4
Access to Data and Intellectual Property: Scientific Exchange in Genome Research
Stephen Hilgartner, Cornell University
Much of the discussion regarding intellectual property policy is framed in terms of a fairly simplistic image of a world of academic science that is very open and a commercial world that is closed and secretive. My goal here is to present a somewhat more nuanced picture of scientific exchange and to explore its implications for the subject of this workshop. I will begin by describing and critiquing a traditional sociological approach, rooted in the work of Robert K. Merton, to analyzing scientific exchange. I will then present an alternative framework for understanding how scientists regulate access to data, illustrating its use with examples from studies of the research community involved in genome mapping and sequencing. To conclude I will suggest some implications of this framework for analyzing how intellectual property policy might affect scientific openness.1
SOCIOLOGICAL DIMENSIONS OF SCIENTIFIC EXCHANGE
Developing a sociological explanation of the systems of exchange in which scientists participate is a complex task. One way to state the problem is to imagine that a naive observer (see Latour and Woolgar 1979), innocent of any understanding of the practices and behavior of biomolecular scientists, walks into
a molecular genetics laboratory. The observer is confronted by a dazzling variety of biological materials, texts, software, and instruments. A sequencing machine sits on a bench. Racks of clones fill the freezers. Laboratory notebooks, computer printouts, reprints, and draft manuscripts cover the desks. When our observer asks the scientists where all this stuff came from, he is told different things: some of it was purchased in open markets, some of it was produced locally in the laboratory, and some of it came from colleagues. Regarding the eventual disposition of these items, he finds further variety. Most of them are of little interest to anyone outside the laboratory but a few are of intense interest to people in the world outside. Some of the items will be submitted to scientific journals. Some might be included in patent applications. Some might be shared with colleagues. Some might be kept quietly in the laboratory, no one being told about their existence.
How can one develop a sociological explanation of the traffic patterns of resources in and out of scientific laboratories? How can one explain the process that shapes who gets what, when, and under what kinds of terms and conditions? This formulation of the problem of scientific exchange focuses attention on the particular entities, or resources, that are involved in exchanges—entities that received little notice in early sociological work on scientific exchange. The traditional approach to these problems is rooted in the work of Robert K. Merton (1942), who, in a classic paper published in 1942, laid out a theory of the normative structure of the scientific community. Merton argued that a normative commitment to producing knowledge that becomes the common property of the scientific community is one of the defining characteristics of science. Free and open scientific exchange is important because it allows knowledge claims to be extensively tested by a skeptical scientific community. Only the claims that survive a period of intense scrutiny become scientifically certified, valid knowledge—a form of knowledge that is inherently public and communally held.
Building on Mertonian theory, Warren Hagstrom (1965) developed a gift-exchange model of scientific exchange in which individual scientists contribute their findings to the scientific community and in return can expect to receive various forms of recognition. The gift-exchange perspective has been recently applied to molecular genetics by Katherine W. McCain (1991), and a discussion of her paper provides a useful starting point for our analysis. McCain's argument is based on two distinctions. The first distinction is between research results and research-related information. Research results are what gets published in journals or technical reports and thereby become the communal knowledge of the scientific community. Research-related information is a residual category that includes all kinds of entities that embody information but cannot be published in journals; it includes clones, algorithms, software, and descriptions of techniques that are too detailed to be included in the methods sections of scientific papers.
The second distinction is between public science and private science. During the early stages of research, the products of research are the private property
of the individual scientist; people might share research products with their immediate colleagues but, in general, they keep their results in the laboratory. Private science, in this model, occupies a temporal phase in the research process: research products remain within the realm of private science until publication occurs. When a researcher decides to publish findings or to present them at meetings, research results cross the private-public boundary. Similarly, at that point, whatever research-related information is needed to replicate the results also becomes public science. In the public science phase, norms of openness govern the exchange of research-related information so that scientists can validate results and advance the field. Problems of lack of openness in this model occur mainly when people fail to provide research-related information after publication of research results. McCain found little indication that that was a widespread practice.
