Summary
The taxonomic status of the modern red wolf, Canis rufus, has been controversial scientifically because modern red wolves contain some amount of coyote ancestry. The same observation has sparked controversy in the realm of conservation. The extensive recovery efforts devoted to extant captive and managed red wolves1 (also referred to in this report as the extant red wolves or extant red wolf populations) are not universally popular, and critics of those efforts have argued that they are misplaced because the modern red wolf is not a valid species. Once common throughout the eastern United States, red wolves were extirpated from the wild by 1980. In 1987, individuals from a captive breeding program were used to re-introduce red wolves into a protected area in North Carolina. The re-introduced individuals established a population, and the U.S. Fish and Wildlife Service (USFWS) now manages this population.
To found the captive breeding colony, the USFWS had selected a small number of individuals from a population of canids discovered in western Louisiana. The USFWS selected individuals for the breeding colony based on their resemblance to wolves in morphology and vocalizations, supported by analyses of genetic variation in enzymes. However, subsequent genetic studies of the managed population indicated that these wolves displayed evidence of significant coyote ancestry. This discovery raised the issue of whether the extant captive and managed red wolves was a valid species that was eligible for conservation and recovery. Opponents of the USFWS effort to restore the red wolf have used questions about the ancestry of the extant populations to argue that the restoration program is flawed.
In March 2019, the National Academies of Sciences, Engineering, and Medicine issued a report entitled Evaluating the Taxonomic Status of the Mexican Gray Wolf and the Red Wolf,2 which
___________________
1 Captive red wolves refer to the approximately 245 red wolves maintained in 43 breeding facilities all throughout the United States. Managed red wolves refer to the individuals comprising the nonessential experimental population in eastern North Carolina.
2 NASEM (National Academies of Sciences, Engineering, and Medicine). 2019. Evaluating the taxonomic status of the Mexican gray wolf and the red wolf. Washington, DC: The National Academies Press. https://doi.org/10.17226/25351.
addressed the taxonomic status and evolutionary history of the red wolf. The report was prepared upon the request of the USFWS and addressed these three questions: (1) Is there evidence that the historical population of red wolves was a lineage distinct from other canids? (2) Is there evidence that the extant (captive and managed) populations of red wolves in the United States represent a lineage of canids distinct from modern gray wolves (C. lupus) and coyotes (C. latrans) (Figure S-1)?, and (3) Is there evidence for genetic continuity between the historical red wolf populations in the eastern United States and the extant (captive and managed) populations of red wolves?
The authors of the 2019 report answered “yes” to the first two questions. With respect to the third question, the authors concluded that there was evidence that the extant populations traced some of their ancestry to the historical red wolf and that this was enough evidence to support species status for the extant red wolf. This finding retained the existing taxonomic designation of the red wolf and reinforced the validity of conserving and restoring red wolves. However, the authors acknowledged that additional genomic evidence from historical specimens could change this assessment by changing the answers to all three questions.
While the 2019 report was being prepared, a group of researchers3 described gene markers indicating red wolf ancestry for a wild, unidentified canid population in southwestern Louisiana. In addition, another group of researchers4 discovered alleles in a canid population in Galveston Island, Texas, that had not been observed previously in coyotes or extant red wolves. These alleles, called “ghost alleles,” could represent genes characteristic of historical red wolves that were lost in the selection of the founders of the breeding colony. If, as these discoveries suggest, these two populations are remnant red wolf populations and harbor genetic information unique to red wolves, then the status and evolutionary history of the red wolf needs to be re-examined.
Prompted by those discoveries about the canids in southwestern Louisiana and Galveston Island, Texas (referred to as Gulf Coast canids or GCC populations in this report), the USFWS requested the National Academies in July 2019 to appoint an ad hoc committee that would carry out two tasks: (1) solicit and review applications to carry out research to determine the taxonomy of the canid population in southern Louisiana, and (2) develop a research strategy to further assess

SOURCE: B. Bartel, U.S. Fish and Wildlife Service (left) and Melba Coleman (middle and right).
