella pneumophila in potable water systems. Applied and Environmental Microbiology 60:1842-1851.
Rohr, U., S. Weber, R. Michel, F. Selenka, and M. Wilhelm. 1998. Comparison of free-living amoebae in hot water systems of hospitals with isolates from moist sanitary areas by identifying genera and determining temperature tolerance. Applied and Environmental Microbiology 64(5):1822-1824.
Rowbotham, T. J. 1980. Preliminary report on the pathogenicity of Legionella pneumophila for freshwater and soil amoebae. Journal of Clinical Pathology 33:1179.
Rowbotham, T. J. 1986. Current views on the relationships between amoebae, legionellae, and man. Israel Journal of Medical Sciences 22:678-689.
Rucinski, S. L., M. P. Murphy, K. D. Kies, S. A. Cunnignham, A. N. Schuetz, and R. Patel. 2018. Eight years of clinical Legionella PCR testing illustrate seasonal pattern. Journal of Infectious Diseases 218(4):669-670.
Rudbeck, M., K. Mølbak, and S. Uldum. 2008. High prevalence of antibodies to Legionella spp. in Danish blood donors: A study in areas with high and average incidence of Legionnaires’ disease. Epidemiology and Infection 136(2):257-262.
Ruiz-Moreno, J. S., L. Hamann, J. A. Shah, A. Verbon, F. P. Mockenhaupt, M. Puzianowska-Kuznicka, J. Naujoks, L. E. Sander, M. Witzenrath, J. C. Cambier, N. Suttorp, R. R. Schumann, L. Jin, T. R. Hawn, B. Opitz, and the CAPNETZ Study Group. 2018. The common HAQ STING variant impairs cGAS-dependent antibacterial responses and is associated with susceptibility to Legionnaires’ disease in humans. PLoS Pathogens 14(1):1-22.
Sahr, T., C. Rusniok, F. Impens, G. Oliva, O. Sismeiro, J. Y. Coppée, and C. Buchrieser. 2017. The Legionella pneumophila genome evolved to accommodate multiple regulatory mechanisms controlled by the CsrA-system. PLoS Genetics 13(2):e1006629.
Sanchez-Buso, L., I. Comas, G. Jorques, and F. Gonzalez-Candelas. 2014. Recombination drives genome evolution in outbreak-related Legionella pneumophila isolates. Nature Genetics 46(11):1205-1211.
Sanden, G. N., W. E. Morrill, B. S. Fields, R. F. Breiman, and J. M. Barbaree. 1992. Incubation of water samples containing amoebae improves detection of legionellae by the culture method. Applied and Environmental Microbiology 58:2001-2004.
Sauer, J.-D., M. A. Bachman, and M. S. Swanson. 2005. The phagosomal transporter A couples threonine acquisition to differentiation and replication of Legionella pneumophila in macrophages. Proceedings of the National Academy of Sciences 102:9924-9929.
Schalk, J. A., A. E. Docters van Leeuwen, W. J. Lodder, H. de Man, S. Euser, J. W. den Boer, and A. M. de Roda Husman. 2012. Isolation of Legionella pneumophila from pluvial floods by amoebal coculture. Applied and Environmental Microbiology 78:4519-4521.
Scheikl, U., H. F. Tsao, M. Horn, A. Indra, and J. Walochnik. 2016. Free-living amoebae and their associated bacteria in Austrian cooling towers: a 1-year routine screening. Parasitology Research 115(9):3365-3374.
Schmitz-Esser, S., P. Tischler, R. Arnold, J. Montanaro, M. Wagner, T. Rattei, and M. Horn. 2010. The genome of the amoeba symbiont Candidatus Amoebophilus asiaticus reveals common mechanisms for host cell interaction among amoeba-associated bacteria. Journal of Bacteriology 192(4):1045-1057.
Schoen, M. E., and N. J. Ashbolt. 2011. An in-premise model for Legionella exposure during showering events. Water Research 45:5826-5836.
