Restoring the Environment Via Bioremediation and Molecular Sciences: Prospects for Better Understanding and New Science-Based Solutions
Michael L. Knotek
Advanced Photon Source Argonne National Laboratory Argonne, Illinois
and
Shirley A. Rawson
William R. Wiley Environmental Molecular Sciences Laboratory Pacific
Northwest National Laboratory Richland, Washington
There is a tremendous national and international legacy of environmental contamination—from defense production, general industrial activity, and agriculture—that poses an enduring threat to the health of humanity and to the survival of ecosystems throughout the biosphere. The bulk of this contamination occurred in a relatively short time between the Second World War and the time in the early 1970s when the long-term adverse effects of such contamination were first realized. Stringent regulations have halted further release of such materials, but the problem of restoring the environment and protecting the health of future generations largely remains, and is proving to be a daunting challenge. While the problem of extracting or passivating contaminants in the environment has proven to be vastly more complex than first imagined, an intractable system of regulations and practices has made it even more difficult to bring the benefits of sound scientific research to actual cleanup efforts. The conservatism built into the system has led to environmental restoration's focus on proven, albeit dated, science and simple technologies. Associated bureaucracies related to the legal requirements have been proven to be ineffective and costly.
However, this is a time of unprecedented progress in scientific understanding of phenomena in the environment, and evolving capabilities in empirical and theoretical research promise a new era in scientific advances, from the molecular to the system level. In addition, our understanding of the risks posed by substances in the environment is evolving rapidly. It is critical at this time to capitalize on these opportunities.
Progress has occurred on numerous fronts. There has been tremendous progress in our understanding of ecosystems in general, and in the ecosystems of subsurface environments especially. The discovery of the existence of extensive deep subsurface microbiology, its complexity and its functionality, is revolutionary. There is an unexpected diversity in microbial communities: Microorganisms and organisms in other phyla exhibiting the ability to express selected genes under stress, and genetic sharing of traits between members of a microbial community (consortium) and between species. Many of the traits exhibited by subsurface communities are common to the traits exhibited by drug resistant microbial communities now bedeviling the health profession. A picture of a robust and complex response by microbial communities (consortia) is emerging, making bioremediation feasible. Whereas genetic engineering was only recently deemed necessary to enable microbes to deal with
pollutants that arise from human activities, the genetic diversity and complex functionality of native communities now suggest that natural attenuation may be the basis for long term solutions, in many cases obviating intervention. Monitoring and modeling replacing active intervention.
Modem tools from the physical sciences, computing sciences, and life sciences promise explosive growth in knowledge in both the predictive and the empirical development of these problems. Computational sciences have shown an exponential growth with the advent of massively parallel systems and are poised to carry us past the reductionist paradigm for science. Computational science will soon be capable of dealing with the systems-level complexities of environmental contingencies. Likewise, the emerging generation of experimental tools that use advanced physical probes—such as synchrotrons, nuclear magnetic resonance, laser-based tools, and surface science—brings revolutionary sensitivity and analytical richness to practical systems studies. And genomics, structural biology, and molecular-level mechanistics are becoming the tools of choice for the study of all advanced biological systems.
An improved mechanistic understanding of health effects and the robustness of genetic defense and repair processes are changing our idea of the ultimate impact of environmental exposures to broad classes of compounds and elements. New classes of persistent manmade chemicals in the environment, e.g. endocrine disruptors, are joining genotoxins and other carcinogens as critical targets for research and eventually for control and mitigation.
Possibly most important, the information revolution promises to cut across historical institutional, and disciplinary boundaries to bring concepts and capabilities together in new ways, focused on problem solving, which will transform the research paradigm. Tools are emerging to bring teams of people and experimental and computational capabilities together in a real time, vertically integrated environment. It is an exciting time for conducting research that can contribute to science-based solutions to the Department of Energy's (DOE) environmental restoration challenges, and the DOE Biological and Environmental Research (BER) program is well-positioned to make significant contributions to those solutions.
