2
Morphology, Behavior, and Ecology
This chapter addresses how the consideration of morphology, behavior, and ecology can help develop a research strategy to understand the evolutionary relationships among historical red wolves, the extant red wolf populations (captive and managed), and the Gulf Coast canid (GCC) populations. Morphological data from ancient and historical red wolves are somewhat limited. Behavioral and ecological data concerning ancient or historical populations of red wolves are even more limited. Nevertheless, an understanding of the morphology, behavior, and ecology of the extant red wolves and the GCC populations could provide insight into their evolutionary relationships. While genomic data will provide the strongest evidence for distinguishing between the four hypotheses described in Chapter 1 (see Figure 1-4), morphological and behavioral data are relevant for (1) determining the extent to which the managed population and GCC populations retain the distinctive features of historical red wolves, (2) evaluating the status of historical and ancient specimens from which genomic data cannot be obtained but which provide the primary data for past distributions of wolves and coyotes, and (3) assessing whether red wolves present and past are ecologically distinct from gray wolves and coyotes.
This chapter discusses the best samples, traits, and methods that can be used to characterize the morphology, behavior, and ecology of historical red wolves and to assess their level of differentiation from other canids and their similarities with (or differences from) the extant red wolf and GCC populations. Specifically, the chapter describes the best available approaches for collecting and analyzing such data to address the following questions:
- Did wolves in the southeastern United States prior to 1920 differ from contemporary gray wolves and coyotes and, if so, in what ways?
- Which hypotheses about the origin and species status of red wolves are supported by morphological evidence?
- How does the morphological and behavioral variation in the extant red wolf and the GCC populations compare with that found in historical red wolves?
MORPHOLOGY
Samples Required to Assess the Morphology of the Historical Red Wolf
Pre-1920 samples from Louisiana, Mississippi, Alabama, Florida, and perhaps Arkansas, which are referred to here as the red wolf’s “core range” (Figure 1-2 in Chapter 1), are the best source of information about its pre-bottleneck morphology (Nowak, 1979, 2002; Chambers et al., 2012).
Hybridization between remnant southeastern wolf populations and coyotes was documented in Texas, Oklahoma, and Missouri in the mid 20th century, making those samples unsuitable for characterizing historical red wolf morphology. The same is true for the extant red wolf populations. The captive breeding colony was founded by only 14 individuals selected from a group of 43 wild-caught animals believed to be red wolves, which were themselves a subset of 400 wild-caught individuals, based on similarity to the historical red wolf phenotype (USFWS, 1990; Phillips et al., 2003). The morphology, behavior, and ecology of the captive and managed red wolves should not be assumed to be representative of historical red wolves because of the possibility of random genetic drift and anthropogenic selection.
Historical samples from central Texas are also unsuitable because hybridization between red wolves and coyotes was probably taking place there as early as the 1850s (Audubon and Bachman, 1851; Nowak, 1979).
Likewise, historical specimens from the northeastern seaboard should be avoided because the taxonomy of pre-disturbance wolves in that area is contested, with some arguing that the red wolf ranged as far north as Maine and New Brunswick, Canada, and others arguing that those populations were gray wolves, hybrids between grays and reds, or even a separate North American species Canis lycaon (cf. Young and Goldman, 1944; Wayne et al., 1995; Wilson et al., 2000; Nowak, 2002; vonHoldt et al., 2016).
Finally, archaeological and paleontological specimens should be avoided for the initial morphological characterization of red wolves. Such specimens often have uncertain dates, they are sporadically distributed through time, they are often fragmentary, and their taxonomy should be treated as uncertain given the subtle level of skeletal differentiation between canid species in light of the fact that the ranges of large mammals can shift thousands of kilometers over millennia (e.g., Graham et al., 1996; Lyons, 2003; Bell et al., 2010). For example, caribou occurred in Alabama as little as 12,000 years ago (Churcher et al., 1989). It is unsafe to presume the identity of a large fossil canid from the southeastern United States just because it lived within the historical range of the red wolf.
Assessing the morphological distinctiveness of the historical red wolf should thus start with pre-1920 specimens from its core range compared with contemporary gray wolf and coyote specimens from well outside the areas with potential hybridization. The same principles used to select red wolf specimens must also be applied to selection of gray wolf and coyote samples—specimens whose taxonomic affinity is not contested and that lived prior to changes in gene flow that may have occurred due to post-European impacts on habitats and populations. Nowak’s (1979) morphological analysis made a careful distinction between historical and later samples and those likely to be affected by hybridization or introgression, and future studies should follow his lead. Appendix B (Red Wolf [C. rufus] Museum Specimens) lists all of the putative museum specimens known to the committee. Of the red wolf, only 12 specimens are known to exist from its core range at the appropriate time period. This sample is very small for characterizing the range of variation that would have been present in such a widespread species, and it will provide only minimal statistical power for testing the similarities and differences between historical red wolves and other groups of canids. For assessing morphological differentiation among populations of the same species, 6 to 10 specimens can be enough to detect differences in skeletal traits if the number of samples is sufficiently numerous and appropriately chosen, if the traits are measured with high precision,
and if multivariate traits with high genetic components variance are considered (e.g., Polly, 2003, 2007b; see discussion below). However, these specimens have the advantage that the individuals are dispersed in space and time and likely to be a random subsample that is unaffected by close kinship or other factors that might bias it. Treating these historical “core” specimens as a cluster for comparison with gray wolves, coyotes, extant red wolf populations, and the GCC populations is operationally the best way to proceed. The amount of comparative material available for historical populations of eastern gray wolves is even more meager, but sufficient material is available from the historical range of coyotes and many other gray wolf populations (Appendix C: Summary of Museum Holdings of Key North American Canids).
Finding 2-1: Previous analyses of the morphological distinctiveness of the red wolf have paid close attention to uncertainties about taxonomy and the possibilities of hybridization. The best material to characterize historical red wolf morphology is from Louisiana, Mississippi, Alabama, Florida, and Arkansas collected between 1800 and 1920. The number of specimens is meager but minimally adequate for many kinds of morphological comparisons.