For defenders of scientific openness, McCain's conclusion might be welcome. But rather than confidently deciding to adjourn our workshop early, we might consider some of the limitations of the gift-exchange perspective. Most important, the perspective gives little consideration to the processes through which scientists decide what entities to move across the private-public boundary and when. To understand the traffic patterns discussed above, it is necessary to look more closely at how, why, and when the entities produced in scientific laboratories cross the private-public divide. In addition, the gift-exchange perspective is based on a rather sharp distinction between public and private science, which is at variance with the many gradations of ''publicness'' that actually occur in scientific practice.
DATA-STREAM PERSPECTIVE
To address those kinds of problems, Sherry Brandt-Rauf and I developed an alternative perspective—the data-stream perspective (Hilgartner and Brandt-Rauf 1994). This framework is informed by a variety of recent social studies of science that emphasize scientific practice and culture, and it is especially amenable to actor-network theory (Callon 1995, Latour 1987). The data-stream perspective conceptualizes data not as isolated objects, but as entities that are embedded in evolving streams of scientific production.
It is important at this point to say something about how Brandt-Rauf and I use the term data. We define it inclusively as the many different entities that scientists produce and use during the process of research. In this usage, data include a wide variety of materials, instruments, techniques, and written inscriptions. Such a broad definition is needed because scientists in every subfield have their own specialized conceptual categories for classifying the resources that they use and produce. Distinctions among data, findings, and results, between samples and materials, and among techniques, software, and instrumentation can become confusing when they cross subfields because the terms are not used uniformly.
Brandt-Rauf and I therefore group all the entities described above under the term data, and we stress the heterogeneity of the category.
Another important point about data is that they are not useful as isolated entities; only when they are connected to a suite of other resources can a scientist use them to accomplish anything. What one needs to perform scientific work is a complex assemblage of resources and techniques. Data are embedded in complex assemblages that weave together many heterogeneous entities. The assemblages are transformed and manipulated as work proceeds, producing evolving streams of products. For example, streams of inscriptions will evolve as the "raw" output of instruments is manipulated mathematically and incorporated into tables, diagrams, and graphs that in turn are explicated and discussed in written texts (see Latour and Woolgar 1979). Streams of materials also result as samples are purified, analyzed, and otherwise processed. The power of scientific research is in the ability of these assemblages to evolve as people produce new entities by reconnecting and recombining many forms of data in many ways.
One of the most important processes that occur as scientific work proceeds is continuing evaluation of the credibility of various pieces of data. At the research front, the assemblages that constitute data-streams contain elements that vary greatly in perceived credibility. Some entities are considered to be well established; people have great confidence in the accuracy of a particular instrument, the reliability of an observation, the robustness of an assay, or the stability of a cell line. Others are of questionable validity. Perceived credibility often fluctuates throughout the process of scientific production (for example, Collins 1985; Collins and Pinch 1993; Knorr-Cetina 1981; Latour 1987; Latour and Woolgar 1979; Lynch 1985). In general, judgments progress toward a definitive resolution. Some data are rejected and others deemed reliable, but the temporal nature of the evaluation process—with the shifting perceptions that often accompany it—adds complexity to data-streams.
The data-stream perspective emphasizes the continuous properties of scientific production: data are conceived as phenomena that are embedded in evolving assemblages, rather than as discrete entities with unshifting boundaries. The point is to shift the level of analysis to the stream as a whole. In trying to explain the kinds of exchanges in which scientists engage, one cannot assume that data arrive on the scene in neatly packaged units that are naturally ready to be disseminated. Each scientific field has its own conventions about what constitutes a publishable paper and what constitutes an interesting result. Within particular communities of researchers, these conventions can be clearly understood and indeed seem obvious to participants. However, conventions are neither identical across fields nor entirely stable. To provide general insights into how data are shared or otherwise exchanged among scientists, one cannot simply assume that the conventions of a single research community constitute the only way to conduct science; instead, the conventions of different research communities become phenomena for social analysts to explain. The data-stream perspective frames
the issue in terms of two central questions: What portions of a given data-stream typically are distributed to whom and under what terms and conditions? How are these portions bounded? In other words, how are discrete entities extracted from the continuously evolving streams of scientific production and entered into exchange relations?