THE NATIONAL WASTE LEGACY
To appreciate the areas in which further scientific advances are needed, the magnitude of contamination must be considered. Across the country there are more than 20,000 declared hazardous material generators. Hazardous waste is routinely sent to more than 5,000 waste treatment, storage, and disposal facilities. Inventories have listed more than 600,000 leaking underground-storage tanks in the nation. As previous hazardous waste sites are characterized, there could be as many as 32,000 national CERCLA sites.1
Furthermore, the nation has responsibility for approximately 6,000 contaminated federal facilities.2 For example, DOE is responsible for waste management and environmental restoration of more than 130 installations across the United States. Contaminated media at DOE's facilities include about 1.85 × 109 m3 of water (99% groundwater), and 79 × 106 m3 of solid media (95% soils and sediments).2 About 56% of the waters show radiologic contamination only, 30% mixed hazardous and radiologic contamination, and 14% hazardous contamination. Radiologic contamination of soils exceeds 70% with equal proportions of mixed and strictly hazardous contamination.2 Many of the contaminants are widely distributed in groundwater and in soil and sediments at low concentrations across many diverse geologic settings.
A broad variety of contaminant classes were disposed at DOE's facilities. 3 The most-common classes were fuel hydrocarbons, chlorinated hydrocarbons, radionuclides, metals, and ketones; the most-common substances were toluene, trichloroethylene, tritium-uranium, lead-chromium, and acetone. More than 50% of DOE's waste sites were contaminated by binary or ternary mixtures, with the most-common binary-contaminant mixture consisting of metals and radionuclides.3 Twelve other common pairings included metals, anions, radionuclides, chlorinated hydrocarbons, polychlorinated biphenyls (PCBs), and ketones in various combinations.
Many of the combinations resulted in mixtures of contaminants that could interact with each other to modify contaminant subsurface geochemical behavior. Mixtures of radionuclides and metals with organic ligands (organic acids or amino-carboxylic chelating agents) can form mobile aqueous complexes in soil and groundwater. Likewise, organic solvents (chlorinated hydrocarbons and ketones), which can mobilize sparingly soluble hydrophobic organic compounds, were disposed of with PCBs. Organic compounds that can stimulate subsurface microbial
activity, thereby modifying speciation of metals and radionuclides, were disposed of simultaneously with metal-and radionuclide-containing wastes.
The volume and extent of contamination of diverse subsurface environments creates a pressing need for effective remedial technology. Early remedial approaches, such as excavation and groundwater pumping and treatment, have proved inadequate to address the problem.4 Attention is now focused on insitu technologies that include bioremediation, intrinsic contaminant degradation, and contaminant containment and treatment with insitu permeable chemical barriers.5 There are several advantages to these approaches: Because they are insitu technologies, the wastes are contained and treated on site, contaminants can be completely transformed to nonhazardous materials, and the public perception is one of "natural processes." These technologies also have disadvantages: Because they are insitu technologies, the methods must be adapted to the specific chemical, physical, and biologic conditions of each site; monitoring and assessment of successful contaminant transformation are difficult, and the concentrations of process end products can exceed regulatory requirements. All these approaches require a detailed understanding of the physical, chemical, and biologic processes that occur in subsurface environments.
The DOE BER program has had a long-term commitment to basic research that provides the scientific basis for environmental-restoration technologies. The success of any of these approaches depends on coupling process-level understanding with long-term monitoring of contaminated systems that are undergoing treatment to pre-established end points. In the case of insitu bioremediation, successful remediation requires knowledge of the diversity and function of indigenous microbial communities, of the expression of degradative genes that might be present in the population, and the chemical reactivity of contaminants. To assess the success of intrinsic contaminant degradation, additional information is needed on the nature of solute and microbial transport in subsurface environments, which in turn requires knowledge of subsurface physical and chemical heterogeneity. Traditional sampling, measurement, and monitoring strategies for subsurface environments are expensive and time-consuming, and they provide a complete view of the range of microbial ecology in the subsurface. When insitu permeable chemical barriers are considered for use at contaminated sites, similar knowledge on the subsurface environments is needed. The gaps in understanding of how complex hydrologic, geochemical, and biologic processes interact to affect subsurface contamination have formed the basis for much of the BER program's research aimed at restoration of DOE lands.
