Proceedings of a Workshop
| IN BRIEF | |
May 2021 |
DATA IN MOTION: NEW APPROACHES TO ADVANCING SCIENTIFIC, ENGINEERING, AND MEDICAL PROGRESS
Proceedings of a Workshop—in Brief
The movement toward open science, data sharing, and increased transparency is being propelled by the need to rapidly address critical scientific challenges, such as the global coronavirus (COVID-19) public health crisis. This movement has supported growth in fields, such as artificial intelligence (AI), which has demonstrated potential to accelerate science, engineering, and medicine in new and exciting ways. To further advance innovation around these new approaches, the National Academies of Sciences, Engineering, and Medicine’s Board on Research Data and Information (BRDI) convened a public virtual workshop on October 14-15, 2020, to address how researchers in different domains are utilizing data that undergo repeated processing, often in real-time, to accelerate scientific discovery. Although these topics were not originally part of the workshop, the impact of COVID-19 prompted the planning committee to add sessions on early career researchers’ perspectives, as well as rapid review and publishing activities as a result of the pandemic. Workshop discussions, summarized below, also explored the advances needed to enable future progress in areas such as AI, cyberinfrastructure, standards, and policies.
WELCOME AND GOALS FOR THE WORKSHOP
James Hendler, Rensselaer Polytechnic Institute, Committee Chair, welcomed participants and opened the session by describing the purpose of the workshop, to accelerate science through the use of information and modern technologies. Hendler noted that the workshop discussions were designed to address issues such as the need for fast response science as well as the scientific infrastructure to support this effort.
The Need for Fast Response Science: Covid-19 and Other Challenges and Drivers
Stuart Feldman, Schmidt Futures, moderated the first panel focusing on key challenges in supporting the acceleration of science, including around the current COVID-19 public health crisis.
David Haussler, University of California, Santa Cruz (UCSC), discussed current obstacles to the flow of scientific data, particularly around the ownership of information in the life sciences, noting that “we won’t be able to get the large data sets and the AI enabled analysis of them unless we solve this problem.” Haussler provided several examples from his work over the past 20 years where the proprietary nature of data has hindered scientific discovery. He described current research in the genetic sequencing of cancer and other human diseases, through such initiatives as the Cancer Genome Atlas, the International Cancer Genome Consortium, and the TopMed project of National Institutes of Health’s (NIH’s) Heart, Lung and Blood Institute. He also described the original UCSC Cancer Genomics Hub (CGHub), which produced the first petabyte-scale international genome sharing. His other efforts related to addressing genetic data sharing include co-founding the Global Alliance for Genomics and Health, co-initiating a project called the BRCA Exchange, and building the Human Genome Browser, a way to collect all information freely available for interactive exploration about the human genome.
While there has been some level of support for developing a public cancer gene trust which would include
![]()
information about cancer genetic variants, treatments and outcomes, this free exchange of data has never taken hold, significantly limiting scientific advancement in this area, Haussler stated. This is also a challenge facing current efforts to build SARS-CoV-2 genome analysis tools, using the greater than 500,000 SARS CoV-2 genomes that have been collected globally. Genome data support scientific understanding of the genetic evolution of the virus in real time, allowing researchers to trace its movements from place to place, and watch for mutations that may cause resistance or intractability to certain therapies or diagnostics. Even here, where rapid response is so essential, arguments about ownership of the data are inhibiting the speedy development of new software tools. Haussler added that data needed to address the health of humanity and our ecosystems must remain open to ensure we can overcome the challenges we face.
Ana Bonaca, Harvard University, discussed how open data and astronomy are enabling new discoveries about the universe, including in research examining the motion of stars. Ninety percent of the Milky Way is made up of dark matter and dark energy, and scientists do not fully understand how these are distributed. Researchers have been continually observing the night sky, measuring the colors and positions of stars over nearly a year and reviewing the resulting images after 22 months of operation. Through this work, Bonaca noted that there is incredible potential to further new discoveries in the field. Recent breakthroughs in research have been enabled by initial investments in basic science and in ensuring the openness of data, which, as Bonaca noted, can be communicated and utilized only a few hours after they have been generated.