___________________
3 S. M. Murphy, J. R. Adams, J. J. Cox, and L. P. Waits. 2019. Substantial red wolf genetic ancestry persists in wild canids of southwestern Louisiana. Conservation Letters 12(2):e12621.
4 E. Heppenheimer, K. E. Brzeski, R. Wooten, W. Waddell, L. Y. Rutledge, M. J. Chamberlain, D. R. Stahler, J. W. Hinton, and B. M. vonHoldt. 2018. Rediscovery of red wolf ghost alleles in a canid population along the American Gulf Coast. Genes 9(12):618.
the taxonomy of the red wolf. The first task was completed in January 2020.5 This report addresses the second task from the USFWS. The full statement of the second task was:
The committee will develop a research strategy to examine the evolutionary relationships between ancient red wolves, the extant red wolf population, and the unidentified canid populations in southern Louisiana. The committee will identify the types of studies needed to improve understanding of genetic ancestry, phylogenetic relationships, morphology, behavior, and ecology, taking into consideration evidence about the role of admixture and hybridization in speciation. Although the strategy will focus on the study of the red wolf, the committee may draw upon knowledge generated from research on other organisms, particularly those with taxonomic questions parallel to those of the red wolf. The strategy will also include key considerations for sampling and analyses to minimize impact on specimens and live organisms when carrying out research.
WHY DO MORE RESEARCH ON THE RED WOLF?
There are three possible explanations for the significant amount of coyote ancestry in modern red wolves. First, the extant red wolves may be hybrids between coyotes and either gray wolves or what were once red wolves. Hybridization may have occurred long ago or in more recent years. Evidence suggests that some hybridization certainly occurred in the 20th century as red wolf populations dwindled in size while coyotes expanded their range eastward. Second, the extant red wolves may have diverged from coyotes recently enough that they continue to share alleles, even though each species has its own forward evolutionary trajectory—a situation referred to as incomplete lineage sorting. Third, hybridization and introgression of coyote genes into red wolves could reflect historical phenomena associated with the ancestral origin of the red wolf. In this case, the presence of coyote alleles would be the result of hybridization between coyotes and gray wolves (or another ancient canid) that gave rise to the red wolf, in which some level of reproductive isolation exists, but is incomplete. These three possible explanations for shared alleles between coyotes and red wolves are not mutually exclusive. Therefore, resolving these possible explanations requires tracing the ancestry of extant red wolves and ascertaining when coyote genes entered the lineage of red wolves.
Another reason why the controversy over the status of red wolves persists is the paucity of data from ancient specimens that could serve as a standard for the red wolf before hybridization with coyotes in the 20th century. There are no morphological data from ancient specimens (specimens dated prior to 1800), and there is no evidence from the nuclear DNA of ancient specimens. There are data for mitochondrial DNA (mtDNA) sequences from only four ancient specimens.6 Those data indicate a close relationship between coyotes and red wolves but not the nature of that relationship.
Data from historical specimens (specimens collected between 1800 and the start of the eastward expansion of coyotes in 1920) do not resolve questions about the ancestry of red wolves. Analyses of morphological data from historical specimens indicate that specimens identified as red wolves are distinguishable from specimens of gray wolves and coyotes. However, those specimens identified as red wolves have many intermediate characteristics of gray wolf and coyote specimens, which is consistent with either a recent divergence of red wolves from coyotes and gray wolves or a hybrid origin of red wolves. Analyses of mtDNA from historical specimens of putative red wolves place them in a lineage with coyotes, but these data are from specimens collected at the western edge of the historical range of red wolves, where the range abutted that of coyotes.7 As a result,
___________________
5 National Academies of Sciences, Engineering, and Medicine. 2020. Evaluation of Applications to Carry Out Research to Determine the Taxonomy of Wild Canids in the Southeastern United States. Washington, DC: The National Academies Press. https://doi.org/10.17226/25661.
6 See Appendix C.
7 Red wolf historical range is illustrated in Figure 1-2 in Chapter 1; red wolf specimens are listed in Appendix D.
whether the mtDNA from historical specimens originates from coyotes or red wolves cannot be resolved without an analysis of nuclear DNA from the same specimens.