BER PROGRAM ACCOMPLISHMENTS
The BER program's studies in subsurface science since 1985 have contributed substantially to the fundamental understanding of microbial ecosystems in pristine aquifers and unsaturated zones. Over the last decade, BER has sponsored research in deep microbiology and bacterial transport at multiple DOE and private field sites and was responsible for developing techniques to collect samples that could be unequivocally demonstrated to host microorganisms representative of indigenous microbial ecosystems. 6-9 Diverse bacterial communities were detected in a range of geologic environments not previously known for hosting microbial communities, including coastal-plain sediments in eastern Georgia, consolidated rock almost 3 km deep in sedimentary basins in Virginia, and basaltic aquifers in central Washington.6,10 Researchers have used 16s ribosomal RNA (rRNA) probes to determine the phylogeny of many of the ecosystems and to relate microbial strains isolated from the subsurface to known general.11,12 In many cases, the large numbers of the organisms within subsurface samples could not be identified; this showed the uniqueness and potential long-term isolation of these environments. Some of these communities produced novel and unique organisms with previously unsuspected capabilities to degrade anthropogenic compounds.13,14 A new chemoheterotrophic strain of Sphingomonas, Sphingomonas aromaticivorans , was identified as part of the consortium present at the DOE's Savannah River site. The species catabolically degrades a broad array of aromatic compounds, including xylene, toluene, and naphthalene; the ability to degrade these compounds was determined to be encoded on a 180-kb plasmid (figure 1).13,15,16 These findings highlight the ability of the subsurface community to degrade and transform particular contaminants.
Many of the organisms have adapted to deep subsurface environments, as indicated by the uniqueness of their metabolism. The BER program supported the discovery of a bacillus, Bacillus infernus, that is a strict anaerobe (it cannot tolerate oxygen).10 The B. infernus strain was isolated from rock fragments collected at depths of 2.65
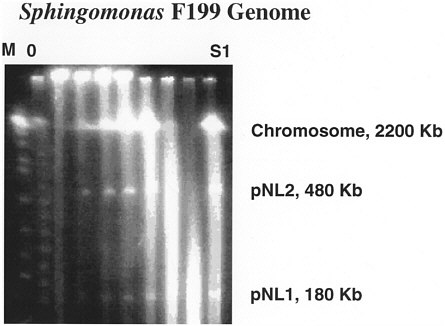
FIGURE 1 Gel blot of F199 genomic DNA showing presence of 2 megaplasmids, including pNL-1 at 180 kb, which appears to encode for degradation of aromatic compounds (Kim and others, 1996).
2.77 km in the Taylorsville Basin of Virginia. The bacterium can grow by fermentation of glucose or oxidation of formate or lactate accompanied by the reduction of Mn(IV) and Fe(III). Elsewhere, program-sponsored research in the biogeochemistry of subsurface systems used laboratory studies to examine the mechanisms of dissimilatory iron reduction, which has been inferred to be responsible for the close spatial association of subsurface microorganisms and secondary iron-beating mineral oxides, such as goethite.17 Experiments demonstrate that such iron oxides can play an important role in the chemical transformation of contaminant complexes.18,19 Some of the concepts of long-lived, subsurface permeable barriers call upon microbially catalyzed iron reduction as an integral part of the remedial treatment of metals and radionuclides.5 The program's research findings indicate that microbial populations with metabolism adapted to perform the kind of subsurface metal oxidation and reduction important to contaminant transformation are already present in the subsurface.