Schoenen, D., R. Schulze-Robbecke, and N. Schirdewahn. 1988. Microbial contamination of water by pipe and tubing material. 2. Growth of Legionella pneumophila. Zentralblatt fur Bakteriologie, Mikrobiologie und Hygiene Serie B, Umwelthygiene, Krankenhaushygiene, Arbeitshygiene, praventive Medizin 186:326-332.
Schrammel, B., S. Cervero-Arago, E. Dietersdorfer, J. Walochnik, C. Luck, R. Sommer, and A. Kirschner. 2018. Differential development of Legionella sub-populations during short- and long-term starvation. Water Research 141:417-427.
Schulze-Robbecke, R., M. Rodder, and M. Exner. 1987. Multiplication and killing temperatures of naturally occurring Legionellas. Zentralblatt fur Bakteriologie, Mikrobiologie und Hygiene Serie B, Umwelthygiene, Krankenhaushygiene, Arbeitshygiene, praventive Medizin 184:495-500.
Schunder, E., N. Gillmaier, E. Kutzner, W. Eisenreich, V. Herrmann, M. Lautner, and K. Heuner. 2014. Amino acid uptake and metabolism of Legionella pneumophila hosted by Acanthamoeba castellanii. Journal of Biological Chemistry 289(30):21040-21054.
Scola, B. L., R. J. Birtles, G. Greub, T. J. Harrison, R. M. Ratcliff, and D. Raoult. 2004. Legionella drancourtii sp. nov., a strictly intracellular amoebal pathogen. International Journal of Systematic and Evolutionary Microbiology 54:699-703.
Shadrach, W. S., K. Rydzewski, U. Laube, G. Holland, M. Özel, A.F. Kiderlen, and A. Flieger. 2005. Balamuthia mandrillaris, free-living ameba and opportunistic agent of encephalitis, is a potential host for Legionella pneumophila bacteria. Applied and Environmental Microbiology 71:2244-2249.
Shaheen, M., and N. J. Ashbolt. 2018. Free-living amoebae supporting intracellular growth may produce vesicle-bound respirable doses of Legionella within drinking water systems. Exposure and Health 10(3):201-209.
Shaheen, M., C. Scott, and N. J. Ashbolt. 2019. Long-term persistence of infectious Legionella with free-living amoebae in drinking water biofilms. International Journal of Hygiene and Environmental Health 222:678-686.
Sharma, L., A. Losier, T. Tolbert, C. S. Dela Cruz, and C.R. Marion. 2017. Atypical pneumonia: Updates on Legionella, Chlamydophila, and Mycoplasma pneumonia. Clinics in Chest Medicine 38(1):45-58.
She, R. C., E. Billetdeaux, A.R. Phansalkar, and C. A. Petti. 2007. Limited applicability of direct fluorescent-antibody testing for Bordetella sp. and Legionella sp. specimens for the clinical microbiology laboratory. Journal of Clinical Microbiology 45(7):2212-2214.
Sheehan, K. B., J. M. Henson, and M. J. Ferris. 2005. Legionella species diversity in an acidic biofilm community in Yellowstone National Park. Applied and Environmental Microbiology 71(1):507-511.
Shu, L., D. A. Brock, K. S. Geist, J. W. Miller, D. C. Queller, J. E. Strassmann, and S. DiSalvo. 2018. Symbiont location, host fitness, and possible coadaptation in a symbiosis between social amoebae and bacteria. eLife 7:e42660.
Simmering, J. E., L. A. Polgreen, D. B. Hornick, D. K. Sewell, and P. M. Polgreen. 2017. Weather-dependent risk for Legionnaires’ disease, United States. Emerging Infectious Diseases 23(11):1843–1851.
Simonsen, Ø., E. Wedege, A. Kanestrøm, K. Bolstad, I. S. Aaberge, E. Ragnhildstveit, and J. Ringstad. 2015. Characterization of the extent of a large outbreak of Legionnaires’ disease by serological assays. BMC Infectious Diseases 15:163. doi:10.1186/s12879-015-0903-2.