TYPES OF TRANSACTIONS: STRATEGIC CONSIDERATIONS
To understand systems of exchange in science, one must consider the different sorts of transactions in which scientists engage. The limits of the gift exchange perspective will also emerge. McCain (1991), for example, emphasizes two kinds of transactions: transactions between an individual laboratory and the entire scientific community (such as publication of a research paper) and transactions between a source and a requestor (as when one laboratory requests a clone from another). However, these are included in a much broader array of transactions that take place in academic science. It is hard to overstate the importance of open publication in academic reward structures, but it must also be remembered that even in academe publication is only one means among many of distributing data. Data are also quietly given to selected colleagues, they are patented, they are transferred when visitors come to the laboratory to learn novel techniques, they are bought and sold, they are privately released to corporate sponsors, and they are retained in the laboratory pending future decisions about their fate (Hilgartner and Brandt-Raub 1994).
It is useful to consider the strategic issues involved in selecting which of those things to do at what time. A key concept here is competitive edge. Using the idealized example of some novel techniques, we can see that it is typical in molecular biology for a new technique initially to be difficult—it might need "magic hands." Later, however, it is increasingly routinized and is packaged in standard protocols; if it is extremely successful, it is incorporated into commercial kits and sold on the market (see Fujimura 1987). In any case, the technique typically becomes available to an increasingly wide circle. As that is going on, the competitive edge that a scientist gets from using the technique in the laboratory declines. At the beginning, because the technique is scarce, a researcher might be able to do things that no one else can do, gaining a short-term competitive advantage that can be important. The competitive edge typically declines as the technique or other initially scarce entity is disseminated.
Various strategies can be used to exploit short-term competitive edges strategically. One of them—widely used by academic scientists—is to restrict access and use data to generate more data that will be cashed in later. Another is to use carefully targeted access; for example, the data can be used as a bargaining chip to negotiate with corporate sponsors or others for resources in return for access. Another is to offer widespread access, say, via open publication, to build one's academic reputation. Often, the question is not necessarily whether to provide
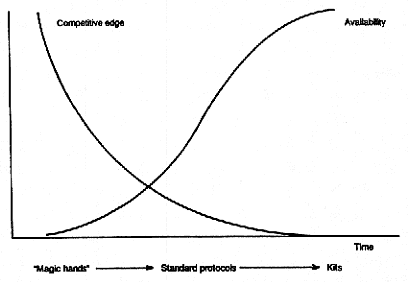
Relationship between the availability of a technique and the competitive edge it conveys. Following initial development, the technique is progressively routinized and becomes available to growing numbers of labs. As this occurs, the comparative advantage it conveys declines. Source: Published with permission from Sage Publications.
access but how much access to provide and when. Here, decision strategies become very complex. Timing can be a key element, both because competitive edge might exist for only a short period of time and because access control interacts in interesting ways with quality control (for discussion in the context of Internet-accessible biomolecular databases, see Hilgartner 1995). Scientists constantly make judgments about whether data are sufficiently reliable to disseminate.
COLLABORATION
Another set of important interactions around the data concerns collaboration (Hilgartner and Brandt-Rauf in press). Scientific collaboration takes many forms, but one important feature of many of them is the merging of portions of data-streams. From this point of view, collaboration involves negotiations in which data serve as bargaining chips in discussions of whether it makes sense to pool resources. Assessing whether or not a collaboration makes sense entails considering the resources that different groups can bring. In addition, because forming a collaboration allows scientists to create a data-stream that spans the boundaries of individual laboratories, negotiations about the terms of the collaboration must
typically be conducted. Someone who controls a unique resource is in a position to dictate the terms of all collaborations involving that resource. As one molecular biologist put it to me, "if the other people don't like those terms, they don't have to collaborate." In other situations, complex and protracted negotiations can arise, especially when groups possess relatively equal resources and there is no clear hierarchy governing the relationship (Hilgartner and Brandt-Rauf in press). In some cases, such as the European yeast genome-sequencing program, policy-makers establish formal rules granting entitlements to portions of a data-stream to manage collaboration among geographically dispersed laboratories (Hilgartner and Brandt-Rauf in press).