Previous research addressed the issue of how microbial ecosystems were established in subsurface environments. Questions about the availability of sufficient nutrients and energy sources and the rates of nutrient and bacterial transport were addressed in transport studies conducted in intermediate-flow cells and in field settings. In intermediate-scale experiments in the laboratory, researchers have studied the transport and distribution of bacteria in physically and chemically heterogeneous porous media specifically constructed to determine the complex interactions between the heterogeneities.20 Microorganisms have been discovered to position themselves at interfaces of physically heterogeneous media because of the chemical heterogeneities in electron-acceptor concentrations that arise at such interfaces. In field studies, the transport of bacteria through aquifer systems often appears to be limited by the physical constraints of the porous media; results from intermediate-flow cell experiments allow scientists to evaluate the magnitude of the contribution made by solute flow to the residence time and availability of nutrients and electron acceptors.20,21
The role of microbial ecosystems in buffering the aqueous geochemistry of aquifers has been explored by BER program researchers using stable-carbon isotopic geochemistry; as a result, the existence of anaerobic
subsurface lithoautotrophic microbial ecosystems that derive energy from geochemically derived hydrogen has been suggested.22 Such ecosystems might constitute a potential model of how life can occur on other terrestrial-type planets, such as Mars. Just recently, the National Aeronautics and Space Administration's exobiology program has been willing to fund exploratory research in this field to explore how far through the solar system microbial communities extend.
The BER program's studies on the microbial genome were influential in providing the definitive genetic sequence to confirm the Woese rRNA determination of the Archaea as a separate major domain, with Bacteria and Eukarya, on the tree of life.23 The determination of the complete genetic sequence for the extremophilic Methanococcus jannaschii revealed a high proportion of previously unknown genes and confirmed the uniqueness of Archaea relative to bacteria. The discovery allows re-examination of the question of the origin of life and assists in confirming the physicochemical conditions under which life first began.24 Extremophilic organisms like M. jannaschii are being intently studied as possible sources of the enzymes important to new industrial processes, and knowledge gained about their natural enzyme systems can support the re-engineering of enzyme systems for specific functions, such as contaminant degradation.
FUTURE DIRECTIONS IN ENVIRONMENTAL RESEARCH
Much still is not known about subsurface microbial ecosystems, the degradative potential of microbial consortia, the mechanisms by which microorganisms degrade contaminants, and the effects of the microorganisms on insitu subsurface chemical and physical processes. A major need exists for an integrative and predictive capability that will allow coupling of molecular-scale mechanistic understanding with processes occurring on larger physical scales. The colonization of subsurface environments by microbial consortia is not yet fully understood. Transfer of genetic material among subsurface species is not well understood. There is also a need to develop the ability to design enzymes that can withstand extreme environmental conditions. The relationships between subsurface microbial ecosystems and macroscopic surface ecosystems have not been conceptualized from the perspective of complexity theory and self-organizing systems.
To address these kinds of research questions, the BER program has initiated another program in Natural and Accelerated Bioremediation Research (NABIR). The NABIR program builds on the BER program's decade of basic research contributions in subsurface microbial ecology and biogeochemistry and is aimed at providing the fundamental research to explain and manipulate the chemical, biologic, and physical processes that contribute to insitu bioremediation. The degradation or transformation of contaminants by microorganisms is seen as having great potential for meeting DOE's restoration challenges.
Modem physical and chemical tools are now available to address some of the unanswered questions about subsurface microbial ecosystems and demonstrate how more scientific knowledge will link to improved remedial approaches. Research sponsored by the NABIR program will use many of these tools.
Using the highly focused x radiation generated at new, third-generation light sources, such as the Advanced Photon Source at Argonne National Laboratory, and zone-plate collimation, scientists can explore the distribution of trace metals in complex microbial ecosystems associated with subsurface environments, such as plant-root systems.25 The transformation of iron valence state as a function of depth within the crystal lattice can also be studied with synchrotron radiation, as evidenced by recent studies of dissimilatory iron-reducing bacteria at the Advanced Light Source at the Lawrence Berkeley National Laboratory.26
The BER program opened its first national scientific user facility, the William R. Wiley Environmental Molecular Sciences Laboratory at Pacific Northwest National Laboratory in Richland, Washington, in October 1997. The mission of the facility is to develop a molecular-level understanding of the physical, chemical, and biologic processes that underlie environmental remediation, water processing and storage, human-health effects, and atmospheric chemistry. The fundamental environmental molecular science that is conducted within the facility provides the knowledge base needed to address DOE's serious environmental issues, including the extensive contamination of soil and groundwater described earlier.