Mark Zelinka, Lawrence Livermore National Laboratory, discussed work to compare results from recent global climate model simulations to determine why these are more sensitive to carbon dioxide increases than their predecessors. Early in 2019, several models simulated values that exceeded the upper limit of the range of scientific consensus about climate sensitivity (the amount of warming that will result from a doubling of the earth’s pre-industrial CO2 levels), which ranges between 1.5 and 4.5 degrees Celsius. These new models indicated that there was a possibility of a five to five and a half degree Celsius warming of the planet. To further establish whether this was a robust signal shared by many models and to get at its root cause, Zelinka worked with colleagues to systematically compare results from dozens of climate model simulations performed at modeling centers around the world. They confirmed that many state-of-the-art models exhibit higher sensitivities than their predecessors, primarily due to differences in modelling how clouds respond to—and thereby amplify—warming. While there is evidence that this is partially due to improvements in how clouds are stimulated in the models, the scientific community is still working to understand these results. A key point is that access to a large ensemble of simulation output from independent climate models is essential for extracting important climate change signals. So how is this facilitated?
The international climate community has—over the course of decades—built an infrastructure that allows investigators to freely access and analyze simulation output from a set of coordinated experiments using more than 100 climate models from 51 institutions in near-real time as results are produced. The infrastructure includes an agreed-upon set of standard experiments that are performed regularly; a standard set of inputs for these experiments; data standards and controlled vocabularies that govern data structure, metadata, nomenclature, formats, etc.; software that checks and prepares output; and a de-centralized federated system for distributing data freely. Figure 1 shows modeling centers contributing to the Coupled Model Intercomparison Project and data nodes available for accessing climate data from the Earth System Grid Federation.
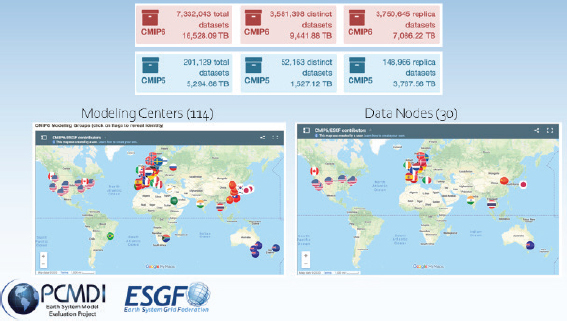
SOURCE: Mark Zelinka, presentation, October 14, 2020. Images from http://esgf-ui.cmcc.it/esgfdashboard-ui/federated-view.html and https://pcmdi.llnl.gov/CMIP6.
Discussion
During the Q&A session, panelists discussed the quality and validity of rapidly produced data compared to data vetted through traditional publication processes. In their experience, Haussler and Bonaca stated that data that are rapidly produced and shared tend to be heavily vetted, often by numerous reviewers, and are generally of high quality. In fact, Bonaca added that her field has developed gold standards related to curating data sets which could be utilized more broadly to support the use of high quality, quickly accessible data. Zelinka noted that the climate community has developed internationally coordinated processes to support vetting data to ensure high quality. Haussler stated that the genome revolution is pushing the concept of openness, serving as a lever to create a movement toward more open data sharing for the rest of the life sciences. The COVID-19 pandemic has reinforced this need for broad sharing of human health data.
IDENTIFYING AND SECURING THE NEEDED SCIENTIFIC INFRASTRUCTURE
Salman Habib, Argonne National Laboratory, moderated a panel discussion focused on identifying and securing the needed scientific infrastructure to support rapid scientific advancement.
Nicholas Schwarz, Argonne National Laboratory, discussed his work related to x-ray light sources, indispensable tools in the exploration of matter. The scientific user facilities of these light sources serve thousands of users every year. These light sources are also undergoing upgrades that will, along with advances in detector data rates, highlight the potential for scientific opportunities in light source science. However, Schwarz noted that addressing some of the most pressing scientific challenges through these light sources requires coupling the intrinsic capabilities of the facilities with advanced data management and analysis. Real time data analysis is required to make decisions.
