To move cloud-based neuroscience research forward, there are many challenges, including technical, financial, legal, regulatory, ethical, and psychosocial, said Magali Haas. During the workshop, topics that came up frequently included sustainability, infrastructure development, training, and funding needs. Enduring data platform software solutions are needed, along with frameworks for future proofing data, said Haas. Sean Hill added that interoperability between platforms and common data models is key.
Haas argued that tangible next steps and action items are needed that can be collectively addressed. She said that when Cohen Veterans Biosciences faced a similar challenge in which they wanted to develop a platform that would facilitate big data modeling and cloud computing with multiple data types, they went through a 2-year exercise of evaluating hundreds of available platforms according to the 45 identified characteristics, winnowed down to the top 10 requirements (see Figure 11-1). Haas said each platform they examined had different benefits and limitations. In addition, some that were funded by government grants ceased to exist, which emphasizes the importance of sustainability.
Ted Willke, director of the Brain-Inspired Computing Lab at Intel, suggested that it will be important to focus on four or five requirements that are most relevant to the types of data and goals of the study. Jane Roskams added that another key step is to prioritize elements that may be more doable and tangible in the short term versus the long term, and then build from there by developing a readable, accessible roadmap.
TECHNOLOGY AND METHODS: PROGRESS AND CHALLENGES
An example of the opportunities and challenges encountered in developing a new technology was provided by Willke. Closed-loop neurofeedback in the cloud is becoming an important treatment approach for many disorders, including depression, schizophrenia, and posttraumatic stress disorder. It enables visualization of brain activity in real time, which requires a lot of preprocessing and analytics in the cloud. Machine learning could also allow automatic selection of regions of interest in the brain, he said. The architecture needed to achieve this would need to be scaled across multiple machines with unlimited processing memory, storage, and communications
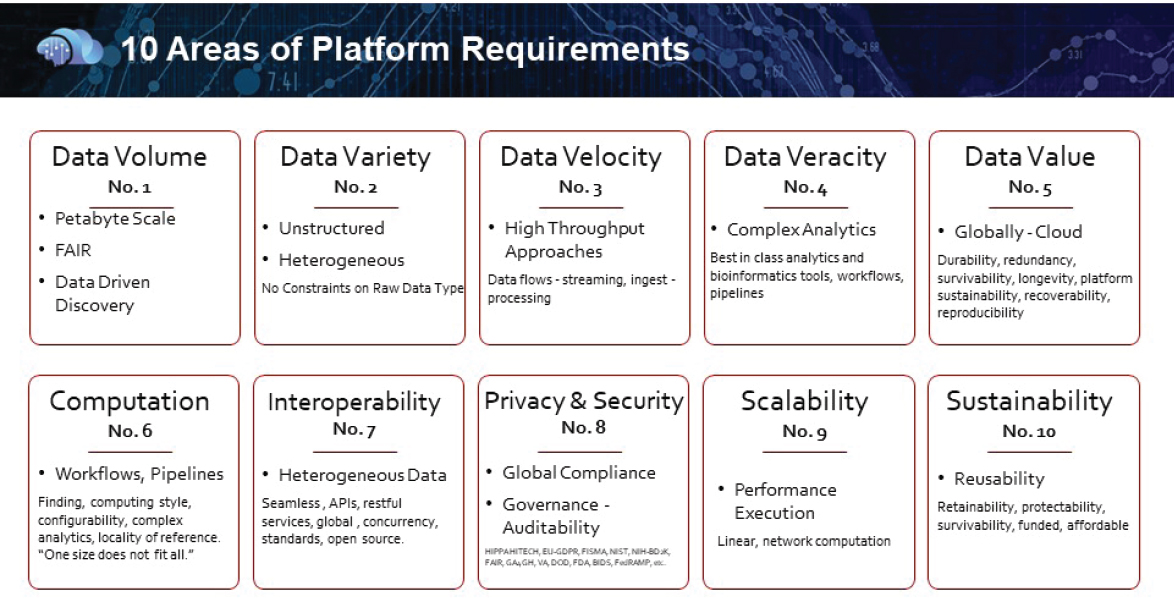
NOTE: API = application program interface; FAIR = findable, accessible, interoperable, reusable.