Singh, N., J. E. Stout, and V.L. Yu. 2004. Prevention of Legionnaires’ disease in transplant recipients: Recommendations for a standardized approach. Transplant Infectious Disease 6(2):58-62.
Sivagnanam, S., and S. A. Pergam. 2016. Legionellosis in transplantation. Current Infectious Disease Reports 18(3):9. doi:10.1007/s11908-016-0517-x.
Sivagnanam, S., S. Podczervinski, S. M. Butler-Wu, V. Hawkins, Z. Stednick, L. A. Helbert, W. A. Glover, E. Whimbey, J. Duchin, G.-S. Cheng, and S. A. Pergam. 2017. Legionnaires’ disease in transplant recipients: A 15-year retrospective study in a tertiary referral center. Transplant Infectious Disease 19(5):1-8.
Skaliy P., and H. McEachern. 1979. Survival of the Legionnaires’ disease bacterium in water. Annals of Internal Medicine 90:662-663.
Soda, E. A., A. E. Barskey, P. P. Shah S. Schrag, C. G. Whitney, M. J. Arduino, S. C. Reddy, J. M. Kunz, C. M. Hunter, B. H. Raphael, and L. A. Cooley. 2017. Vital signs: Health care–associated Legionnaires’ disease surveillance data from 20 states and a large metropolitan area—United States, 2015. Morbidity and Mortality Weekly Report 66:584–589.
Söderberg, M. A., O. Rossier, and N. P. Cianciotto. 2004. The type II protein secretion system of Legionella pneumophila promotes growth at low temperatures. Journal of Bacteriology 186(12):3712-3720.
Sopena, N., L. Pedro-Botet, L. Mateu, G. Tolschinsky, C. Rey-Joly, and M. Sabrià. 2007. Community-acquired Legionella pneumonia in elderly patients: Characteristics and outcome. Journal of the American Geriatrics Society 55(1):114-119.
St-Martin, G., S. Uldum, and K. Mølbak. 2013. Incidence and prognostic factors for Legionnaires’ disease in Denmark 1993–2006. ISRN Epidemiology Volume 2013, Article ID 847283, 8 pages. http://dx.doi.org/10.5402/2013/847283.
States, S. J., L. F. Conley, M. Ceraso, T. E. Stephenson, R. S. Wolford, R. M. Wadowsky, A. M. McNamara, and R. B. Yee. 1985. Effects of metals on Legionella pneumophila growth in drinking water plumbing systems. Applied and Environmental Microbiology 50:1149-1154.
Steinert, M., L. Emody, R. Amann, and J. Hacker. 1997. Resuscitation of viable but nonculturable Legionella pneumophila Philadelphia JR32 by Acanthamoeba castellanii. Applied and Environmental Microbiology 63:2047-2053.
Stewart, C. R., O. Rossier, and N. P. Cianciotto. 2009. Surface translocation by Legionella pneumophila: a form of sliding motility that is dependent upon type II protein secretion. Journal of Bacteriology 191(5):1537-1546.
Storey, M. V., T. A. Stenström, and N. J. Ashbolt. 2004a. Biofilms, thermophilic amoebae and legionellae—A quantitative risk assessment for distributed water. Water Science and Technology 50(1):77-82.
Storey, M. V., J. Winiecka-Krusnell, N. J. Ashbolt, and T. A. Stenström. 2004b. The efficacy of heat and chlorine treatment against thermotolerant acanthamoebae and legionellae. Scandinavian Journal of Infectious Diseases 36(9):656-662.
Storey, M. V., C. E. Kaucner, M. L. Angles, J. R. Blackbeard, and N. J. Ashbolt. 2008. Opportunistic pathogens in drinking and recycled water distribution systems. Water, Australian Water Association 35(1):38-45.
Stout, J. E., and V. L. Yu. 2003. Hospital-acquired Legionnaires’ disease: New developments. Current Opinion in Infectious Diseases 16(4):337-341.
Stout, J. E., M. G. Best, and V. L. Yu. 1986. Susceptibility of members of the family Legionellaceae to thermal stress: Implications for heat eradication methods in water distribution systems. Applied and Environmental Microbiology 52(2):396-399.