Data and Methods for Assessing Morphological Affinities
Phenotypes are a key component of species differentiation and ecological specialization. Determining the extent to which red wolves, both extant and historical, are distinct from other canids can be informed by morphology as well as genetics. While much work has been done on morphological traits in North American canids (see review in NASEM, 2019), the last comprehensive study of morphological differentiation within and between North American canids was the work of Nowak (1979) using discriminant function analysis (DFA) of skull measurements. Skull data have several strengths for measuring differentiation among groups, including their developmental and genetic complexity, but other traits have the potential to add to the phenotypic and ecological understanding of differentiation among canids (Box 2-1). Since Nowak’s study was published, geometric morphometrics has been added to the toolbox for quantifying morphology (Box 2-2), and new analytical strategies have been developed for comparing the morphological structuring of populations and species with genetic structure and for inferring ecological parameters from museum specimens (Box 2-3). These methods have considerable potential to increase the understanding of whether historical red wolves were a distinct entity and of what patterns of continuity they share with the extant red wolf and GCC populations.
The original description of red wolves in central Texas by Audubon and Bachman (1851) focused on their small size, slender proportions, and long limbs. Much attention has been given to differentiation in skull morphology, but very little attention has been given to postcranial variation in North American canids with respect to evaluating the genetic and ecologic distinctiveness of the red wolf, although it has been studied with respect to coyote and wolf evolutionary ecology since the last glacial maximum (Meachen and Samuels, 2012; Meachen et al., 2014; Tomiya and Meachen, 2018). It is known that proportional foot length and foot area affect the locomotor performance of coyotes in snow and therefore their behavior on the landscape (Murray and Boutin, 1991; Murray and Larivière, 2002; Gese et al., 2013). Few comparative data are available, but those data suggest that there may be measurable differences in foot proportions among North American canids (Polly, 2010; Polly et al., 2017). Such data could illuminate both the degree of historical differentiation between red wolves and their contemporary canids and the comparative ecological roles of red wolves and GCC populations in past and present.
For example, one fundamental question about the historical red wolf is whether it was a distinct species or the product of hybridization. Nowak (1979) addressed the question using DFA of linear measurements of the skulls of a very large sample of pre-1930 canids from North America
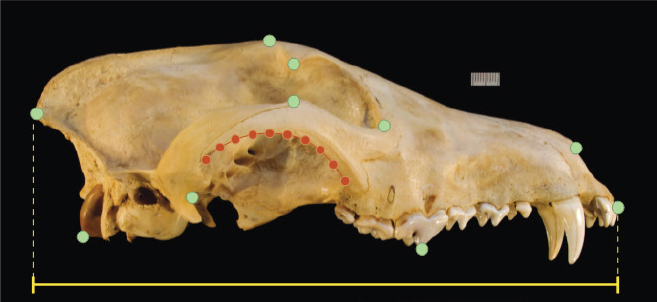
(Box 2-2). After identifying the characteristic features of gray wolves, coyotes, and domestic dogs, Nowak demonstrated that red wolves had skull morphologies intermediate between gray wolves and coyotes but distinct from both; Nowak further found that some pre-1900 specimens from central Texas and some post-1960 specimens from east Texas and western Louisiana were not distinct and thus likely to be hybrids with coyotes but that some of the individuals that founded the managed population had morphologies consistent with being pure red wolves. The DFA provided strong evidence that historical red wolves in their core range were distinct, but it was unable to choose among the competing hypotheses for red wolf origins outlined in Chapter 1. Some authors have rightfully argued that the consistent intermediate position of red wolves could indicate a hybrid origin rather than ancient species status (e.g., Wayne, 1992). DFA alone cannot resolve this difference in interpretation because this technique can create the illusion that the morphology of a previously unclassified individual is perfectly intermediate between two groups even if it belongs to a third group with its own uniquely distinctive features. The reason is that DFA axes are defined explicitly by the differences between the known groups. Unclassified individuals are projected onto those axes with the expectation that they belong to one of the known groups. If they actually belong to a group that was not used to construct the axes, then their projection may fall between the other groups if their unique trait combinations were not part of the DFA parameterization and they share little variation (e.g., Albrecht, 1992). Thus, the unknowns in DFA (red wolves in this case) could appear to be perfectly intermediate between known groups (in this case, coyotes and gray wolves) when they are not. Furthermore, DFA alone cannot determine whether hybridization is likely to produce gradational intermediates or completely distinct ones.
There are several approaches that can supplement DFA to provide greater power for distinguishing between competing interpretations.
A hierarchical analysis of quantitative phenotypic traits (QST) can be used to determine whether historical red wolves in their core range are more divergent from gray wolves than gray wolf subspecies are from one another. This approach can be used with any of the morphological traits described in Box 2-1. Hierarchical analysis can also be used to determine whether the divergence of the captive and managed red wolf populations and the GCC populations from core range red wolves is greater than expected relative to divergence within coyotes and gray wolves, as it might if hybridization has substantially affected their morphology.
Furthermore, QST/FST comparisons (Box 2-2) can be combined with genetic data from the same populations to determine whether the morphological differences among gray wolves, coyotes, and red wolves are the result of selection or drift. Importantly, this approach can help determine whether the morphological divergence of the captive and managed red wolf populations and the GCC populations from historical red wolf populations is greater than expected, given their genetic divergence (which would have to be measured in historical populations with ancient DNA techniques). The selection choices made in the early generations of the captive breeding program, which was intended to recover the red wolf phenotype from the founder population (USFWS, 1990; Phillips et al., 2003), may well have resulted in some unique differentiation that was not present in historical populations. QST/FST will provide an index of whether the morphological effects of the recovery breeding program are proportional to its genetic effects.
Affinities among taxa can also be evaluated by an analysis of the distributions of pairwise morphological distances between samples. If two groups, such as red and gray wolves, are divergent, then the distribution of pairwise distances between them will have a greater mean and variance than the distributions of distances within either taxon. The advantage of using this approach for multivariate phenotypes is that it is based on the complete set of morphological differences rather than on any particular axis as in DFA. This approach can also help assess the degree of morphological continuity between the extant red wolf populations, GCC populations, and pre-1920 red wolves from their core range.
Finally, cluster analysis can be used to recover the hierarchy of morphological similarity that arises from phylogenetic and phylogeographic processes. Cluster analysis shows which populations are most similar, as opposed to DFA, which focuses on differences and thus complements genomic analyses. Because hybridization is likely to play a role in the extant red wolf and GCC populations and perhaps in historical eastern and red wolves, relationships among populations may have a network structure rather than a fully tree-like structure. Maximal clique enumeration networks (Butenko and Wilhelm, 2006) or minimum evolutionary steps networks (White et al., 2010) can easily be adapted to multivariate morphometric data to determine whether morphological similarities are fully branching (as would be expected if taxa have independent branching histories) or reticulating (as would be expected if hybridization influenced their morphology).