SOURCE: Presentation by Magali Haas, September 24, 2019.
among systems, and could enable a full brain correlation analysis on one research participant or multiple research participants, possibly in less than 1 second.
Developing these cloud-based technologies requires collaborators to share tools, methods, and data across multiple institutions, and could provide complex systems to researchers who lack the computer science and engineering expertise needed to create such systems themselves, said Willke. Putting all this in the cloud is necessary because it would be too difficult to develop and maintain in an institution’s private cloud or in a container environment. Rather, a parallel distributed system with associated dependencies would be needed, said Willke.
Challenges unique to the cloud, such as network and service delays, will require new stream-based systems frameworks and impact how preprocessing and machine learning models are dealt with, said Willke. Real time brings additional challenges, including being able to describe temporal data and the conditions under which those data were collected. He said much work needs to be done to develop data and processing standards to enable the development of these technologies. Validating that the pipeline will run as fast as needed will also be required. He added that cost will be an important consideration: The system should run as fast as possible, but not use any more resources than necessary.
Willke said there has been progress in developing such real-time cloud systems, including a depression study involving investigators at Princeton, Penn Med, University of Chicago, and Intel. He said this is one of the first uses of running the cloud in one area of the country and servicing a clinical study at a medical school in another.
Other examples of successful open-source data sharing between industry and academia were described by Douglas Landsman, vice president of research at the National Multiple Sclerosis Society (NMSS). In one initiative, NMSS helped organize a large project to develop new validated outcomes for Phase 3 clinical trials using federated data from multiple industry-sponsored trials. More recently, they are leading a multistakeholder international initiative aiming to identify biomarkers of disability progression to speed Phase 2 clinical trials in progressive MS. This project will combine two existing data management and data-sharing systems by combining LORIS (see Chapter 4) with SPINE (Structured Planning and Implementation of New Explorations), a web-based, virtual laboratory designed to support the design and distributed execution of experiments centered on specific scientific questions.1 Using a federated database and cloud computing, this project aims to create local data repositories at multiple sites,
___________________
1 To learn more about SPINE, see https://cni.bwh.harvard.edu/front-and-back-end-development-spine-0 (accessed January 21, 2020).
enabling investigators to conduct experiments and collaboratively review results.
Collecting data from multiple trials and companies has presented numerous challenges, including legal hurdles and a substantial amount of data cleaning and formatting to enable harmonization, said Landsman. These problems are recurrent, predictable, and solvable. There will always be some manual data cleaning required, which will probably require more resources than originally anticipated, said Landsman. Nick Weber advocated structuring data to be as open and accessible as possible and in a way that ensures distribution of costs so that the burden does not fall only on those producing the data. He suggested developing a central clearinghouse of cloud-based resources specific to neuroscience. Maryann Martone added that it will also be important to change the mindset of researchers to understand that data they are producing will be reused at some point, so that they collect and manage data with that in mind.
TRAINING THE NEXT GENERATION OF SCIENTISTS
Training the next generation of neuroscience researchers in the use of cloud-based technologies is essential across different domains and stages, from graduate students new to the field to more senior scientists who may need retraining, said Deanna Barch. Educating the next generation of researchers to understand cloud-based tools and start using them in early stages of their research could accelerate clinical use, said Willke. He noted that both Princeton and Yale have begun integrating cloud-based toolkits into their curricula. Haas added that training is also needed across a broad range of stakeholder groups to increase understanding of the regulatory implications of cloud technologies. Story Landis, former director of NINDS, suggested that it might be possible to engage professional societies and meeting organizers to conduct such trainings at conferences.
FUNDING: CURRENT COMMITMENTS AND FUTURE NEEDS
Fundraising from governmental and private sources is needed to accomplish foundational work for data standardization, hosting, and reproducibility, and to build advanced cloud infrastructure, said Willke. Jonathan Cohen noted that the 5-year effort Willke described was funded entirely by Intel and is entirely open source, making it a model of academic–industrial collaboration.