Stout, J. E., C. Brennen, and R. R. Muder. 2000. Legionnaires’ disease in a newly constructed long-term care facility. Journal of the American Geriatrics Society 48(12):1589-1592.
Swanson, M., G. Reguera, M. Schaechter, and F. C. Neidhardt. 2016. Microbe. Washington, DC: ASM Press.
Temmerman, R., H. Vervaeren, B. Noseda, N. Boon, and W. Verstraete. 2006. Necrotrophic growth of Legionella pneumophila. Applied and Environmental Microbiology 72(6):4323–4328.
Tesauro, M., F. Petrelli, A. Lizioli, F. Pregliasco, C. Masia, G. Cossellu, G. Farronato, M. Consonni, and F. Sisto. 2018. Presence of Legionella spp. in human dental plaque. Annali di Igiene 30(5):387-390.
Thornley, C. N., D. J. Harte, R. P. Weir, L. J. Allen, K. J. Knightbridge, and P. R. T. Wood. 2017. Legionella longbeachae detected in an industrial cooling tower linked to a legionellosis outbreak, New Zealand, 2015; possible waterborne transmission? Epidemiology and Infection 145(11):2382-2389.
Tijet, N., P. Tang, M. Romilowych, C. Duncan, V. Ng, D. N. Fisman, et al. 2010. New endemic Legionella pneumophila serogroup I clones, Ontario, Canada. Emerging Infectious Diseases 16(3):447-454.
Tison, D. L., D. H. Pope, W. B. Cherry, and C. B. Fliermans. 1980. Growth of Legionella pneumophila in association with blue-green algae (cyanobacteria). Applied and Environmental Microbiology 39:456-459.
Tobin, J. O., C. L. Bartlett, S. A. Waitkins, G. I. Barrow, A. D. Macrae, A. G. Taylor, R. J. Fallon, and F. R. Lynch. 1981. Legionnaires’ disease: Further evidence to implicate water storage and distribution systems as sources. British Medical Journal 282(6263):573.
Torre, I., R. Alfano, T. Borriello, O. De Giglio, C. Iervolino, M. T. Montagna, M. S. Scamardo, and F. Pennino. 2018. Environmental surveillance and in vitro activity of antimicrobial agents against Legionella pneumophila isolated from hospital water systems in Campania, South Italy: A 5-year study. Environmental Research 164(April):574-579.
Tossa, P., M. Deloge-Abarkan, D. Zmirou-Navier, P. Hartemann, and L. Mathieu. 2006. Pontiac fever: An operational definition for epidemiological studies. BMC Public Health 6:1-10.
Travis, T. C., E. W. Brown, L. F. Peruski, D. Siludjai, P. Jorakate, P. Salika, G. Yang, N. A. Kozak, M. Kodani, A. K. Warner, C. E. Lucas, K. A. Thurman, J. M. Winchell, S. Thamthitiwat, and B. S. Fields. 2012. Survey of Legionella species found in Thai soil. International Journal of Microbiology 2012, Article ID 218791. doi:10.1155/2012/218791.
Trigui, H., P. Dudyk, J. Oh, J. I. Hong, and S. P. Faucher. 2015. A regulatory feedback loop between RpoS and SpoT supports the survival of Legionella pneumophila in water. Applied and Environmental Microbiology 81(3):918-28.
Tyml, T., K. Skulinova, J. Kavan, O. Ditrich, M. Kostka, and I. Dykova. 2016. Heterolobosean amoebae from Arctic and Antarctic extremes: 18 novel strains of Allovahlkampfia, Vahlkampfia and Naegleria. European Journal of Protistology 56:119-133.
Tyndall, R. L., and E. L. Domingue. 1982. Co-cultivation of Legionella pneumophila and free-living amoebae. Applied and Environmental Microbiology 44:954-959.