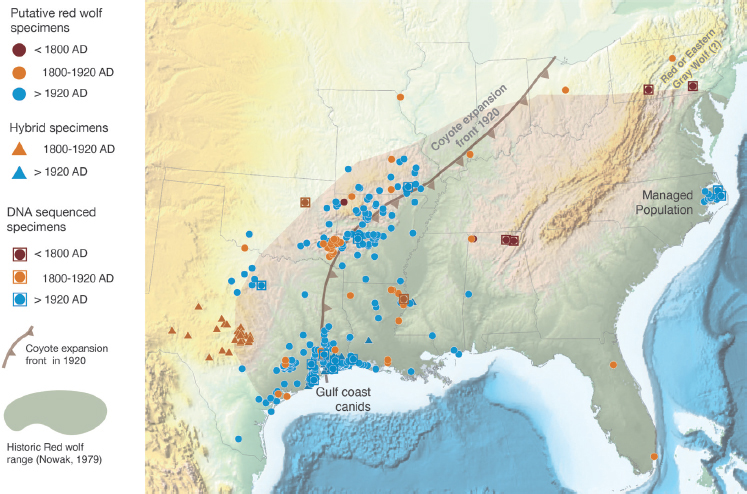
Genetic and genomic data of extant putative red wolves come from populations at the southwestern edge of the historical range (western Louisiana and eastern Texas) and the extant red wolf populations (Figure S-2). The populations at the southwestern edge of the historical ranges are the ones from which the progenitors of the managed population in North Carolina were selected. It is unclear how well those populations represent what was once a numerically large, geographically widespread population of canids.
EXAMINING THE EVOLUTIONARY RELATIONSHIPS AMONG ANCIENT RED WOLVES, THE EXTANT RED WOLF POPULATIONS, AND THE UNIDENTIFIED GULF COAST CANID POPULATIONS
The guiding principle for the research strategy described in this report is that it must address the relationships among ancient red wolves, historical red wolves, the extant red wolves, and the unidentified canids along the Gulf Coast (GCC populations). These relationships will indicate whether red wolves are, or once were, a distinct lineage (a species sharing a common ancestor with either coyotes or gray wolves or a species that has an ancient, hybrid origin) or whether “red
wolves” are recent hybrids between gray wolves and an expanding coyote population. They will also clarify how the extant red wolves are related to the GCC populations. Resolving those relationships requires answering five research questions:
- Are the ancient red wolves (i.e., specimens dated prior to 1800) a distinct lineage from the lineages of gray wolves and coyotes?
- Are the historical red wolves (i.e., those in the 19th century, before the eastward expansion of coyotes) a continuous lineage to ancient specimens in the eastern United States?
- Are the red wolves from the extant captive and managed populations a continuous lineage with those in the eastern United States before the coyote expansion in the 20th century?
- Are the Gulf Coast canid populations part of the distinct lineage of canids that lived in the eastern United States before the coyote expansion?
- Is the genetic diversity of the extant red wolf populations a subset of the diversity of the Gulf Coast canids?
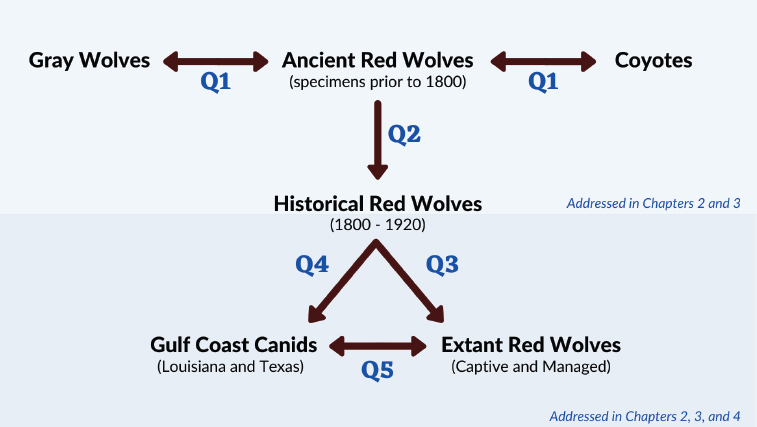
To clarify the relationships among these canids, new research should employ three types of data: morphological data from ancient specimens and extant animals, ancient DNA, and genomic data from the unidentified canids and the extant red wolf populations. Figure S-3 outlines the five questions, the canids to which each question pertains, and where this report describes the necessary data. The next sections summarize the committee’s findings and conclusions regarding the contribution of each type of data to addressing the questions outlined in the statement of task requested by the USFWS.