Even once negotiations are concluded, the resulting collaborations can be fragile and require continuing attention. There seem to be several reasons for that. Many collaborations break down just because the work fails to extend data-streams in the expected directions. Another problem stems from concern that the exchanges among different laboratories have become uneven. A third is ambiguity about what portions of a data-stream are included in a collaboration; this is a problem that is often unavoidable because it might be infeasible to negotiate these matters too tightly in advance. Most of the time, the limits and nature of a collaboration are based on a shared understanding, but shared understandings can easily break down. Social psychologists have shown that in a variety of situations involving collective work (such as basketball games, marriages, and joint writing projects), people tend to perceive their own contribution as being larger than the other participants perceive it, perhaps because it is easier to recall the details of one's own activity than to recollect what other people did (see, for instance, Ross and Sicoly 1982). For all reasons, the negotiations that create and sustain collaborations are not a stage in the research process, but rather a continuing process.
THE DATA-STREAM PERSPECTIVE APPLIED: A SCHEMATIC EXAMPLE
A schematic example might help to illustrate how the data-stream perspective can be used to explain access practices in a particular field of science (from Hilgartner and Brandt-Rauf in press for fuller discussion). Consider the case of hunting for disease genes in the late 1980s or early 1990s, when creating physical maps in the region of a disease gene required a massive amount of work. From high altitude, the data-stream for mapping disease genes can be understood as an evolving assemblage of landmarks in the region where the gene is believed to lie, in which the landmarks become more densely interconnected as work proceeds. Access to such a data-stream could be provided in many ways. At one extreme, a laboratory could use the Internet and Federal Express to provide daily access to its entire data-stream. At the opposite extreme, a laboratory could withhold the entire data-stream and even keep secret the fact that it is looking for the gene.
Those observations suggest a couple of questions, which I will attempt to answer below: What are the typical practices through which gene hunters regulate access to their evolving maps? Can those practices be explained in terms of the structures of data-streams?
Turning first to typical practices, there has been considerable variation within the disease-gene mapping community as to the level of access provided. However, there is no doubt that it takes place only within a portion of the range between the theoretical extremes. In particular, a culture of extreme openness is not to be found: no one provides unrestricted daily access to evolving data-streams in this line of research. Instead, three practices for providing access to data-streams are typical. The first is nonrelease, in which scientists hold the data in the laboratory while production proceeds. A second is delayed release, in which a large gap is maintained between the time when the data are generated in the laboratory and the time when it is shared outside the laboratory. The third is isolated release, in which a laboratory provides access to portions of the data-stream that have been specifically bounded in ways that make the data-stream difficult or impossible to extend.
To explain these practices it is useful to take note of the competitive structure of disease-gene mapping, which was complex and laborious in the late 1980's and early 1990's. Success was by no means guaranteed. Moreover, in the case of the major Mendelian disorders, multiple groups competed to find each gene, and people aptly referred to gene hunts as races. In short, gene hunting was a high-stakes game with a well-defined goal that only one group could reach. It was characterized by intense zero-sum competition.
The second issue to consider is the potential value that different portions of the data-stream have to competitors. In the case of genetic maps that are produced with publicly available clones, published data are of immediate utility to competitors because combining multiple sources of data will usually lead to an increase in the quality of the map in the region. People not only can instantly catch up, but if they are keeping their own data secret at the same time, they might be able to seize the lead. Even if competitors wanted to check the data carefully, the economies are favorable because it takes fewer tests to verify a map than were required to produce it in the first place. It is therefore not surprising that gene hunters do not provide daily releases, and nonrelease can be explained as a consequence of zero-sum competition.
How, then, can we account for delayed release? Because one cannot engage in nonrelease forever! Eventually, it becomes necessary to demonstrate progress to colleagues and funders to sustain a long-term gene hunting effort. Another reason is that scientists use publication as a hedge against the possibility that another group will identify a given gene first, which is fairly likely in many of these races. Although a paper titled "Mapping in the Region of the Gene for Deadly Disease" will be minor in comparison with a paper titled "Positional Cloning of the Gene for Deadly Disease," it has some value. The practice of
delayed release thus can be viewed as a strategy for managing a tradeoff between the advantages of publication and the risks of competitors' taking the lead. If the delay is managed carefully, the risks will be minimal.