Special attention needs to be paid to issues of statistical power and balance of sample size. The number of specimens of historical red wolves is small, and the number of historical eastern grays is very small. Similar to the discriminatory power provided by analyses of full genomes, multivariate analysis of morphology helps counteract the limitations of small sample size, but nevertheless there are limits imposed by the historical samples that may not be possible to overcome. The discussions herein focus on questions and techniques that are the most likely to be fruitful, but they will need to be accompanied by statistical power analyses, rarefaction, and down-sampling strategies to equalize sample sizes, and careful attention will need to be given to standard errors on statistical parameters.
Finding 2-2: Prior analyses of the morphological differentiation of red wolves are comprehensive and their conclusions are well supported, but the kinds of data and analyses that have been applied are limited. Geometric morphometrics can pick up subtle differences in shape that cannot be captured by linear measurements, traits such as teeth and postcrania can capture fine-level genetic differences or ecologically relevant specializations not represented by skull data, and analytical methods such as QST and cluster analysis can provide measures of morphological differentiation that are directly comparable to results from genetic analysis.
Detecting Hybridization from Morphology
Hybridization may require special data to detect. Most authors agree that the small, slender, tawny wolves that Audubon and Bachman (1851) described on the Edwards Plateau were a hybrid swarm formed by an admixture between coyotes and core range red wolves (Young and Goldman, 1944; Nowak, 1979), and some authors argue that red wolves as a whole are the product of hybridization between gray wolves and coyotes (see Chapter 1). Signatures of hybridization can sometimes be detected in morphological traits. Most often studies of morphological hybrids are possible when there is a sharp cline between the two parent populations, such as when a hybrid tension zone exists (Butlin et al., 1991; Gay et al., 2008; Polly et al., 2013). However, the large geographic ranges and dispersal distances of North American canids would at best produce massively wide tension zones and more likely result in widespread hybrid swarms rather than orderly phenotypic clines. Other approaches to hybrid recognition from morphology are needed.
One such approach involves examining the frequencies of abnormal non-metric traits. Ackermann and colleagues have demonstrated by using captive baboons and wild populations of other mammals that frequencies of dental anomalies and unusual skull suture patterns are higher in hybrid individuals (Ackermann, 2010; Ackermann et al., 2006, 2014, 2018). Supernumary teeth, rotated teeth, and atypical skull sutures are more frequent in F1 and F2 hybrid individuals than in the background populations of baboons, and comparative evidence suggests that the same may be true for other mammal groups. How long such high frequencies of abnormal traits persist in hybrid populations is as yet unknown.
Interestingly, Young and Goldman (1944) anecdotally reported seemingly high frequencies of tooth anomalies in pre-1920 red wolf individuals from central Texas and Missouri, both areas where early hybridization swarms are suspected to have occurred, in their description of individual variation in wolves. Given that they examined hundreds of skulls of gray and red wolves from across the whole of North America, it is intriguing that the only reports of such anomalies were ascribed to these populations where there would likely be many F1 hybrid individuals.
This kind of data requires cautious interpretation because elevated frequencies of abnormal metric traits can occur through two other avenues. First, inbreeding can produce elevated frequencies (Berry, 1968; Trinkaus, 1978). Given the small population sizes of putative red wolves and the effects of prior bottlenecks in population size, this is not a negligible possibility. Second, elevated frequencies of abnormal non-metric traits have also been explained by environmental disturbance that leads to developmental perturbations, sometimes through inbreeding (e.g., Trinkaus, 1978; Vasilyev and Vasilyeva, 1995). A systematic study of non-metric trait frequencies may illuminate the history of hybridization, especially if it is supplemented by genomic data.
Finding 2-3: The skull measurements used in past morphological analyses have less power than non-metric traits for detecting hybridization. Anecdotal reports of some historical red wolf samples suggest the presence of hybrid individuals, but no attempt has been made to systematically analyze morphological hybrid signatures in historical red wolf specimens.
Sexual Dimorphism
The degree of sexual dimorphism in a species, especially in body size, is often linked to size-assortative mating, which in turn can affect the likelihood of hybridization between closely related sympatric species (Hinton et al., 2018). The degree of overlap of body size between males and females in sympatric populations of two species may be a predictor of the likelihood of hybridization. This is especially so when there is positive assortative mating by body size in each species, which is the case with both coyotes and red wolves (see Mating Behavior section below). Size dimorphism has the potential to produce asymmetries in hybridization whereby most hybrid individuals are consistently fathered by one species and mothered by the other, thus potentially biasing introgression of mitochondrial haplotypes compared with nuclear genes. Size dimorphism can, in principle, be studied from skulls and dentitions in historical populations using museum specimens (e.g., Gay and Best, 1996; Van Valkenburgh and Sacco, 2002; Gittleman and Van Valkenburgh, 2009).
Finding 2-4: Sexual dimorphism in morphology has been noted in red wolves, but it has neither been systematically evaluated nor used to assess the likelihood of hybridization or of sex-biased introgression processes.
BEHAVIOR AND ECOLOGY
While differences in behavioral traits among groups can offer insight into the evolutionary relationships among those groups, these data require cautious interpretation. One reason is that only some types of behavioral differences, specifically mating behaviors, are directly relevant for understanding evolutionary relationships. Other behaviors, such as non-mating social behavior and foraging tactics, can, in some circumstances, contribute to identifying important distinctions among groups and the level at which those distinctions occur—among populations of the same species or among species—or they might offer insights into evolutionary relationships only when combined with other types of data. A second reason is that many behavioral traits, such as foraging tactics, are sensitive to local ecological circumstances, so only consistent differences among groups across a
range of ecological conditions can reveal genuine differences among species. A third reason is that it is difficult to predict which behaviors, if any, would be characteristic of recent hybrids. Finally, it is impossible to know the behaviors of historical red wolves so, unlike the use of morphology and, potentially, genomics, there is no standard against which data from extant groups can be assessed.
However, given that those data can be helpful in conjunction with other data, the committee reviewed their potential contributions and limitations. But in light of the challenges described above, collecting and assessing behavioral data should be a lower priority than obtaining genomic and morphological data.
Mating Behavior
Differences between groups in their mating behaviors are important because they can contribute to pre-mating isolation—that is, the prevention of cross-species mating—between the groups. Behavior can lower the likelihood of mating between two sympatric groups in three ways. First, two populations may occupy different habitats, and individuals from each population may search for mates only in its own habitat. Second, individuals may have positive assortative mating in which individuals choose mates who are similar to them in a phenotypic trait such as body size. When species differ in body size, positive assortative mating will promote reproductive isolation. Third, the timing of mating and reproduction can differ between populations such that individuals from each population search for mates in different periods.