Morphological, Behavioral, and Ecological Data
Morphological data from ancient and historical red wolves are somewhat limited, but even more limited are behavioral and ecological data. Nevertheless, an understanding of these types of data collected from ancient and historical red wolves, extant red wolf populations, and the GCC populations could be helpful in understanding their evolutionary relationships and the consequences of those relationships.
Findings
2-1: Previous analyses of the morphological distinctiveness of the red wolf have paid close attention to uncertainties about taxonomy and possibilities of hybridization. The best material to characterize historical red wolf morphology was collected from Louisiana, Mississippi, Alabama, Florida, and Arkansas between 1800 and 1920. The number of specimens is meager but minimally adequate for many kinds of morphological comparisons.
2-2: Prior analyses of the morphological differentiation of red wolves are comprehensive and their conclusions are well supported, but the kinds of data and analyses that have been applied are limited. Geometric morphometrics can pick up subtle differences in shape that cannot be captured by linear measurements, traits such as teeth and postcrania can capture fine-level genetic differences or ecologically relevant specializations not represented by skull data, and analytical methods such as QST and cluster analysis can provide measures of morphological differentiation that are directly comparable to results from genetic analysis.
2-3: The skull measurements used in past morphological analyses have less power than non-metric traits for detecting hybridization. Anecdotal reports of some historical red wolf samples suggest the presence of hybrid individuals, but no attempt has been made to systematically analyze morphological hybrid signatures in historical red wolf specimens.
2-4: Sexual dimorphism in morphology has been noted in red wolves, but it has neither been systematically evaluated nor used to assess the likelihood of hybridization or of sex-biased introgression processes.
2-5: Behavioral differences between species can affect the likelihood of hybridization, choice of habitats, and choice of prey. Furthermore, these behaviors may depend on the environmental context in which an individual finds itself and thus be plastically modified as habitats change. Much has been recorded about red wolf behavior, but most of it has been based on individuals in the hybrid swarm of east Texas in the 1960s and in the managed population, neither of which may be representative of historical red wolf behaviors.
Conclusions
2-1: To distinguish among the origin hypotheses for the red wolf using morphology, additional kinds of data and analysis are required. The existing studies based on skulls can be supplemented by similar analyses of teeth, which are likely to better capture genetic relationships among taxa, and of postcrania, which are likely to supplement the understanding of ecological differentiation of red wolves from coyotes and gray wolves. Geometric morphometric data will enable finer level discrimination based on details of shape that were not captured in the original analyses.
2-2: New analytical techniques are required to assess the congruence between morphological and genetic evidence for the distinctiveness of the red wolf. Prior analyses relied solely on discriminant function analysis, which is a powerful tool for determining what is different between known groups, but it is less powerful for determining which groups are more similar, how variation at the subspecific level compares with differentiation between species, or whether differences are due to hybridization. The same hierarchical analyses of variance, cluster analyses, and pairwise distance analyses that are commonly used on genetic data can be applied to morphological data to facilitate these comparisons.
2-3: The behavioral data that exist appear to be consistent with the hypothesis that hybridization happened late in the red wolf’s history, not at its inception, but behavioral data for historical red wolves are sparse. The behavioral data that exist deserve to be more comprehensively analyzed with regard to hypotheses about hybridization in canids, and morphological indicators of behavior are needed to supplement direct observations of behavior.