Underwood, A. P., W. Bellamy, B. Afshar, N. K. Fry, and T. G. Harrison. 2006. Development of an online tool for European working group for Legionella infections sequence-based typing, including automatic quality assessment and data submission. Pp. 163-166 In: Legionella. N. Cianciotto, Y. Kwaik, P. Edelstein, B. Fields, D. Geary, T. Harrison, C. Joseph, R. Ratcliff, J. Stout, and M. Swanson (eds.). Washington, DC: ASM Press.
Vaccaro, L., F. Izquierdo, A. Magnet, C. Hurtado, M. A. Salinas, T. Santos Gomes, S. Angulo, S. Salso, J. Pelaez, M. I. Tejeda, A. Alhambra, C. Gómez, A. Enríquez, E. Estirado, S. Fenoy, and C. del Aguila. 2016. First case of Legionnaire’s disease caused by Legionella anisa in Spain and the limitations on the diagnosis of Legionella non-pneumophila infections. PLoS ONE 11(9):e016293.
Valcin¸ a, O., D. Pūle, I. Lucenko, D. Krastin¸ a, Ž. Šteingolde, A. Krūmin¸ a, and A. Bērzin¸ š. 2015. Legionella pneumophila seropositivity-associated factors in Latvian blood donors. International Journal of Environmental Research and Public Health 13(1):ijerph13010058.
Valster, R. M., B. A. Wullings, and D. van der Kooij. 2010. Detection of protozoan hosts for Legionella pneumophila in engineered water systems by using a biofilm batch test. Applied and Environmental Microbiology 76:7144-7153.
Valster, R. M. 2011. Free-living protozoa in drinking water supplies. PhD thesis Wageningen University.
Vandenesch, F., M. Surgot, N. Bornstein, J. C. Paucod, D. Marmet, P. Isoard, and J. Fleurette. 1990. Relationship between free amoeba and Legionella: studies in vitro and in vivo. Zentralblatt für Bakteriologie 272:265-275.
van der Kooij, D., H. R. Veenendaal, N. P. G. Slaats, and D. Vonk. 2002. Biofilm formation and multiplication of Legionella on synthetic pipe materials in contact with treated water under static and dynamic conditions. Pp. 176-180 In: Legionella. R. Marre, Y. Abu Kwaik, C. Bartlett, N. P. Cianciotto, B. S. Fields, M. Frosch, J. Hacker, and P. C. Luck (eds.). Washington, DC: ASM Press.
van der Kooij, D., H. R. Veenendaal, and W. J. Scheffer. 2005. Biofilm formation and multiplication of Legionella in a model warm water system with pipes of copper, stainless steel, and cross-linked polyethylene. Water Research 39:2789-2798.
van der Kooij, D. 2014. Legionella in drinking-water supplies. Pp. 127-175 In: Microbial growth in drinking water supplies. Problems, causes, controls and research needs. D. Van der Kooij and P. W. J. J. van der Wielen (eds.). London, UK: IWA Publishing.
van der Kooij, D., A. J. Brouwer-Hanzens, H. R. Veenendaal, and B. A. Wullings. 2016. Multiplication of Legionella pneumophila sequence types 1, 47, and 62 in buffered yeast extract broth and biofilms exposed to flowing tap water at temperatures of 38°C to 42°C. Applied and Environmental Microbiology 82:6691-6700.
van der Kooij, D., G. L. Bakker, R. Italiaander, H. R. Veenendaal, and B. A. Wullings. 2017. Biofilm composition and threshold concentration for growth of Legionella pneumophila on surfaces exposed to flowing warm tap water without disinfectant. Applied and Environmental Microbiology 83(5):e02737-16.
van der Kooij, D., H. R. Veenendaal, R. Italiaander, E. J. van der Mark, and M. Dignum. 2018. Primary colonizing Betaproteobacteriales play a key role in the growth of Legionella pneumophila in biofilms on surfaces exposed to drinking water treated by slow sand filtration. Applied and Environmental Microbiology 84(24):e01732-18.
van der Lugt, W., S. M. Euser, J. P. Bruin, J. W. den Boer, J. T. Walker, and S. Crespi. 2017. Growth of Legionella anisa in a model drinking water system to evaluate different shower outlets and the impact of cast iron rust. International Journal of Hygiene and Environmental Health 220(8):1295-1308.