The complement of research equipment and general laboratory infrastructure within the Wiley Environmental Molecular Sciences Laboratory are grouped into three national user facilities: the High Field Magnetic Resonance
Facility, the High Field Mass Spectrometry Facility, and the Molecular Sciences Computing Facility. These facilities contain several one-of-a-kind and first-of-a-kind instruments that will support scientific advances in a variety of disciplines. For example, the High Field Magnetic Resonance Facility will contain one of the world's first 900-MHz nuclear magnetic resonance (NMR) spectrometers and a state-of-the-art 750-MHz NMR spectrometer. These instruments support studies of the molecular structure of enzymes, proteins, and DNA as related to bioremediation and cellular-response effects. The facility also supports a suite of modem pulsed electron paramagnetic resonance instruments that allow scientists to perform structural analyses of metal clusters important to enzymatic reduction of groundwater contaminants. Near-field optical microscopy permits the fluorescence imaging of single-chromophore probe molecules in specific ambient environments with nanometer spatial resolution. Detailed information on chemical reactions of individual molecules can be obtained through the combination of time-resolved spectroscopy and near-field optics. Environmentally important research includes the examination of membrane systems in plants, which can be extended by analogy to the membrane systems in microorganisms important to bioremediation.27-29 A more-extensive discussion of the use of near-field microscopy for understanding biodegradation of chlorinated hydrocarbons is given by Dunning elsewhere in this volume. The Molecular Sciences Computing Facility contains one of the nation's fastest massively parallel computers, an IBM Scalable POWER Parallel System, which expands the capability to perform ab initio calculations of molecular structure for larger and larger single molecules. For example, in 1993 it was possible to perform an ab initio calculation of the structure of 18-crown-6 ether on a Cray supercomputer over a period of months; the computer in the new Molecular Sciences Computing Facility allows one to perform a similar computation on a Still's crown ether (a much larger molecule) within an hour or two. That example equates to an increase in computing capability by a factor of 100-1,000, which suggests that the nonlinearities and complexities of subsurface flow and transport can also be addressed more easily on such a machine. Dunning provides details of some of the research aimed at environmental restoration being addressed with the computational capability of this machine.
As the BER program initiates research in its NABIR program and brings the power of modem physics and biology to bear on key environmental processes related to bioremediation, the scientific community has the opportunity to contribute fundamental scientific research of direct relevance to addressing a major societal issue, the environmental legacy of the Cold War. The BER program's new national scientific user facility, the Environmental Molecular Sciences Laboratory is expected to provide resources to research teams nationwide for improving the understanding of molecular science related to the environment. It plans eventually to develop a field-research center for bioremediation that will allow questions of implementing bioremediation, including transport of microorganisms, to be addressed. These major opportunities for scientific advancement will be but the first of many sponsored by the BER program as it continues to foster basic scientific research into the next century.
ACKNOWLEDGMENTS
The research described in this paper has been supported by several offices within the DOE Office of Energy Research. The authors gratefully acknowledge the support and encouragement of Ari Patrinos, associate director for biological and environmental research (BER); and Michelle Broido, director of the BER Program's Environmental Sciences Division; and Robert Marianelli, director of the Chemical Sciences Division of the Office of Basic Energy Sciences. The authors also extend thanks and appreciation to the contributions of the many scientific investigators supported by the BER program.
REFERENCES
1. MacIlwain C. Science seeks weapons cleanup role. Nature 1996;383:375-9.
2. US Department of Energy. Linking legacies: connecting the cold war nuclear weapons production processes to their environmental consequences. DOE/EM-0319. 1997.
3. Riley RG, Zachara JM. Chemical contaminants of DOE lands and election of contaminant mixtures for subsurface science research. DOE/ER-0546T. Washington, DC: US Department of Energy, Office of Energy Research. 1992.