The mating systems of wolves and coyotes differ in two of these ways. Red wolves have always been reported as favoring forested habitats, whereas coyotes have been reported as preferring open areas with fields and mixed woods (summarized in Waples et al., 2018). Occupying different habitats would contribute to behavioral isolation. In studies of eastern wolves and coyotes, the species occupied different habitats, akin to the distinction between red wolves and coyotes, and hybrids occupied intermediate habitats (Otis et al., 2017). Both species display positive assortative mating based on body size (Hinton et al., 2018); red wolves in North Carolina are larger than coyotes (Hinton and Chamberlain, 2014), which will also contribute to reproductive isolation. Red wolves will aggressively displace coyotes and identified hybrids from their territories (Gese and Terletzky, 2015).
Distinctions in mating behaviors between populations erode when anthropogenic effects have altered habitat structures, local ecology, or the animals’ social systems in such a way that individuals from different species encounter one another in novel habitats or in novel circumstances. For example, normal habitat differences between the sister taxa green treefrogs and barking treefrogs preclude hybridization. However, in altered wetlands both species attempt to breed in the same locations, and hybridization is often the result (Lamb and Avise, 1986, 1987). Hybridization rates among waterfowl species have increased as a result of human habitat alteration (Rhymer and Simberloff, 1996).
Similar effects may be important in the comparison of red wolves and coyotes. The historical habitat separation of these two canids—forests for red wolves and mixed woods and open areas for coyotes—is threatened by increased fragmentation of forest habitat and increased encirclement of forest habitat by large areas of fields and scrub woodlands. This brings individuals from different species into contact with one another at a higher rate than would happen otherwise and promotes hybridization (Hinton et al., 2018). Hybridization may also be promoted by the hunting mortality of one member of a breeding pair during the breeding season, but the evidence for that is equivocal (Hinton et al., 2017b, 2018).
Observations of mating behavior in red wolves and coyotes can establish only whether, in the absence of habitat disruption, individuals from the two groups are likely to be reproductively isolated. Nonetheless, these are critical data because they could identify whether modern hybrid
matings occur only in altered habitats or via the ill-timed loss of a partner. If so, those observations would be consistent with red wolves and coyotes having been distinctive, isolated groups before the advent of coyote expansion and anthropogenic habitat alteration.
Social Structure
Social structure represents another feature that one can examine in order to understand the distinctions between red wolves and coyotes. In particular, the social structure of breeding units can distinguish populations. Many animals have a social structure in which individuals other than parents contribute to raising young. In some groups these “helpers” are close relatives such as siblings, whereas in others they can be distant relatives or unrelated individuals. The presence or absence of helpers and the number of cooperating individuals can reflect an adaptive response to ecological conditions in which parents would otherwise struggle to raise young successfully or an adaptive response to a saturated environment in which the cooperating individuals would struggle to find territories or mates.
Red wolves and coyotes have different social structures, although the social structure is quite variable in coyotes so that structure can be closer to the social structure of red wolves in some cases than in others. Coyotes can live as isolated individuals, as mated pairs, and as members of small packs (Bekoff and Wells, 1982). Coyote groups consist of a mated pair, the young of that year, and, sometimes, young from the previous year (Bowen, 1981; Bekoff and Wells, 1982; Phillips and Henry, 1992). Group sizes average 1.5 individuals in summer and up to 3 individuals in winter (Bowen, 1981). Larger group size facilitates defense of pups from predators (Bekoff and Wells, 1982) and the ability to prey on larger animals such as mule deer (Bowen, 1981). Coyotes typically mature and begin breeding when 1 year old (Phillips et al., 2003).
By contrast, red wolves in the managed population live in packs, which consist of a mated pair, offspring of the year, and offspring from previous years (Phillips and Henry, 1992; Sparkman et al., 2011, 2012). Red wolves do not mature and begin to breed until they are at least 2 years old (Phillips et al., 2003). Pack sizes are therefore larger than in coyotes. Larger packs, which result from the delayed dispersal of previous years’ offspring, are more successful at raising pups (Sparkman et al., 2011). In addition, delayed dispersal has a direct benefit for those offspring in facilitating their own ability to eventually establish a territory and attract a mate (Sparkman et al., 2011, 2012).
Foraging Habit and Prey Choice
The third behavioral feature that could inform a diagnosis about the nature of differences between red wolves and coyotes is foraging habit and prey choice. This behavioral feature reflects the most relevant potential ecological distinction for understanding the differences between the two species. Differences between populations in foraging habit and prey choice can reflect adaptive responses to prey availability, density, and defense. In effect, differences in these behaviors can reflect the occupancy of different ecological niches, which could make the populations distinctive ecological groups and, in several species concepts, unquestionably different species.
There are three challenges to using these behavioral features to distinguish populations for accurate taxonomy. First, there can be variation among conspecific populations in all of the behaviors. This variation can occur on a geographic scale or even on a local scale. When this is so, in order to use these behavioral features to support taxonomic diagnoses, the differences among the populations under scrutiny must be greater than the typical level of variation among conspecific populations.
This is a difficult challenge to answer in the case of red wolves and coyotes. There are few populations of putative red wolves to evaluate for population differences. Moreover, the taxonomic
statuses of newly discovered populations in Missouri or along the Louisiana–Texas border area are themselves unclear. If consistent differences in social structure are observed among these groups, these would be consistent with species distinctions. On the other hand, inconsistent differences might reflect population variation or different taxonomic statuses of different populations.
The second challenge is that variation among populations, whether in general or between particular pairs of populations, may reflect plastic responses to current conditions. For example, individuals from two populations may have different diets based solely on local differences in prey availability, not on any genetically based differences in prey preference (Bowen, 1981; Hinton et al., 2017a). This problem may also occur in a converse situation. A short-term study of diets may indicate that two populations have similar diets simply because it was conducted in a period in which some prey items were especially abundant and individuals from each population capitalized on that abundance (McVey et al., 2013).
One useful approach to this challenge is to ask whether historical red wolves were ecologically distinct from other North American canids, especially where they occurred in sympatry; whether managed red wolves and GCC populations share those distinctions; and whether those distinctions might be reflected in tooth morphology. Today gray wolves are more predacious, favoring larger ungulate prey, and coyotes are more opportunistic feeders on small mammals, larger ungulates, and carrion (Arjo et al., 2002; Bartel and Knowlton, 2005; Morey et al., 2007). The frequency with which teeth encounter bone, gristle, and meat may therefore differ on average between species, and the microscopic wear patterns on teeth may therefore differ with diet. DeSantis and colleagues (2015) found that the patterns of wear on the teeth of coyotes are less consistent and more variable than the patterns in gray wolves. This would be expected from the more varied diet of coyotes; the diet of gray wolves, which is one of mostly soft tissue and some bone, produces a more predictable and consistent pattern of tooth wear. The two species also appear to distribute wear differently along their tooth rows (Tanis et al., 2018). Wear analysis of teeth in museum specimens may therefore offer insights into diets in historical populations.