If we turn to isolated release, the strategic incentives are similar to those for delayed release. However, the practices that enable people to make data-streams impossible to extend are often controversial. A good example of a means of data isolation in the arena of disease-gene mapping is the renaming of clones: by changing the name of a publicly available clone, one can make it unrecognizable, and maps that use the novel name cannot be extended. Renaming clones is considered inappropriate by many researchers, so doing so entails risks to reputation. For some combination of moral and strategic reasons, many disease-gene mappers refrain from such practices, although there are exceptions.
This stripped-down discussion shows how a strategic analysis that looks at the empirical structure of data-streams can go a long way toward explaining scientists' practices regarding access to data. However, this kind of analysis has important limitations. First, at this level of generality, the analysis applies to disease-gene mapping in general and does not take into account the differences in the histories and personalities associated with different genes, chromosomes, and so on. Second, this kind of strategic analysis does not include any discussion of the rhetoric that is used in access negotiations, which can be important in shaping outcomes. Third, the focus on strategic incentives clearly needs to be broadened to include collective definitions of appropriate conduct in science and how those definitions are applied to new situations and renegotiated during their application.
Despite those limitations, the example provides a sense of how scientific exchange can be analyzed from the perspective of data-streams. In addition, it suggests how different this kind of analysis is from the kind that results when one assumes that academic science is governed simply by a culture of openness.
INTELLECTUAL PROPERTY AND OPENNESS
I have argued that rather than merely assuming that academic science is governed by openness, analysts should try to understand the processes that shape what gets made public, what is kept private, and what is deployed in transactions that fall between these extremes. I now want to consider the implications of the more nuanced picture of scientific exchange that I advocate for issues of intellectual property protection. In particular, I want to explore the question of whether we should expect an emphasis on intellectual property in academic science to cause a reduction in scientific openness.
To look at that question, one clearly needs to consider how intellectual property considerations influence a number of aspects of scientists' practices. Do intellectual property considerations influence what portions of data-streams are provided, to whom, and when? Do they introduce new sources of delay? Do they
change the kinds of restrictions that are placed on the use of data? Do they increase the complexity and formality of negotiations over access to data? Do they make collaboration more unstable or difficult to form? Do they complicate the development and maintenance of shared understandings about control over data-streams that are collectively produced? Without a doubt, the answers to those questions will vary with the particular context, and many case studies will be needed before these issues are fully understood. However, I want to conclude with several assertions about the likely effects of intellectual property on openness in academic science.
First, there is little reason to believe that intellectual property protection is likely to lead to an increase in openness among academic scientists. In the world of commerce, patents are perceived as promoting openness because they are seen as an alternative to trade secrets, which clearly constitute a more restrictive legal mechanism. A patent offers downstream readers an opportunity to extend a technology by providing details about how an invention works—details that would be unavailable under a regime of trade secrecy. However, in the world of academic science, the restrictions on openness motivated by possible commercial exploitation might tend to propagate upstream from the point of potential patent back into the research process. Consequently, one would expect access to portions of data-streams that are believed to be precursors of potentially patentable products to be relatively tightly controlled.
Second, there is also considerable empirical evidence that intellectual property considerations actually reduce openness, at least on occasion. Michael Mackenzie, Peter Keating, and Alberto Cambrosio, for example, show how the expansion of what was considered patentable in the realm of hybridoma and monoclonal antibodies was accompanied by reductions in the free flow of scientific information (Mackenzie and others 1990). My own interviews with molecular geneticists suggest that at least minor delays in publication sometimes occur while scientists, university technology-transfer offices, and patent lawyers make assessments of the potential commercial value and patentability of results. Survey research on academic-industrial relations has also suggested that biotechnology faculty with commercial involvements are more likely to have engaged in practices that restrict scientific openness (Blumenthal 1992) and some evidence suggests that concerns about intellectual property protection can complicate negotiations about scientific exchange and in some cases make it more difficult to form and maintain collaboration. Indeed, one way to interpret efforts to develop and put into use standard "material-transfer agreements" is as an attempt to reduce the impact of such complexities on scientific collaboration.