Recommendation: The committee recommends that existing morphological datasets be submitted to new analytical techniques that are directly comparable to genetic results, that ancient DNA sequencing and morphological analyses be performed on the same individuals, and that new morphological data be collected that are relevant to assessing the ecological and behavioral distinctiveness of historical red wolves in their core range.
Genomic Data from Historical and Ancient Canids
Genomic data from ancient and historical canids would enable the investigation of the ancestry and population history of red wolves. To be useful, genomic data must be sufficiently fine-grained to make it possible to discern whether the red wolf lineage is distinct from other wolf lineages in North America and to detect the occurrence and timing of historical hybridization events among lineages. For such analyses, a well-planned selection of samples and careful data collection and filtering will be essential. A sampling strategy wherein specimens are collected from individuals from across North America prior to 1800 (“ancient” specimens) and from individuals from post-1800 to 1920 (“historical” specimens) will yield a latitudinal and longitudinal survey of genetic diversity in canids across North America. The data could indicate whether:
- the red wolf is a distinct taxonomic lineage that shares a most recent common ancestor with either coyotes or gray wolves;
- the red wolf is a lineage that resulted from ancient hybridization between two canid species (e.g., between coyote and gray wolf or some now-extinct canid lineages); and
- the animals termed “red wolves” are possibly the result of recent (i.e., occurring within historical, not ancient, time) hybridization between coyotes and gray wolves.
Findings
3-1: To collect a data set maximally useful for discerning the taxonomic status of the red wolf, it will be necessary to include canid samples of broad geographic and temporal provenance, and to prioritize individual samples that can yield high-quality morphological and genomic data.
3-2: Many steps in the collection and analysis of genetic data can introduce bias. Methods that sequence the entire genome, rather than individual sites, will be maximally informative for addressing questions of lineage identity and history.
3-3: In work with ancient DNA samples, specialized methods are necessary to minimize the risk of contamination and preserve the integrity of DNA molecules.
3-4: Filtering genomic data collected from ancient samples is essential to eliminate contaminating sequences and biases arising from damage accrued over time.
3-5: To assess questions of lineage origin and continuity for the red wolf, analytic approaches will need to account for the possibility of previous introgression and uncertainties in its timing.
Conclusions
3-1: Biases in collection and analysis of genetic data can be mitigated through care in selecting samples, application of appropriate methods to isolate DNA, and analysis of data from the entire genome rather than from small subsets of the genome.
3-2: Genomic data from ancient and historical canids would make it possible to investigate the ancestry and population history of red wolves. In particular, high-resolution whole-genome data will be needed to determine with high confidence whether the historical red wolf lineage is distinct from other wolf lineages in North America and to screen for the occurrence and timing of possible hybridization among lineages.
3-3: To discern the taxonomic identity and relationships of contemporary and late 20th century red wolf populations with each other and with earlier canids, it will be necessary to have genomic data from individuals that lived prior to the coyote expansion to the eastern United States. In addition, genomic data from other canids that lived along the proposed range boundaries and from farther afield, across a range of time periods, will be needed to provide a data set for comparison.
Genomic Analysis of Extant Canids
The examination of the evolutionary relationships among ancient red wolves, the extant red wolf populations, and the GCC populations will require placing these populations in the context of genomic data from a broader continuum of modern canid populations. Different types of genomic data (e.g., whole-genome, RAD-seq, and SNP arrays) differ in their power to address questions about lineage continuity among the extant red wolves, the GCC populations, and the historical red wolf populations in the southeastern United States.
Findings
4-1: Three main types of genomic data have been used to analyze the taxonomic identity of the red wolf. While all of these data types can provide useful information, whole-genome sequence data constitute the least biased data type that can provide maximum genomic representation and strong inferential power to address questions related to the taxonomic identity and continuity among extant and historical canid populations.
4-2: A large number of canid genomes are available that can be used to determine the level of genetic continuity among the extant red wolf populations, GCC populations, and historical red wolf populations, but further whole-genome sequencing will be required to achieve a more complete representation of the genetic variation across space for some taxa. In particular, additional whole-genome data from one Mexican wolf and one Eastern wolf are required as well as whole-genome
data from a larger number of individuals from each of the extant red wolf populations and the GCC populations.