Another possible approach to this challenge is to use stable isotope data from the canids and their prey (Fox-Dobbs et al., 2012). Stable isotopes can reflect the integration of an individual’s diet over its lifetime and, as a result, show less sensitivity to seasonal or annual variation in prey abundance. Isotope analyses can be performed on collagen samples from older specimens and thus offer potential insight into diets in the past. Values of enriched nitrogen can indicate whether populations occupy different trophic levels, and values of enriched carbon can point to different distributions of prey type if those prey themselves have different foraging habits and diets.
The third challenge is that it is unclear how these behavioral features would appear in recent hybrids. Even if consistent differences between the behaviors of red wolves and coyotes are based in part on genetic differences, there is no a priori reason to expect that hybrids would exhibit some type of intermediate behavior.
This last issue is an acute challenge because the controversy is whether all extant red wolves represent hybrids that are sufficiently recent that red wolves cannot be considered a distinct species. In this sense, there is no standard for comparing coyotes, “genuine” red wolves, and putative hybrids. Without behavioral data on historical red wolves, which are impossible to obtain, it cannot be said that behavior of the extant red wolf populations offers a standard against which, along with behavior of coyotes, the behavior of suspected hybrids can be evaluated.
The Ideal Behavioral Data
In an ideal world, experimentation could distinguish patterns of behavior induced by anthropogenic effects from those found in unaffected habitats. This is not possible for the red wolf–coyote situation. In a world less than ideal but still desirable, behaviors could be compared between these
groups when they are found in unaltered habitat and in altered habitat. This is also not possible for either the extant red wolf populations or the GCC populations. The best that is likely to be possible is the long-term observation of mating patterns, social structure, and prey use patterns to determine whether these southeastern canids occupy different ecological niches and, if so, whether those differences are consistent between different locations in which coyotes come into contact with red wolves. For example, are differences observed in the areas in which the managed red wolves have contact with coyotes seen also in areas occupied by the GCC populations and coyotes?
The large home ranges of carnivores such as these suggest that differentiation among conspecific populations will be low. In this light, it might be useful to compare distinctions among other populations of wolves and coyotes over large geographic areas. Large differences between pairs of wolf and coyote populations in the same area could be a useful diagnostic tool if the typical differences among populations within each species are small in magnitude.
A large challenge for interpreting data on red wolf populations is that the populations are either small (canids in Louisiana and Missouri) or founded by a small number of individuals (i.e., the extant captive breeding colonies and the managed red wolves in North Carolina). Genetic drift of such small, or bottlenecked, populations will quickly result in large genetic differences among populations of the same species. Large distinctions among populations of the same taxon make it difficult to interpret, unambiguously, apparent differences between putatively different taxa.
In that light, it would be informative to conduct long-term behavioral observations on animals from whom tissue samples can be obtained for genomic analyses and from whose photographic images key morphological data can be collected. The joint distribution of differences in these three arenas—genomic, morphological, and behavioral—can inform the interpretation of the entire set of data as well as provide valuable calibration for considering the interpretation of one or two of these types of data in other taxa.
Finding 2-5: Behavioral differences between species can affect the likelihood of hybridization, choice of habitats, and choice of prey. Furthermore, these behaviors may depend on the environmental context in which an individual finds itself and thus be plastically modified as habitats change. Much has been recorded about red wolf behavior, but most of it has been based on individuals in the hybrid swarm of east Texas in the 1960s and in the managed population, neither of which may be representative of historical red wolf behaviors.
CONCLUSIONS AND RECOMMENDATIONS
Conclusion 2-1: To distinguish among the origin hypotheses for the red wolf using morphology, additional kinds of data and analysis are required. The existing studies based on skulls can be supplemented by similar analyses of teeth, which are likely to better capture genetic relationships among taxa, and postcrania, which are likely to supplement the understanding of the ecological differentiation of red wolves from coyotes and gray wolves. Geometric morphometric data will enable finer discrimination based on details of shape that were not captured in the original analyses.
Conclusion 2-2: New analytical techniques are required to assess the congruence between morphological and genetic evidence for the distinctiveness of the red wolf. Prior analyses relied solely on discriminant function analysis, which is a powerful tool for determining what is different between known groups, but it less powerful for determining which groups are more similar, how variation at the subspecific level compares to differentiation between species, or whether differences are due to hybridization. The same hierarchical analyses of variance, cluster analyses, and pairwise distance analyses that are commonly used on genetic data can be applied to morphological data in order to facilitate these comparisons.
Conclusion 2-3: The behavioral data that exist appear to be consistent with the hypothesis that hybridization happened late in the red wolf’s history, not at its inception, but behavioral data for historical red wolves are sparse. The behavioral data that exist deserve to be more comprehensively analyzed with regard to hypotheses about hybridization in canids, and morphological indicators of behavior are needed to supplement direct observations of behavior.
Recommendation 2-1: The committee recommends that existing morphological data sets (e.g., Young and Goldman, 1944; Nowak 1979) be submitted to new analytical techniques that are directly comparable to genetic results, that ancient DNA sequencing and morphological analyses be performed on the same individuals, and that new morphological data be collected that are relevant to assessing the ecological and behavioral distinctiveness of historical red wolves in their core range.
REFERENCES
Ackermann, R. R. 2010. Phenotype traits of primate hybrids: Recognizing admixture in the fossil record. Evolutionary Anthropology 19:258–270.
Ackermann, R. R., J. Rogers, and J. Cheverud. 2006. Identifying the morphological signatures of hybridization in primate and human evolution. Evolution 64:271–290.
Ackermann, R. R., L. Schroeder, J. Rogers, and J. M. Cheverud. 2014. Further evidence for phenotypic signatures of hybridization in descendant baboon populations. Journal of Human Evolution 76:54–62.
Ackermann, R. R., M. L. Arnold, M. D. Baiz, J. A. Cahill, L. Cortés-Ortiz, B. J. Evans, B. R. Grant, P. R. Grant, B. Hallgrimsson, R. A. Humphreys, C. J. Jolly, J. Malukiewicz, C. J. Percival, T. B. Ritzman, C. Roos, C. C. Roseman, L. Schroeder, F. H. Smith, K. A. Warren, R. K. Wayne, and D. Zinner. 2018. Hybridization in human evolution: Insights from other organisms. Evolutionary Anthropology 28:189–209.