Over the next ten years, U.S. light sources are projected to generate about 1 exabyte of data per year requiring tens of petaflops to one exaflop of peak computing power. Schwarz added that to handle these data, networks are needed that connect a robust software infrastructure to routinely analyze and process data in real time. These advances will require a seamlessly connected infrastructure. Machine learning can be used to increase performance, opening the ability to compute at the edge, or close to where the information is produced or used. We need to look at data as living and continually evolving as part of this process, stated Schwarz.
Peter Fox,1 Rensselaer Polytechnic Institute, discussed reexamining science infrastructures in Earth and space science. Based on high-dimensional data, we currently have over 5,600 discovered datasets on minerals that have existed for over 4.5 billion years, and the need is highlighted for a scientific infrastructure that allows for dimensional integration of data across space and time. Tools that exist to capture these data through spreadsheets, tables, e.g., Python, R, have not been able to sufficiently render the needed graphs or extract data into a network supportive of scientific discovery. However, Fox noted that there have been advances in scientific data visualization to address these challenges.2 For example, to advance discovery, there is a need to exploit patterns in network subcommunities, hubs and other global and local metrics. Data visualization tools that use Jupyter notebooks and other tools have produced lower-dimensional renderings that contribute to the scientific insight gained from minerals in Earth and planetary bodies as well as other areas.

SOURCE: Andrew Connolly, presentation, October 14, 2020.
___________
1 Sadly, Dr. Peter Fox died on March 27, 2021. Our deepest thoughts are with his family and friends. See https://president.rpi.edu/news/memo/03/28/2021/passing-dr-peter-fox.
2 Fox cited two papers: Fox, P., and J. Hendler. 2014. The Science of Big Data. Big Data 2(2):68-70; and Fox, P., and J. Hendler. 2011. Changing the equation on scientific data visualization. Science 331(6018):705-708.
Through this work, Fox noted there is a move toward representing these new scientific results in a more interactive way where the data are always in motion.
Andrew Connolly, University of Washington, discussed the scientific infrastructure needed to achieve research goals in astronomy over the next decade (Figure 2 shows astronomical surveys over this period). Many use cases in the field require additional existing data to classify and annotate sources and enable real time processing of the data. Scaling science for large volumes of data is also a key challenge and requires more than just scaling hardware, said Connolly. Existing tools may not be robust enough to scale for today’s analytical needs. In high energy physics, analysis codes that individual scientists write or use to ask specific questions can be three to four times less computationally efficient than software tailored to the processing of the data, which hinders effective use of large datasets.
Connolly discussed the ecosystem for scientific computing infrastructure for the coming decade, noting that this will include extending existing analysis frameworks and infrastructure to meet research requirements. A cloud ecosystem for science is needed, but developing applications that work with the volume of data generated from new experiments is not something that individual researchers can easily accomplish. Development of analysis tools and frameworks should be coordinated within and potentially across science domains. In order for research communities to move into these new domains and make effective use of these analysis frameworks, training will be needed at the graduate student level and continuing across the full career trajectories of researchers.
Discussion
During the Q&A session, panelists discussed efforts to develop data visualizations for large data streams, including whether these can be made available to citizen scientists. Connolly said that large data poses computational challenges in terms of both its volume and dimensionality. The re-rendering of data returned to the individual user can support visualization of large, complex data sets. How data are represented in the lower dimensional space is a key challenge, offering an opportunity to connect dimensionality reduction techniques with visualization tools. Schwarz added that there is also an opportunity for machine learning to augment data visualization. Habib added that the data visualization community needs to connect more closely with data intensive science to bridge this gap.
Panelists discussed data preservation efforts and archiving, noting that there has been some progress in this area in some fields. However, as Connolly noted, a key challenge with data preservation is obtaining and maintaining funding to support these efforts. The scientific community needs to recognize the importance of archiving and data preservation; the computing and data infrastructure needed to support this effort “has to be elevated to a first-class citizen,” stated Schwarz.