4. Rittman BE, editor. In situ bioremediation: when does it work? Washington, DC: National Academy Press. 1993.
5. US Department of Energy. Subsurface contaminants focus area: technology summary. DOE/EM-0296. 1996.
6. Fredrickson JF, Onstott TC. Microbes deep inside the earth. Sci Am 1996:68-73.
7. Ghiorse W. Subterranean life. Science 1997;275:789-90.
8. Lovley D, Chappelle F. Deep subsurface microbial processes. Geophysics 1995;33:365-81.
9. Vasco DW, Peterson JE, Majer EL. Beyond ray tomography: wavepaths and fresnel volumes. Geophysics 1995;60:1790-1804.
10. Boone DR, Yitai L, Zhao Z, Balkwill DL, Drake GR, Stevens TO, Aldrich HC. Bacillus infernus sp.nov., an Fe(III)-and Mn(IV)-reducing anaerobe from the deep terrestrial subsurface. Int J System Bacteriol 1995;45:441-8.
11. Fredrickson JK, McKinley JP, Nierzwicki-Bauer SA, White DC, Ringelberg DB, Rawson SA, Li SW, Brockman FJ, Bjomstad BN. Microbial community structure and sediment: implications for long-term microbial survival. Mol Ecol 1995;4:619-26.
12. Reeves RH, Reeves JY, Balkwill DL. Strategies for phylogenetic characterization of subsurface bacteria. J Microbiol Meth 1995; 21:235-51.
13. Fredrickson JF, Balkwill DL, Drake GR, Romine MF, Ringelberg DB, White DC. Aromatic-degrading Sphingomonas from the deep subsurface. Appl Environ Microbiol 1995;61:1917-22.
14. Fredrickson JF, Brockman FJ, Workman DJ, Li SW, Stevens TO. Isolation and characterization of a subsurface bacterium capable of growth on toluene, naphthalene, and other aromatic compounds. Appl Environ Microbiol 1991;57:796-803.
15. Balkwill DL, Drake GR, Reeves RH, Fredrickson JK, White DC, Ringelberg DB, Chandler DP, Romine MF, Kennedy DW, Spadoni CM. Taxonomic study of aromatic-degrading bacteria from deep terrestrial subsurface sediments and description of Sphingomonas aromaticivoran sp.nov., Sphingomonas subterranea sp.nov., and Sphingomonas stygia sp.nov. Int J System Bacteriol 1997;47:191-201.
16. Stillwell LC, Thurston S J, Schneider RP, Romine MF, Fredrickson JK, Saffer JD. Physical mapping and characterization of a catabolic plasmid from the deep-subsurface bacterium, Sphingomonas sp. Strain F199. J Bacteriol 1995;177:4537-39.
17. Roden EE, Zachara JM. Microbial reduction of crystalline Fe(III) oxides: influence of oxide surface area and potential for cell growth. Environ Sci Technol 1996;30:1618-28.
18. Zachara JM, Smith SC, Kuzel LS. Adsorption and dissociation of Co-EDTA complexes in Fe oxide-containing subsurface sands. Geochim Cosmochim Acta 1995;59:4825-44.
19. Zachara JM, Smith SC, Fredrickson JK. Adsorption and dissociation of Co(II)EDTA2-in subsurface sediments under conditions of bacterial iron reduction. Geochim Cosmochim Acta 1997.
20. Murphy EM, Ginn TR, Chilakapati A, Resch CT, Phillips JL, Wietsma TW Spadoni CM. The influence of physical heterogeneity on microbial degradation and distribution in porous media. Water Resources Res. 1997.
21. Williams V, Fletcher M. Pseudomonas fluorescens adhesion and transport through porous media are affected by lipopolysaccharide composition. Appl Environ Microbiol 1996;62:100-4.