Albrecht, G. H. 1980. Multivariate analysis and the study of form, with special reference to canonical variate analysis. American Zoologist 20:679–693.
Arjo, W. M., D. H. Pletscher, and R. R. Ream. 2002. Dietary overlap between wolves and coyotes in northwestern Montana. Journal of Mammalogy 83:754–766.
Atchley, W. R., and B. K. Hall. 1991. A model for development and evolution of complex morphological structures. Biological Reviews 66:101–157.
Audubon, J. J., and J. Bachman. 1851. The quadrupeds of North America, Vol. 2. New York: V. G. Audubon.
Bartel, R. A., and F. F. Knowlton. 2005. Functional feeding responses of coyotes, Canis latrans, to fluctuating prey abundance in the Curlew Valley, Utah, 1977–1993. Canadian Journal of Zoology 83:569–578.
Bekoff, M., and M. C. Wells. 1982. Behavioral ecology of coyotes—Social organization, rearing patterns, space use, and resource defense. Zeitschrift Fur Tierpsychologie-Journal of Comparative Ethology 60:281–305.
Bell, C. J., J. A. Gauthier, and G. S. Bever. 2010. Covert biases, circularity, and apomorphies: A critical look at the North American Quaternary herpetofaunal stability hypothesis. Quaternary International, 217:30–36.
Berry, R. J. 1968. The biology of non-metrical variation in mice and men. In D. R. Brothwell (ed.), The skeletal biology of earlier human populations. London: Pergamon Press. Pp. 103–133.
Bookstein, F. L. 1992. Morphometric tools for landmark data: Geometry and biology. Cambridge, UK: Cambridge University Press.
Bowen, W. D. 1981. Variation in coyote social organization—The influence of prey size. Canadian Journal of Zoology 59:639–652.
Boyer, D. M., J. Puente, J. T. Gladman, C. Glynn, S. Mukherjee, G. S. Yapuncich, and I. Daubechies. 2015. A new fully automated approach for aligning and comparing shapes. Anatomical Record 298(1):249–276.
Butenko, S., and W. E. Wilhelm. 2006. Clique-detection models in computational biochemistry and genomics. European Journal of Operational Research 173:1–17
Butlin, R. K., M. G. Ritchie, and G. M. Hewitt. 1991. Comparisons among morphological characters and between localities in the Chorthippus parallelus hybrid zone (Orthoptera, Acrididae). Philosophical Transactions of the Royal Society of London B 334:297–308.
Cardini, A. 2019. Integration and modularity in Procrustes shape data: Is there a risk of spurious results? Evolutionary Biology 46:90–105.
Caumul, R., and P. D. Polly. 2005. Phylogenetic and environmental components of morphological variation: Skull, mandible and molar shape in marmots (Marmota, Rodentia). Evolution 59:2460–2472.
Chambers, S. M., S. R. Fain, B. Fazio, and M. Amaral. 2012. An account of the taxonomy of North American wolves from morphological and genetic analyses. North American Fauna 77:1–67.
Churcher, C. S., P. W. Parmalee, G. L. Bell and J. P. Lamb, J. 1989. Caribou from the late Pleistocene of northwestern Alabama. Canadian Journal of Zoology 67:1210–1216.
DeSantis, L. R., B. W. Schubert, E. Schmitt-Linville, and P. S. Ungar, S. L. Donohue, and R. J. Haupt. 2015. Dental microwear textures of carnivorans from the La Brea Tar Pits, California, and potential extinction implications. Contributions in Science, Los Angeles County Museum of Natural History 42:37–52.
Dryden, I., and K. V. Mardia. 1998. Statistical shape analysis. New York: Wiley.
Dumont, E. R., J. Piccirillo, and I. R. Grosse. 2005. Finite element analysis of biting behavior and bone stress in the facial skeletons of bats. Anatomical Record 283:319–330.
Evans, A. R., G. P. Wilson, M. Fortelius, and J. Jernvall. 2007. High-level similarity of dentitions in carnivorans and rodents. Nature 445:78–81.
Felsenstein, J. 1973. Maximum-likelihood estimation of evolutionary trees from continuous characters. American Journal of Human Genetics 25:471–492.
Felsenstein, J. 1988. Phylogenies and quantitative characters. Annual Review of Ecology and Systematics 19:445–471.
Fox-Dobbs, K., A. A. Nelson, P. L. Koch, and J. A. Leonard. 2012. Faunal isotope records reveal trophic and nutrient dynamics in twentieth century Yellowstone grasslands. Biology Letters, 8(5), 838–841.
Gambaryan, P. P. 1974. How mammals run. New York: John Wiley & Sons.
Gay, L., P. A. Crochet, D. A. Bell, and T. Lenormand. 2008. Comparing clines on molecular and phenotypic traits in hybrid zones: a window on tension zone models. Evolution 62:2789–2806.
Gay, S. W., and T. L. Best. 1996. Relationships between abiotic variables and geographic variation in skulls of pumas (Puma concolor: Mammalia, Felidae) in North and South America. Zoological Journal of the Linnean Society 117:259–282.
Gese, E. M., and P. A. Terletzky. 2015. Using the “placeholder” concept to reduce genetic introgression of an endangered carnivore. Biological Conservation 192:11–19.
Gese, E. M., J. L. Dowd, and L. M. Aubry. 2013. The influence of snowmobile trails on coyote movements during winter in high-elevation landscapes. PLOS ONE 8(12):e82862.
Gittleman, J. L., and B. Van Valkenburgh. 2009. Sexual dimorphism in the canines and skulls of carnivores: Effects of size, phylogeny, and behavioural ecology. Journal of Zoology 242:97–117.
Gómez-Robles, A., M. Martinón-Torres, J. B. De Castro, A. Margvelashvili, M. Bastir, J. L. Arsuaga, A. Pérez-Pérez, F. Estebaranz, and L. M. Martínez. 2007. A geometric morphometric analysis of hominin upper first molar shape. Journal of Human Evolution 53:272–285.
Graham, R. W., E. L. Lundelius, M. A. Graham, E. K. Schroeder, R. S. Toomey, E. Anderson, A. D. Barnosky, J. A. Burns, C.S. Churcher, D. K. Grayson and R. D. Guthrie. 1996. Spatial response of mammals to late Quaternary environmental fluctuations. Science 272:1601–1606.
Gunz, P., P. Mitteroecker, and F. L. Bookstein. 2005. Semilandmarks in three dimensions. In D. J. Slice (ed.), Modern morphometrics in physical anthropology. Boston: Springer. Pp. 73–98.