YOUNG RESEARCHER PERSPECTIVES
Feldman moderated a panel with researchers from the Schmidt Futures fellowship program, a competitive program offering postdoctoral experiences to recent Ph.D. graduates in fields different from their existing expertise.
Amy Shepherd, Schmidt Science Fellow, provided an overview of her background and experience with the program. Initially from New Zealand, she completed her Ph.D. in behavioral neuroscience in 2019 at the University of Melbourne in Australia. Currently, Shepherd is a postdoctoral research fellow at Boston Children’s Hospital where she is examining how the nervous system, the immune system, and the gastrointestinal tract modulate each other. Like many other young scientists, Shepherd noted that her research was significantly impacted by the COVID-19 pandemic as she was no longer able to gain access to the lab to perform experiments. She pointed out that evaluation of postdocs and other early career faculty should take the impacts of the pandemic into account. As a scientific community, Shepherd added, there is a need to more carefully examine how we judge the quality of science and scientists. Relying on the impact factor of the scientific journals in which scientists publish is no longer a strong and reliable measure for judging performance, particularly given the limitations that COVID-19 has placed in terms of research in the lab. This will likely have long term impacts on science.
Shepherd added that lottery grants may serve as a mechanism for supporting early career researchers during this time. These grants, like Explorer grants used in New Zealand, require researchers to develop short grant proposals which are sorted into “declined” or “suitable for funding” categories by an expert panel. Funding is then allocated based on a lottery system, serving as a more efficient mechanism to support early career researchers affected by this crisis.
Ina Anreiter, Schmidt Science Fellow, Stanford University, completed her bachelor’s in biology and a master’s in molecular genetics at the University of Lisbon in Portugal. She completed her Ph.D. in behavioral genetics at the University of Toronto and became a Schmidt Science Fellow where she studied computational biology for a year. She is
also currently a fellow at Stanford University. Anreiter noted that when the COVID-19 pandemic hit, she was conducting several crucial experiments, which then had to be halted due to lack of access to the lab. Echoing Shepherd’s comments, she added that early career scientists have been particularly hard hit by the crisis. The pandemic has also limited opportunities for early career researchers to present at conferences. While the virtual conference format has allowed for greater diversity in terms of international participants, it has also limited networking opportunities. Anreiter noted that the research community should consider building conferences with both in-person and virtual components. She added that the move toward pre-print publications, while not a substitute for peer review, may be one mechanism for improving how science is disseminated, ultimately increasing opportunities for early career researchers. Postdoctoral funding opportunities are also limited, posing a significant barrier. “We should give postdocs the means to work independently” by supporting an allocation of independent research funds directed to early career scientists, stated Anreiter.
Evan Zhao, Schmidt Science Fellow, stated that he received his undergraduate degree at California Institute of Technology and Ph.D. at Princeton University in chemical engineering. Currently, he works on synthetic biology approaches for therapeutics at the Massachusetts Institute of Technology. The pandemic has shifted the focus away from research toward other challenges in academia, for example, mental health issues on campus and the lack of diversity. Strengthening mentoring programs is seen as one way to address these challenges. However, as Zhao noted, for mentorship to be impactful, the power dynamic between a mentor and mentee must be considered. Ph.D. students have little power in this relationship, particularly given that there are few formal mechanisms for evaluating mentors in academia. As a result, mentees are more likely to be treated poorly, which has broader implications for mental health and diversity on campus. These limitations have been exacerbated by pandemic. To address this, Zhao discussed the need for students to request recommendation letters that can be saved to a cloud server and accessed at any time. Zhao also discussed the need for an evaluation system for mentors. Effectiveness as a mentor should be a particularly important factor in the initial hiring of faculty.
Discussion
During the Q&A session, panelists discussed challenges related to mentoring, obtaining research funding as a postdoc, and the long-term career implications of the pandemic for early career scientists. Anreiter stated that as mentors play a significant role in the careers of postdocs, including, as Shepherd noted, in obtaining tenure, there should be an evaluation system and consequences for mentors who are not supportive of their mentees.