Vandewalle-Capo, M., C. Massip, G. Descours, J. Charavit, J. Chastang, P. A. Billy, S. Boisset, G. Lina, C. Gilbert, M. Maurin, S. Jarraud, and C. Ginevra. 2017. Minimum inhibitory concentration (MIC) distribution among wild-type strains of Legionella pneumophila identifies a subpopulation with reduced susceptibility to macrolides owing to efflux pump genes. International Journal of Antimicrobial Agents 50(5):684-689.
van Heijnsbergen, E., J. A. C. Schalk, S. M. Euser, P. S. Brandsema, J. W. den Boer, and A. M. de Roda Husman. 2015. Confirmed and potential sources of Legionella reviewed. Environmental Science and Technology 49:4797-4815.
van Heijnsbergen, E., A. van Deursen, M. Bouwknegt, J. P. Bruin, A. M. de Roda Husman, and J. A. C. Schalk. 2016. Presence and persistence of viable, clinically relevant Legionella pneumophila bacteria in garden soil in The Netherlands. Applied and Environmental Microbiology 82:5125-5131.
van Hoof, J., L. M. Hornstra, E. van der Blom, O. W. Nuijten, and P. van der Wielen. 2014. The presence and growth of Legionella species in thermostatic shower mixer taps: an exploratory field study. Building Services Engineering Research and Technology 35(6):600-612.
Varner, T. R., P. B. Bookstaver, C. N. Rudisill, and H. Albrecht. 2011. Role of rifampin-based combination therapy for severe community-acquired Legionella pneumophila pneumonia. Annals of Pharmacotherapy 45(7-8):967-976.
Vekens, E., O. Soetens, R. De Mendonca, F. Echahidi, S. Roisin, A. Deplano, L. Eeckhout, W. Achtergael, D. Piérard, O. Denis, and I. Wybo. 2012. Sequence-based typing of Legionella pneumophila serogroup 1 clinical isolates from Belgium between 2000 and 2010. Eurosurveillance 17(43):9-14.
Venezia, R. A., M. D. Agresta, E. M. Hanley, K. Urquhart, and D. Schoonmaker. 1994. Nosocomial legionellosis associated with aspiration of nasogastric feedings diluted in tap water. Infection Control and Hospital Epidemiology 15(8):529-533.
Veríssimo, A., G. Marrão, F. G. da Silva, and M. S. da Costa. 1991. Distribution of Legionella spp. in hydrothermal areas in continental Portugal and the island of São Miguel, Azores. Applied and Environmental Microbiology 57(10):2921-2927.
Viasus, D., S. Di Yacovo, C. Garcia-Vidal, R. Verdaguer, F. Manresa, J. Dorca, F. Gudiol, and J. Carratalà. 2013. Community-acquired Legionella pneumophila pneumonia: A single-center experience with 214 hospitalized sporadic cases over 15 years. Medicine 92(1):51-60.
Vickers, R. M., V. L. Yu, S. S. Hanna, P. Muraca, W. Diven, N. Carmen, and F. B. Taylor. 1987. Determinants of Legionella pneumophila contamination of water distribution systems: 15-hospital prospective study. Infection Control and Hospital Epidemiology 8(9):357-363.
Volk, C. J., and M. W. LeChevallier. 2000. Assessing biodegradable organic matter. Journal of the American Water Works Association 92(5):64-76.
von Baum, H., S. Ewig, R. Marre, N. Suttorp, S. Gonschior, T. Welte, and C. Lück. 2008. Community-acquired Legionella pneumonia: New insights from the German competence network for community acquired pneumonia. Clinical Infectious Diseases 46(9):1356-1364.
Wadowsky, R. M., and B. B. Yee. 1985. Effect of non-Legionellaceae bacteria on the multiplication of Legionella pneumophila in potable water. Applied and Environmental Microbiology 49:1206-1210.