Hammer, Ø., and D. A. T. Harper. 2006. Palaeontological data analysis. Oxford, UK: Blackwell Publishing.
Hanken, J., and B. K. Hall. 1993. The skull. Chicago: University of Chicago Press
Hinton, J. W., and M. J. Chamberlain. 2014. Morphometrics of Canis taxa in eastern North Carolina. Journal of Mammalogy 95:855–861.
Hinton, J. W., A. K. Ashley, J. A. Dellinger, J. L. Gittleman, F. T. van Manen, and M. J. Chamberlain. 2017a. Using diets of Canis breeding pairs to assess resource partitioning between sympatric red wolves and coyotes. Journal of Mammalogy 98:475–488.
Hinton, J. W., K. E. Brzeski, D. R. Rabon, and M. J. Chamberlain. 2017b. Effects of anthropogenic mortality on critically endangered red wolf Canis rufus breeding pairs: Implications for red wolf recovery. Oryx 51:174–181.
Hinton, J. W., J. L. Gittleman, F. T. van Manen, and M. J. Chamberlain. 2018. Size-assortative choice and mate availability influences hybridization between red wolves (Canis rufus) and coyotes (Canis latrans). Ecology and Evolution 8:3927–3940.
Hlusko, L. J., M. L. Maas, and M. C. Mahaney. 2004. Statistical genetics of molar cusp patterning in pedigreed baboons: Implications for primate dental development and evolution. Journal of Experimental Biology B 302B:268–283.
Jernvall, J. 1995. Mammalian molar cusp patterns: Developmental mechanisms of diversity. Acta Zoologica Fennica 198:1–61.
Lamb, T., and J. C. Avise. 1986. Directional introgression of mitochondrial DNA in a hybrid population of tree frogs—The influence of mating-behavior. Proceedings of the National Academy of Sciences 83:2526–2530.
Lamb, T., and J. C. Avise. 1987. Morphological variability in genetically defined categories of anuran hybrids. Evolution 41:157–165.
Lande, R. 1992. Neutral theory of quantitative genetic variance in an island model with local extinction and colonization. Evolution 46:381–389.
Lele, S. 1991. Some comments on coordinate free and scale invariant methods in morphometrics. American Journal of Physical Anthropology 85:407–417.
Lyons, S. K., 2003. A quantitative assessment of the range shifts of Pleistocene mammals. Journal of Mammalogy 84:385–402.
Maga, A. M., N. Navarro, M. L. Cunningham, and T. C. Cox. 2015. Quantitative trait loci affecting the 3D skull shape and size in mouse and prioritization of candidate genes in-silico. Frontiers in Physiology 6:92.
McVey, J. M., D. T. Cobb, R. A. Powell, M. K. Stoskopf, J. H. Bobling, L. P. Waits, and C. E. Moorman. 2013. Diets of sympatric red wolves and coyotes in northeastern North Carolina. Journal of Mammalogy 94:1141–1148.
Meachen, J. A., and J. X. Samuels. 2012. Evolution in coyotes (Canis latrans) in response to the megafaunal extinctions. Proceedings of the National Academy of Sciences 109:4191–4196.
Meachen, J. A., A. C. Janowicz, J. E. Avery, and R. W. Sadleir. 2014. Ecological changes in coyotes (Canis latrans) in response to the Ice Age megafaunal extinctions. PLOS ONE 9(12):e116041.
Morey, P. S., E. M. Gese, and S. Gerht. 2007. Spatial and temporal variation in the diet of coyotes in the Chicago metropolitan area. American Midland Naturalist 158:147–161.
Murray, D. L., and S. Boutin. 1991. The influence of snow on lynx and coyote movements: Does morphology affect behavior? Oecologia 88:463–469.
Murray, D. L., and S. Larivière. 2002. The relationship between foot size of wild canids and regional snow conditions: Evidence for selection against a high footload? Journal of Zoology 256:289–299.
NASEM (National Academies of Sciences, Engineering, and Medicine). 2019. Evaluating the taxonomic status of the Mexican gray wolf and the red wolf. Washington, DC: The National Academies Press. https://doi.org/10.17226/25351.
Nowak, R. M. 1979. North American Quaternary Canis. Monographs of the Museum of Natural History, University of Kansas 6:1–154.
Nowak, R. M. 2002. The original status of wolves in eastern North America. Southwestern Naturalist 1:95–130.
Otis, J. A., D. Thornton, L. Rutledge, and D. L. Murray. 2017. Ecological niche differentiation across a wolf–coyote hybrid zone in eastern North America. Diversity and Distributions 23:529–539.
Phillips, M. K., and V. G. Henry. 1992. Comments on red wolf taxonomy. Conservation Biology 6:596–599.
Phillips, M. K., V. G. Henry, and B. T. Kelly. 2003. Restoration of the red wolf. In L. D. Mech and L. Boitani (eds.), Wolves: Behavior, ecology, and conservation. Chicago, IL: University of Chicago Press. Pp. 272–288.
Polly, P. D. 2003. Paleophylogeography: The tempo of geographic differentiation in marmots (Marmota). Journal of Mammalogy 84:369–384.
Polly, P. D. 2007a. Chapter 15: Limbs in mammalian evolution. In B. K. Hall (ed.), Fins into limbs: Evolution, development, and transformation. Chicago: University of Chicago Press. Pp. 245–268.
Polly, P. D. 2007b. Phylogeographic differentiation in Sorex araneus: Morphology in relation to geography and karyotype. Russian Journal of Theriology 6:73–84.
Polly, P. D. 2008. Adaptive zones and the pinniped ankle: A 3D quantitative analysis of carnivoran tarsal evolution. In E. Sargis and M. Dagosto (eds.), Mammalian evolutionary morphology: A tribute to Frederick S. Szalay. Dordrecht, The Netherlands: Springer. Pp. 165–194.
Polly, P. D. 2010. Tiptoeing through the trophics: Geographic variation in carnivoran locomotor ecomorphology in relation to environment. In A. Goswami and A. Friscia (eds.), Carnivoran evolution: New views on phylogeny, form, and function. Chicago: University of Chicago Press. Pp. 374–410.
Polly, P. D. 2019. Geometric morphometric tests for phenotypic divergence between chromosomal races. In J. B. Searle, J. Zima, and P. D. Polly (eds.), Shrews, chromosomes and speciation. Cambridge, UK: Cambridge University Press. Pp. 336–364.