4-3: The allele frequency spectrum of variation of the reference historical red wolf population and the statistical test of Schraiber can be used to analyze whole-genome data from the extant red wolf populations and the GCC populations to determine whether these populations represent a continuous line of descent from the historical population. Similarly, this analytical approach could be used to determine whether the extant red wolves represent a subset of the GCC populations.
4-4: A number of analytical approaches can provide evidence of hybridization using either RADseq or whole-genome data. However, analyses to determine whether hybridization is recent or ancient among North American canid populations will require unbiased whole-genome data sets in order to make strong demographic inferences throughout the genome as well as to identify and determine the size of ancestry tracts.
Conclusions
4-1: To trace the genetic history of extant red wolf populations (captive and managed) and the GCC populations, using whole-genome sequencing methods on modern fresh tissue samples will provide the most informative data with which to assess both genetic continuity between populations over space and time and historical admixture with other North American canids.
4-2: Although expensive to produce and computationally intensive to analyze, whole-genome data will provide the best chance to identify introgressed genomic fragments (i.e., ancestry haplotypes) and to estimate the time scale at which gene flow occurred.
Recommendation: The committee recommends that new whole-genome sequence data be built upon the existing data with a carefully constructed set of contemporary samples including all North American canid genetic lineages and the full geographic range of all North American canids (red wolf, gray wolf, jackal, coyote, and domestic dog).
COMPONENTS OF THE RESEARCH STRATEGY
Taken together, the five research questions presented earlier address the problems outlined in the statement of task. Each question addresses the relationships between particular pairs of canids (among three canid groups for Question 1) within a specific period.8 Obtaining answers to the first four questions require integrating new genomic data with new morphological data.9 Answering the fifth question will require further analyses of some of the data collected to answer the other questions. The combined results will describe the relationships among the key canid populations with respect to historical continuity and contemporary relatedness.
OVERALL RECOMMENDATIONS
Beyond the specific findings, conclusions, and recommendations in each chapter about the types of data and analyses best suited to address the five research questions, the committee developed and offers the following four overarching recommendations.
___________________
8 See Figure 5-1 in Chapter 5.
9 See Figure 5-2 in Chapter 5.
5-1: The U.S. Fish and Wildlife Service (USFWS) should adopt the proposed research strategy. The USFWS should encourage researchers to form collaborations that include experts in the extractions and analyses of ancient DNA, the collection and analyses of genomic data, the collection and analyses of morphological data, and the collection and analyses of behavioral and ecological data. The call for such a collaborative effort reflects the committee’s conviction that the integration of multiple types of data is needed to answer the questions in a compelling manner.
5-2: The USFWS should support the research strategy via funding. The funding should be sufficient to allow teams of researchers to perform the sampling necessary to answer each question. While data from some individual samples can contribute to answering more than one question, any attempt to economize by attempting to use one set of samples for all questions is unlikely to resolve any of the individual questions.
5-3: The USFWS should be open-minded about refining the strategy as initial results emerge. Research strategies are templates based on knowledge in hand. As new knowledge is gained, the emphases of a strategy may change, new elements may be appropriate, and old elements may appear unnecessary or at least far less important. The USFWS should make the execution of this strategy an iterative process in which researchers can reconsider priorities as a sufficient amount of new data becomes available.
5-4: The USFWS should work with the research community, including professional societies, to create opportunities for larger community involvement in the synthesis and dissemination of research results. In particular, having a shared space for dialogue among researchers with different interpretations of the existing data may produce improvements in the research strategy or in the plans for its individual components. It can also lead to a robust conclusion about the taxonomic status of the canids identified now as red wolves. Species delimitation remains a difficult and often controversial topic, especially when taxa have recently diverged and there is evidence of past admixture. An open-minded approach to all of the data is the best approach, and open-mindedness is aided when the community of experts are engaged in interpretation, discussion, and debate.