Wadowsky, R. M., L. J. Butler, M. K. Cook, S. M. Verma, M. A. Paul, B. S. Fields, G. Keleti, J. L. Sykora, and R. B. Yee. 1988. Growth-supporting activity for Legionella pneumophila in tap water cultures and implication of hartmannellid amoebae as growth factors. Applied and Environmental Microbiology 54:2677-2682.
Wadowsky, R. M., R. Wolford, A. M. McNamara, and R. B. Yee. 1985. Effect of temperature, pH, and oxygen level on the multiplication of naturally occurring Legionella pneumophila in potable water. Applied and Environmental Microbiology 49:1197-1205.
Wadowsky, R. M., T. M. Wilson, N. J. Kapp, A. J. West, J. M. Kuchta, S. J. States, J. N. Dowling, and R. B. Yee. 1991. Multiplication of Legionella spp. in tap water containing Hartmannella vermiformis. Applied and Environmental Microbiology 57(7):1950-1955.
Walker, J. T. 2018. The influence of climate change on waterborne disease and Legionella: A review. Perspectives in Public Health 138(5):282-286.
Wallensten, A., I. Oliver, K. Ricketts, G. Kafatos, J. M. Stuart, and C. Joseph. 2010. Windscreen wiper fluid without added screenwash in motor vehicles: a newly identified risk factor for Legionnaires’ disease. European Journal of Epidemiology 25(9):661-665.
Wallis, L., and P. Robinson. 2005. Soil as a source of Legionella pneumophila serogroup 1 (Lp1). Australian and New Zealand Journal of Public Health 29(6):518-520.
Wang, C., M. Saito, T. Tanaka, K. Amako, S.-I. Yoshida. 2015a. Comparative analysis of virulence traits between a Legionella feeleii strain implicated in Pontiac fever and a strain that caused Legionnaires’ disease. Microbial Pathogenesis 89:79-86.
Wang, H., S. Masters, J. O. Falkinham, M. A. Edwards, and A. Pruden. 2015b. Distribution system water quality affects responses of opportunistic pathogen gene markers in household water heaters. Environmental Science and Technology 49:8416-8424.
Wang, S. P., J. S. Wang, and H. F. Li. 1995. A study on the risk factors of Legionella infection in children. Zhonghua Liu Xing Bing Xue Za Zhi 16(2):88-91.
Wang, H., M. A. Edwards, J. O. Falkinham, and A. Pruden. 2013. Probiotic approach to pathogen control in premise plumbing systems: a review. Environmental Science and Technology 47(18):10117-10128.
Wang, H., S. Masters, Y. Hong, J. Stallings, J. O. Falkinham, M. A. Edwards, and A. Pruden. 2012. Effect of disinfectant, water age, and pipe material on occurrence and persistence of Legionella, mycobacteria, Pseudomonas aeruginosa, and two amoebas. Environmental Science and Technology 46(21):11566-11574.
Warren, W. J., and R. D. Miller. 1979. Growth of Legionnaires’ disease bacterium (Legionella pneumophila) in chemically defined medium. Journal of Clinical Microbiology 10:50-55.
Wei, S. H. 2014. Nosocomial neonatal legionellosis associated with water in infant formula, Taiwan. Emerging Infectious Diseases 20(11):1921-1924.
Weissenmayer, B. A., J. G. Prendergast, A. J. Lohan, and B. J. Loftus. 2011. Sequencing illustrates the transcriptional response of Legionella pneumophila during infection and identifies seventy novel small non-coding RNAs. PLoS ONE 6(3):e17570.
Wery, N., V. Bru-Adan, C. Minervini, J. P. Delgenes, L. Garrelly, and J. J. Godon. 2008. Dynamics of Legionella spp. and bacterial populations during the proliferation of L. pneumophila in a cooling tower facility. Applied and Environmental Microbiology 74:3030-3037.
Whiley, H., and R. Bentham. 2011. Legionella longbeachae and legionellosis. Emerging Infectious Diseases 17(4):579-583.
Whiley, H., A. Keegan, H. Fallowfield, and K. Ross. 2014. Uncertainties associated with assessing the public health risk from Legionella. Frontiers in Microbiology 5(SEP):1-8.