22. Stevens TO, McKinley JP. Lithoautotrophic microbial ecosystems in deep basalt aquifers. Science 1995;270:450-54.
23. Bult CJ, White O, Olsen GJ, Zhou L, Fleischmann RD, Sutton GG, Blake JA, Fitzgerald LM, Clayton RA, Gocayne JD, Kerlavage AR, Dougherty BA, Tomb JF, Adams MD, Reich CI, Overbeek R, Kirkhess EF, Weinstock KG, Merrick JM, Nguyen D, Utterback TR, Kelley JM, Peterson JD, Sadow PW, Hanna MC, Cotton MD, Roberts KM, Hurst MA, Kaine BP, Borodovsky M, Klenk HP, Fraser CM, Smith HO, Woese CR, Venter JC. Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii. Science 1996;273:1058-72.
24. Pace NR. A molecular view of diversity and the biosphere. Science 1997;276:734-40.
25. Lu HP, Xie XS. Single-molecule spectral fluctuations at room temperature. Nature 1997;385:143-6.
26. Nealson K. New microscopic and spectroscopic methods applied to the study of metal precipitates: solution and dissolution at the microscale. Invited presentation. American Society for Microbiology, 97th General Meeting. Miami, FL. May 4-8, 1997. p 157.
27. Dunn RC, Holtom GR, Mets L, Xie XS. Nearfield fluorescence imaging and fluorescence lifetime measurement of light harvesting complexes in intact photosynthetic membranes. J Phys Chem 1994;98:3094-8.
28. Xie XS. Single-molecule spectroscopy and dynamics at room temperature. Acc Chem Res 1996;29:598-606.
29. Xie XS. Probing single molecule dynamics. Science 1994;265:361-4.
Discussant
Thom H. Dunning, Jr.
William R. Wiley Environmental Molecular Sciences Laboratory
Pacific Northwest National Laboratory
Richland, Washington
In the late 1980s, recognizing that the Department of Energy (DOE) would face enormous technical challenges in addressing its energy and environmental missions in the 21st century, Bill Wiley, the director of the Pacific Northwest National Laboratory (PNNL), and his senior management drafted plans for a unique new research facility. This facility, which was intended to be one of DOE's scientific-user facilities, 1 would bring together the scientists and capabilities needed to understand, at a molecular level, the physical, chemical, and biologic processes critical to DOE's environmental missions especially the cleanup of its former weapons-production sites (for example, the Hanford site adjacent to PNNL in southeastern Washington state and the Savannah River site in southeastern South Carolina). On October 1, 1997, this facility, now called the William R. Wiley Environmental Molecular Sciences Laboratory (EMSL), began operation under the sponsorship of DOE's Biological and Environmental Research program. It brings a new level of research capability to bear on the environmental problems facing the DOE.
To address DOE's complex environmental problems, scientists in the EMSL will capitalize on the continuing revolution in the experimental, theoretical, and computational molecular sciences. Using a multidisciplinary approach involving chemists, materials scientists, geochemists, structural and molecular biologists, computer scientists, and applied mathematicians—experimentalists, theoreticians, and computational scientists-resident EMSL staff and visiting scientists will conduct research in
-
The molecular sciences, to provide the fundamental understanding of complex environmental systems necessary to address DOE's environmental problems.
-
Biogeochemistry and chemical processing, to focus this new knowledge on specific environmental problems.
-
Computer science and technology, to provide the infrastructure needed to support widely distributed collaborative-research teams.
The research carried out in the EMSL in the next decade can be expected to have a profound effect on the understanding and solution of the environmental problems that confront DOE, this nation, and the world.
RESEARCH CAPABILITIES IN THE EMSL
The complement of research equipment and laboratory infrastructure in the EMSL can support a broad multidisciplinary effort in the environmental molecular sciences. Three major facilities house specialized and unique capabilities:
-
The High Field Magnetic Resonance Facility, which houses a 750-MHz nuclear magnetic resonance (NMR) spectroscope, 2 600-MHz NMR spectrometers; and numerous 300-, 400-, and 500-MHz NMR spectroscopes. In addition, as of the middle of 1998, we have operational a 900-MHz NMR spectrometer, which was built by Oxford Instruments. These instruments support work in structural biology, interfacial chemistry, and microimaging.