Polly, P. D., A. V. Polyakov, V. B. Ilyashenko, S. S. Onischenko, T. A. White, N. A. Shchipanov, N. S. Bulatova, S. Pavlova, P. M. Borodin, and J. B. Searle. 2013. Phenotypic variation across chromosomal hybrid zones of the common shrew (Sorex araneus) indicates reduced gene flow. PLOS ONE 8(7):e67455.
Polly, P. D., J. Fuentes–Gonzales, A. M. Lawing, A. K. Bormet, and R. G. Dundas. 2017. Clade sorting has a greater effect than local adaptation on ecometric patterns in Carnivora. Evolutionary Ecology Research 18:61–95.
Reyment, R. A., R. E. Blackith, and N. A. Campbell. 1984. Multivariate morphometrics. New York: Academic Press.
Rhymer, J. M., and D. Simberloff. 1996. Extinction by hybridization and introgression. Annual Review of Ecology and Systematics 27:83–109.
Sokal, R. R., and F. J. Rohlf. 1994. Biometry: The principles and practices of statistics in biological research. New York: W.H. Freeman.
Sparkman, A. M., J. R. Adams, T. D. Steury, L. P. Waits, and D. L. Murray. 2011. Direct fitness benefits of delayed dispersal in the cooperatively breeding red wolf (Canis rufus). Behavioral Ecology 22:199–205.
Sparkman, A. M., J. R. Adams, T. D. Steury, L. P. Waits, and D. L. Murray. 2012. Pack social dynamics and inbreeding avoidance in the cooperatively breeding red wolf. Behavioral Ecology 23:1186–1194.
Spitze, K. 1993. Population structure in Daphnia obtusa: Quantitative genetic and allozymic variation. Genetics 135:367–374.
Tajima, F. 1983. Evolutionary relationship of DNA sequences in finite populations. Genetics 105:437–460.
Tanis, B. P., L. R. DeSantis, and R. C. Terry. 2018. Dental microwear textures across cheek teeth in canids: Implications for dietary studies of extant and extinct canids. Palaeogeography, Palaeoclimatology, Palaeoecology 508:129–138.
Tatsuoka, M. M. 1971. Multivariate analysis: Techniques for educational and psychological research. New York: Wiley and Sons.
Thesleff, I. 2003. Developmental biology and building a tooth. Quintessence International 34:613–620.
Tomiya, S., and J. A. Meachen. 2018. Postcranial diversity and recent ecomorphic impoverishment of North American gray wolves. Biology Letters 14:20170613.
Trinkaus, E. 1978. Bilateral asymmetry of human skeletal non metric traits. American Journal of Physical Anthropology 49:315–318.
Ungar, P. S. 2010. Mammal teeth: Origin, evolution, and diversity. Baltimore, MD: Johns Hopkins University Press.
USFWS (U.S. Fish and Wildlife Service). 1990. Red wolf recovery/species survival plan. Atlanta, GA: U.S. Fish and Wildlife Service.
Van Valkenburgh, B. 2009. Costs of carnivory: Tooth fracture in Pleistocene and recent carnivorans. Biological Journal of the Linnean Society, 96:68–81.
Van Valkenburgh, B., and T. Sacco. 2002. Sexual dimorphism, social behavior, and intrasexual competition in large Pleistocene carnivorans. Journal of Vertebrate Paleontology 22:164–169.
Vasilyev, A. G., and I. A. Vasilyeva. 1995. Non-metric variation in red vole populations within the East Ural Radioactive Track (EURT) zone. Acta Theriologica 40:55–64.
Vitek, N. S., C. L. Manz, T. Gao, J. I. Bloch, S. G. Strait, and D. M. Boyer. 2017. Semi supervised determination of pseudocryptic morphotypes using observer free characterizations of anatomical alignment and shape. Ecology and Evolution 7:5041–5055.
VonHoldt, B. M., J. A. Cahill, Z. Fan, I. Gronau, J. Robinson, J. P. Pollinger, B. Shapiro, J. Wall, and R. K. Wayne. 2016. Whole-genome sequence analysis shows that two endemic species of North American wolf are admixtures of the coyote and gray wolf. Science Advances 2:e1501714.
Wainwright, P. C., M. E. Alfaro, D. I. Bolnick, and C. D. Hulsey. 2005. Many-to-one mapping of form to function: A general principle in organismal design? Integrative and Comparative Biology 45:256–262.
Wakeley, J. 1996a. Distinguishing migration from isolation using the variance of pairwise distances. Theoretical Population Biology 49:369–386.
Wakeley, J. 1996b. The variance of pairwise nucleotide differences in two populations with migration. Theoretical Population Biology 49:39–57.
Waples, R. S., R. Kays, R. J. Fredrickson, K. Pacifici, and L. S. Mills. 2018. Is the red wolf a listable unit under the U.S. Endangered Species Act? Journal of Heredity 109:585–597.
Wayne, R. K. 1992. On the use of morphologic and molecular genetic characters to investigate species status. Conservation Biology 6:590–592.
Wayne, R. K., N. Lehman, and T. K. Fuller. 1995. Conservation genetics of the gray wolf. In L. N. Carbyn, S. H. Fritts, and D. R. Seip (eds.), Ecology and conservation of wolves in a changing world. Edmonton, AB: Canadian Circumpolar Institute. Pp. 399–408.
White, T. A., M. Bordewich, and J. B. Searle. 2010. A network approach to study karyotypic evolution: The chromosomal races of the common shrew (Sorex araneus) and house mouse (Mus musculus) as model systems. Systematic Biology 59:262–276.
Whitlock, M. C., 2008. Evolutionary inference from QST. Molecular Ecology 17:1885–1896.
Wilson, P. J., S. Grewal, I. D. Lawford, J. N. M. Heal, A. G. Granacki, D. Pennock, J. B. Theberge, M. T. Theberge, D. R. Voigt, W. Waddell, R. E. Chambers, P. C. Paquet, G. Goulet, D. Cluff, and B. N. White. 2000. DNA profiles of the eastern Canadian wolf and the red wolf provide evidence for a common evolutionary history independent of the gray wolf. Canadian Journal of Zoology 78:2156–2166.
Wójcik, A. M., P. D. Polly, M. D. Sikorski, and J. M. Wójcik. 2006. Selection in a cycling population: Differential response among skeletal traits. Evolution 60:1925–1935.
Wright, S. 1951. The genetic structure of populations. Annals of Eugenics 15:323–354.
Young, S. P., and E. A. Goldman. 1944. The wolves of North America. Washington, DC: American Wildlife Institute.
This page intentionally left blank.


