Zhao noted that while there are fewer faculty positions and less funding available as a result of the pandemic, there has been more attention paid to mental health and the need for increased pay for postdocs, which has started productive conversations around these issues. Also, Zhao added that with COVID-19 there has been a push toward open science, pre-prints, and crowdsourced evaluation of science, all movements in a positive direction. The pandemic has taught us lessons about how we can improve science. There is hope that science will shift in a positive direction based on these lessons.
DISSEMINATION AND SHARING
Amy Brand, MIT Press, moderated a session related to the rapid dissemination and sharing of science, including fast response publishing.
Jessica Polka, ASAPbio, discussed the role of pre-prints, noting that they have played a major role in communicating life sciences research during the pandemic. Her organization, ASAPbio, is working to catalyze culture change around the use of pre-prints and open peer review. Currently, pre-prints are reaching monthly posting rates of about 8 percent of the volume of PubMed and experiencing significant growth (see Figure 3 which shows the growth of preprints between 2013-2020). Through this approach, colleagues in the field can begin reviewing and providing informal feedback from a broad range of perspectives and can incorporate these findings into their own work. The urgent need to communicate during COVID-19 is facilitating this process. Pre-prints save time and accelerate scientific discovery. Polka added that these innovations may extend to other areas of publishing. For example, as people are moving meetings online and presenting to broader audiences, there may be opportunities for broader publication of materials such as immediate dissemination of a conference poster. There is a need, however, for an infrastructure to support this move toward pre-prints. The use of a “version note” function which documents changes over time in a manuscript has helped to document changes over the life of a publication. Polka added that there has also been a shift from funders, such as NIH, to explicitly allow pre-prints to be cited in place of journal articles. Pre-prints have been demonstrated to be a highly visible way of disseminating work to the community.
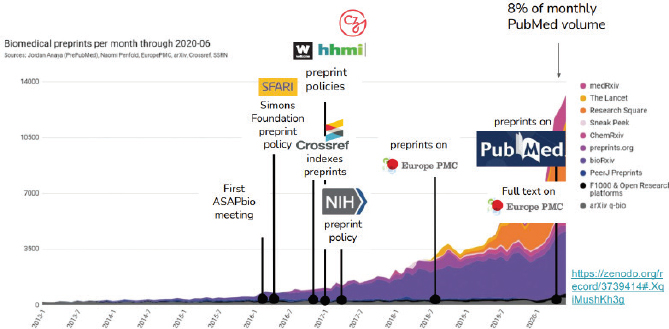
SOURCE: Jessica Polka, presentation, October 15, 2020.
Vilas Dhar, Patrick J. McGovern Foundation, stated that that there is a need to build mechanisms to create public awareness and access to science, including supporting innovations in dissemination and publishing. For example, Dhar added that there is a need to reconceptualize the concept of digital first, where a publication is seen as the start of a public conversation that is continually informed by new scientific advances. AI can support this process through the use of data algorithms to review large data sets at the time of publication. The field of academic publishing should move toward a process of crowdsourcing of papers, allowing researchers to continue to develop and transform their work as new data is collected, said Dhar.
Dhar noted that there is also a need to acknowledge transaction costs associated with moving large amounts of data, including the time needed to standardize data sets. Fundamental questions about ownership of data and the need to acknowledge and credit creators should be considered. Finally, Dhar discussed the importance of funders; philanthropic institutions and science funders stepping forward to create new mechanisms for dissemination and distribution of data and research. There is a need to apply standards and provide the technological capacity to ensure that data sets are broadly accessible to researchers and the public.