Congress has appropriated $20 million for the development of a data warehouse primarily for opioid-related activities at FDA, said Silvana Borges. FDA hopes to include not only data from different sources, such as data related to distribution, prescription, and abuse patterns; safety data;
and data from social media. Integrating these types of data adds complexity to the construction of the warehouse because the data may be in non-standardized forms and from unfamiliar or distant sources. Building an infrastructure for data linkage and the analytical tools needed to integrate these different sources of information should help address the opioid crisis while also providing a model for future data integration efforts in other disease areas, said Borges. However, she acknowledged concerns about the sustainability funding for these efforts.
Landsman said NMSS is eager to partner with investigators and provide funding for good, well-planned, and potentially long-term data management projects. He suggested that funders, including both NIH and private sources, could require data management plans in future projects as a condition of funding.
POTENTIAL NEXT STEPS: WORKING GROUPS TO MOVE THE FIELD FORWARD
Individual workshop participants discussed potential next steps that could help researchers and institutions navigate using the cloud for neuroscience data storage and computation.
Articulating long-term strategic goals and prioritizing them will ultimately be the most productive way to make progress, stated Michael Hawrylycz. He added that it is clear that informatics and the cloud will become increasingly important in neuroscience. It will therefore be more productive to proactively imagine where the field would ideally head and plan for that, rather than implementing short-term fixes.
Some specific areas that could benefit from such proactive strategic planning that individual workshop participants raised included
- Developing common data model frameworks and support for data transformations between platform or software versions, said Haas. Hill agreed that common data models are an important part of a potential solution that would address interoperability among platforms.
- Defining and advocating for a policy about, and potentially protections against, re-identification of patients based on pooled data, said Haas.
- Building sustainability models for software, infrastructure, and funding, including support for enduring data platforms. For example, Haas noted that in an evaluation performed by Cohen Veterans Bioscience, some well-run platforms supported by government grants disappeared unexpectedly when funding lapsed.
To design future platforms and tools, Walter Koroshetz, director of NINDS, said that focusing on utility for users can help prioritize the most valuable actions or resources, and inform both scientific and financial decisions. Pilot examples from the BRAIN initiative, and the AMPs on AD and PD, could be informative starting places. Other approaches to learning from existing initiatives could be to catalog existing platforms, data, and pipelines. Haas said that existing platforms could be evaluated for specific capabilities and as models, such as which platforms have the “best-in-class solutions” for different data types or models. This work could be coordinated with existing coordinating bodies, such as NIH or INCF.
To support broader and easier use of the cloud, multiple individual workshop participants highlighted the benefits of assembling existing resources and examples of consent forms and governance materials that could be used as templates. Stacia Friedman-Hill said that the NIMH Clinical Research Toolbox is an existing resource that links out to information about regulations for clinical research. That website could potentially also be a resource to collect or post consent form language in a centralized repository.
Similarly, other areas that could benefit from collecting, evaluating, and comparing existing practices that individual participants discussed included reporting incidental findings in genetic analyses, data storage and prioritization, and sharing code with protections. Michael Milham noted that establishing best practices for assigning credit and incentivizing data sharing would be beneficial, particularly for tenure and promotion decisions.
Some participants also discussed training as a mechanism to support greater use of the cloud, noting that training could be tailored to different groups and encompass different levels of familiarity. Several examples were for new trainees such as graduate students; for those who have some experience, but could benefit from additional information; for funders of research; or for oversight groups such as IRBs, said Barch. Haas said that redesigned training could facilitate understanding of implications for using the cloud, including broad data sharing or regulatory implications. Roskams added that an essential component to a successful training initiative would be to develop a widely accessible “toolkit” for use, especially for institutions that may have fewer resources available to develop training programs de novo.
Planning for widespread use of the cloud in the future will benefit from engaging a broad group of users now. Koroshetz suggested involving users who may not be as familiar with cloud-based tools, and who are not data scientists. This step could improve overall utility and guide iterations of future products based on existing platforms. Huerta noted that it would be good to engage leaders of existing data platforms in any follow-on activities that might be organized after the workshop.
Although many challenges still need to be addressed, from technical to legal to ethical to cultural, there is also a great deal of excitement about what is possible, said Haas, and there is the will and ability to take on the challenges and usher in better—and more widespread—use of the cloud for neuroscience data.