A third point about the likely effects of intellectual property considerations on academic science is that the effects will not be uniform across all scientific fields. That is true not only in the trivial case of comparing lines of research with different commercial potential, but much more generally. The perspective outlined above suggests that the effects of intellectual property concerns will be
mediated by the prevailing structure of data-streams in particular lines of research. Access practices are probably most intensively shaped not at the level of the discipline or field, but at the level of much-narrower links of research, such as disease-gene mapping, that can be defined in terms of a characteristic data-stream and a particular competitive structure.
If that is true, then one might ask in which lines of research one would expect to find intellectual property considerations producing the largest reductions in openness. The data-stream perspective suggests that the answer might depend in large part on the specific competitive structure of a field of research. In a field characterized by races with intense zero-sum competition, commercial concerns will probably not have a pronounced effect; even in the absence of the potential for profit, the reasons for restricting access are already strong. For example, even if disease genes could not be patented, the winner of the race to find an important gene in the late 1980s could expect substantial rewards. At that time, few human disease genes had been cloned, and cloning one constituted a major achievement.
However, it is important to recognize that intense zero-sum competition is not the typical situation in academic research. In many fields, scientific goals might not be sufficiently well defined, agreed on, and focused on identifiable targets to inspire races among rivals with the same finish line in mind. Instead, research might be exploratory; and in some cases, only one laboratory might be pursuing a given line of investigation. In the absence of focused competition among research groups, openness might be relatively unrestricted. Consequently, one might expect the greatest reductions in academic openness to be provoked by introducing the prospect of commercialization into less-competitive situations.
REFERENCES
Blumenthal D. 1992. Academic-industry relationships in the life sciences: extent, consequences, and management. J Amer Med Assoc 268(23): 3344–3349.
Callon M. 1995. Four models for the dynamics of science. In: Jasanoff S, et al., editors. Handbook of science & technology studies. Newbury Park, CA: Sage Publications. Pp 29–63.
Collins HM. 1985. Changing Order: Replication and Induction in Scientific Practice. University of Chicago Press.
Collins HM, and Pinch, T. 1993. The Golem. Cambridge Univ Press.
Fujimura JH. 1987. Constructing 'do-able' problems in cancer research. Social Studies of Science 17: 257–293.
Hagstrom WO. 1965. The scientific community. New York: Basic Books.
Hilgartner S. (in press). Data access policy in genome research. In: Thackray A, editor. Private science. University of Pennsylvania Press.
Hilgartner S and Brandt-Rauf SI. 1994. Data access, ownership, and control: toward empirical studies of access practices. Knowledge: Creation, Diffusion, Utilization 15(4): 355–72.
Hilgartner S. 1995. Biomolecular databases: new communication regimes for biology?" Sci Comm 17(2): 240–63.
Hilgartner S, Brandt-Rauf SI. in press. Controlling data and resources: access strategies in molecular genetics In: David P and Steinmueller E, editors. A productive tension: university-industry research collaborations. Stanford University Press.
Knorr-Cetina KD. 1981. The manufacture of knowledge. New York: Pergamon.
Latour B. 1987. Science in action. Cambridge, MA: Harvard University Press.
Latour B and Woolgar, S. 1979. Laboratory life. Beverly Hills, CA: Sage Publications.
Lynch M. 1985. Art and artifact in laboratory science. London: Routledge and Kegan Paul.
Mackenzie M, Keating P. Cambrosio A. 1990. Patents and free scientific information: making monoclonal antibodies proprietary. Sci Tech Human Values 15(1): 6–83.
McCain KW. 1991. Communication, competition, and secrecy: the production and dissemination of research-related information in genetics. Sci Tech Human Values 16(4): 491–516.
Merton RK 1973 [1942]. Science and technology in a democratic order, reprinted as The normative structure of science. In Merton RK, editor. The sociology of science. Chicago: University of Chicago Press.
Ross M, Sicoly F. 1982. Egocentric biases in availability and attribution. In: Kahneman D, et al., editors. Judgments under uncertainty: heuristics and biases. Cambridge Univ Press.