Stefano Bertozzi, University of California, Berkeley, discussed issues related to rapid review and publishing activities, including pre-prints. He also discussed the use of AI to support rapid publishing efforts. AI can support the ability to identify published papers that suggest close domain expertise, including authors of those papers. Bertozzi mentioned that the journal he edits, RR:C19, works with teams of Ph.D. students who identify new preprint manuscripts that may be good candidates for rapid peer review. These peer reviews are then published by RR:C19. The reviews are useful to authors to help them address issues prior to submitting the manuscript for publication; to journals seeking quality manuscripts (and potentially abbreviating their internal peer review process); to scientists who seek to build on the work; and to journalists, policy makers and the public that don’t know whether the reported results can be relied upon. This approach aims to minimize the number of successive journal submissions and successive rounds of peer review that occur (usually in secret) prior to publication, thus also improving both the speed and efficiency of publication.
Discussion
During the Q&A session, panelists discussed how the move to pre-prints and open science has changed the peer review process. Noting that peer review is a time consuming and taxing process, panelists discussed the importance of attribution and acknowledgement for reviewers. Polka added that an important component of recognition of reviewers is to publish the peer review, a step that is already being implemented with rapid reviews and pre-prints; however, she added some reviewers are not comfortable with open review, especially revealing their identities. Polka also discussed
the idea of compensating peer reviewers, noting that while many are in favor of this, a considerable number believe this may be problematic and may create negative incentives.
Panelists discussed how societies are supporting the transition to open access. Polka noted that there are examples of open science activities supported by societies, for example, ChemRxiv. Also, she added, the Howard Hughes Medical Institute/Wellcome Learned Society Curation Awards were designed to incentivize societies to create opportunities to review and curate pre-prints.
WHAT DOES IT MEAN?
Hendler moderated a panel summarizing new approaches to rapidly advancing progress in science, engineering, and medicine.
Julia Lane, New York University, addressed data in motion from the perspective of economics and social sciences, noting the importance of government data collection in these fields. There is currently potential to improve the way in which we collect and make use of government data. This includes improving and expanding access to the Federal Statistical System, a federal data collection system that collects, organizes, and structures data, making it available to researchers and the public. Lane added that she has been trying to develop mechanisms to promote access to these data in a secure environment. There is tremendous potential in terms of the use of the data if these barriers, including those related to the confidentiality of data related to individuals, can be removed. Other key challenges include encouraging researchers to use these data as well as more clearly identifying what data are available. Several federal agencies including NIH, the National Science Foundation (NSF), and the U.S. Department of Agriculture, are working to examine their data and determine how and if these are being used by researchers. Lane added that there is currently work under way to develop a machine learning model to enable the identification and reuse of federal data sets.
Hiroaki Kitano, Sony Computer Science Laboratories, Inc., began by discussing his current efforts related to AI, simulation, and modeling around COVID-19 for the Japanese government.3 The objective of this work is to build models and simulations to reduce high risk COVID-19 contact and infection, increase early detection and treatment, reduce the severity of symptoms for those infected, treat patients with severe COVID-19 status, and conduct follow up of recovered patients. This works feeds into an integrated data platform and real-time monitoring system. The system incorporates international standards for testing methods and precision and supports efforts to prepare for future pandemics. He is also working on largescale text mining systems to review data and capture recent research related to COVID-19. Through this work, he has been able to identify a global network of researchers and develop a database of related publications. Kitano added that AI can strengthen research discovery, transform science, support the movement toward alternative forms of scientific discovery, and accelerate science at an unprecedented speed.
William Regli, University of Maryland, described how data can impact science using his prior experience at the Defense Advanced Research Projects Agency (DARPA) and NSF. A key challenge identified through his work with the research community is the lack of awareness of the value of data, including inconsistencies in preserving and curating data. Regli noted that to address this, data should be curated at the time of creation. Regli noted that DARPA has advanced data management plans to help ensure that data are clearly tagged and saved with future users in mind. He also described efforts to pre-register experiments to ensure reproducibility, helping to inform the process of saving data for future use. Regli noted that a significant cost is often the loss of the data at the time of creation and there is a need to shift the economic burden to the future data user. Regli added that as AI advances, so too will the process of data curation and archiving.