Whitesides, M. D., and J. D. Oliver. 1997. Resuscitation of Vibrio vulnificus from the viable but nonculturable state. Applied and Environmental Microbiology 63:1002-1005.
Williams, K., A. Pruden, J. Falkinham, and M. Edwards. 2015. Relationship between organic carbon and opportunistic pathogens in simulated glass water heaters. Pathogens 4:355-372.
Woodhead, M., F. Blasi, S. Ewig, G. Huchon, M. Leven, A. Ortqvist, T. Schaberg, A. Torres, G. van der Heijden, and T. J. M. Verheij. 2011. Guidelines for the management of adult lower respiratory tract infections—full version. Clinical Microbiology and Infection 17(Suppl 6):E1-59.
World Health Organization (WHO). 2018. Legionellosis. http://www.who.int/mediacentre/factsheets/fs285/en.
Wullings, B. A., and D. van der Kooij. 2006. Occurrence and genetic diversity of uncultured Legionella spp. in drinking water treated at temperatures below 15C. Applied and Environmental Microbiology 72(1):157-166.
Wullings, B. A., R. Italiaander, and P. W. J. J. van der Wielen. 2016. Distinction between dead and live bacteria using PMA and EMA in combination with qPCR. BTO report 2016.072, KWR Watercycle Research Institute, Nieuwegein, The Netherlands (in Dutch).
Yanagihara, K., S. Kohno, and T. Matsusima. 2001. Japanese guidelines for the management of community-acquired pneumonia. International Journal of Antimicrobial Agents 18(Suppl 1):S45-S48.
Yee, R. B., and R. M. Wadowsky. 1982. Multiplication of Legionella pneumophila in unsterilized tap water. Applied and Environmental Microbiology 43:1330-1334.
Yiallouros, P. K., T. Papadouri, C. Karaoli, E. Papamichael, M. Zeniou, D. Pieridou-Bagatzouni, G. T. Papageorgiou, N. Pissarides, T. G. Harrison, and A. Hadjidemetriou. 2013. First outbreak of nosocomial Legionella infection in term neonates caused by a cold mist ultrasonic humidifier. Clinical Infectious Diseases 57(1):48-56.
Yu, V. 1993. Could aspiration be the major mode of transmission for Legionella? American Journal of Medicine 95:13-15.
Yu, V. L., and T. C. Lee. 2010. Neonatal legionellosis: the tip of the iceberg for pediatric hospital-acquired pneumonia? The Pediatric Infectious Disease Journal 29(3):282-284.
Yu, V. L., J. F. Plouffe, M. C. Pastoris, J. E. Stout, M. Schousboe, A. Widmer, J. Summersgill, T. File, C. M. Heath, D. L. Paterson, and A. Chereshsky. 2002. Distribution of Legionella species and serogroups isolated by culture in patients with sporadic community-acquired legionellosis: an international collaborative survey. Journal of Infectious Diseases 186:127-128.
Yu, V. L., J. Ramirez, J. Roig, and M. Sabria. 2004. Legionnaires’ disease and the updated IDSA guidelines for community-acquired pneumonia. Clinical Infectious Diseases 39(11):1734-1737.
Zähringer, U., Y. A. Knirel, B. Lindner, J. H. Helbig, A. Sonesson, R. Marre, and E. T. Rietschel. 1995. The lipopolysaccharide of Legionella pneumophila serogroup 1 (strain Philadelphia 1): Chemical structure and biological significance. Progress in Clinical and Biological Research 392:113-139.
Zhang, Q., H. Zhou, R. Chen, T. Qin, H. Ren, B. Liu, X. Ding, D. Sha, and W. Zhou. 2014. Legionnaires’ disease caused by Legionella pneumophila serogroups 5 and 10, China. Emerging Infectious Diseases 20(7):1242-1243.


























