Discussion
During the Q&A session, panelists discussed the divide between research produced through traditional approaches and research characterized by rapid data production. These distinct approaches need to meet in the middle to create an engine to accelerate scientific discovery, stated Regli. Hendler discussed the role of AI in terms organizing and analyzing the breadth of scientific knowledge that is anticipated in the future. While AI will be able to model and cluster data, there will still be a need to involve human concepts in organizing data, he said. Kitano agreed, adding that while AI will allow us to explore new and emerging areas, the match between the machine and the human will be important moving forward. Regli stated that we cannot make progress in these areas without the confluence of data, AI, and human beings.
___________
SUMMARY REMARKS AND CONCLUDING DISCUSSION
Hendler summarized the workshop discussion, noting that we continue to teach our children and students an outdated scientific model based on an individual research model. We are moving into a world where models will be shared and compared, changing the research landscape. As such, the concept of the experiment has a different meaning in a world of change and complex data. The notion of distinct disciplines is also changing, driven in part by the complexity and interconnectedness of the problems that science must address, such as the COVID-19 pandemic.
Hendler identified several themes he heard during the workshop, including:
- Modern science is driven by an increasing scale of data and growing demand for rapid research, so more computing power is needed to analyze these data (e.g., simulations, machine learning, etc.).
- Research activity is increasingly undertaken by large teams of researchers as opposed to individual labs.
- The model of the journal as the primary mechanism to share scientific knowledge needs to change to match the speed and complexity of science.
- New models of the dissemination of science are evolving, but not yet fully formed.
- Given the demands placed on science, communication should flow not just between scientists but also between scientists and society.
- Machine learning from data is changing science. New models are being driven by machine learning and other “bottom up” correlative reasoning from data.
- A major challenge is linking traditional “top down” causal science with correlative reasoning.
- Importantly, data itself is and must be treated as a “first class citizen.”
Additionally, Hendler said, to address challenges faced by early career researchers, there is a need to highlight the importance of the advisor-advisee or mentor-mentee relationship. Discipline and country-specific differences are also critical to consider in this relationship. Increasing the independence of junior researchers is a pressing need, and this should cause us to re-think post-doc funding approaches. New ways are needed to evaluate students and faculty that make proper allowances for personal or public events that interfere with progress, handle different stages of publication, and allow credit for mentoring. Mentoring is itself a skill and will require evaluation. Hendler concluded by observing that career hopes and concerns are changing with the times. Senior scientists need to be aware that the world they grew up in is not the world their students and postdocs are now living in.
DISCLAIMER: This Proceedings of a Workshop—in Brief was prepared by Thomas Arrison, Jennifer Saunders, and Emi Kameyama as a factual summary of what occurred at the meeting. The statements made are those of the rapporteurs or individual meeting participants and do not necessarily represent the views of all meeting participants; the planning committee; or the National Academies of Sciences, Engineering, and Medicine.
REVIEWERS: To ensure that it meets institutional standards for quality and objectivity, this Proceedings of a Workshop—in Brief was reviewed in draft form by Anita de Waard, Elsevier, and Ian Foster, Argonne National Laboratory. The review comments and draft manuscript remain confidential to protect the integrity of the process.
PLANNING COMMITTEE: James A. Hendler (Chair), Rensselaer Polytechnic Institute; Amy Brand, MIT Press; Stuart Feldman, Schmidt Futures; and Salman Habib, Argonne National Laboratory.
STAFF: Thomas Arrison, director, Board on Research Data and Information (BRDI); George Strawn, scholar, BRDI; Ester Sztein, deputy director, BRDI; and Emi Kameyama, program officer, BRDI.
SPONSORS: This workshop was supported by the Arnold Ventures and National Institutes of Health as part of BRDI activities.
Suggested citation: National Academies of Sciences, Engineering, and Medicine. 2021. Data in Motion: New Approaches to Advancing Scientific, Engineering, and Medical Progress: Proceedings of a Workshop—in Brief. Washington, DC: The National Academies Press. https://doi.org/10.17226/26203.
Policy and Global Affairs

Copyright 2021 by the National Academy of Sciences. All rights reserved.