-
The High Field Mass Spectrometry Facility, which houses the world's first 11.5-tesla Fourier-transform-ion cyclotron resonance (FT-ICR) mass spectrometer, 7-tesla and 3-telsa FT-ICR mass spectrometers, and a number of other advanced mass spectrometers. These instruments support a wide range of activities, including combinatorial chemistry and gene sequencing.
-
The Molecular Science Computing Facility (MSCF), which houses a 512-processor IBM Scalable POWER
-
Parallel computer system, a 20-terabyte EMASS hierarchical data-storage system, a state-of-the-art graphics and visualization laboratory, and systems that incorporate IBM's next-generation multiprocessor technology. In addition, the users of the MSCF are supported by a new generation of molecular-modeling software that takes full advantage of the computer systems.
In addition to the above major facilities, the EMSL offers, in 1 setting, a comprehensive collection of state-of-the-art equipment for research in the environmental molecular sciences. The capabilities embodied in this collection can be integrated as needed to address fundamental problems in 4 major categories:
-
Nanostructural materials, for the design and synthesis of model materials for environmental studies.
-
Interfacial structures and composition, for studies of the composition and structure of interfacial regions.
-
Reactions at interfaces, for studies of chemical and radiologic processes at interfaces.
-
Gas-phase monitoring and detection, for the development of new techniques for detecting and monitoring molecular species.
As one of DOE's national scientific-user facilities, the EMSL makes its research instruments available to researchers worldwide for studies in the environmental molecular sciences or related disciplines of national importance. For additional information on how to apply for time on the research instruments in the EMSL, visit our Web site (http://www.emsl.pnl.gov/).
RESEARCH IN THE ENVIRONMENTAL MOLECULAR SCIENCES
I cannot do justice here to the wide range of research being carried out in the EMSL. I will therefore pick a few examples on which to focus. I will use them to illustrate 3 of the major research themes in the EMSL: remediation of contaminated soils and groundwater, adverse health effects of exposure to hazardous chemicals, and processing of high-level wastes.
SINGLE-MOLECULE MICROSCOPY AND THE BIODEGRADATION OF CHLORINATED HYDROCARBONS
The most-prevalent class of contaminant on DOE's weapons-production sites is not, as one might suppose, radionuclides, but rather chlorinated hydrocarbons (CHCs).2 In fact, CHCs—because of their widespread use as solvents, degreasers, and so on—are estimated to constitute a large fraction of the inventory of chemical pollutants in the United States. It has proved very difficult to remediate CHCs. However, a few strains of microrganisms have been found that slowly degrade them, and some of the enzymes responsible for the dechlorination steps have been identified. Deeper insights into the enzymatic processes involved in degradation will be necessary before we will be able to understand the nature of the rate-limiting steps and, if possible, develop or stimulate more-efficient processes.
For several years, Xiaoliang (Sunney) Xie and his collaborators in our laboratory have been developing methods to study single-molecule spectroscopy and dynamics.3 Single-molecule sensitivity has been achieved at room temperature by using a combination of a laser, an inverted fluorescence microscope, and a photon-counting apparatus. The single-molecule measurements have provided unique and detailed information on condensed-phase systems, especially biologic systems. For measurements of single enzyme molecules, the molecules are immobilized in an agarose gel containing 99% water in which the substrate molecules are still mobile. A dilute solution is used to ensure that there is only enzyme molecule in the laser focus field.
For the last 2 yr, Xie, H. P. Lu, and L. Xun have been studying the enzyme responsible for the dechlorination of pentachlorophenol (PCP), pentachlorophenol-4-monooxygenase4 (which has also been found to degrade trichloroethylene, one of the most-abundant contaminants on DOE sites). The active site of this enzyme is flavin adenine dinucleotide, which is fluorescent in its oxidized state. When the enzyme oxidizes the CHC, it gains an electron, and in this state it does not fluoresce. Eventually, the monooxygenase loses the extra electron, and the catalytic
cycle starts anew. Because of this property of the flavoenzyme, Xie and his co-workers have been able to follow the enzymatic reactions of a single molecule, similar to monooxygenase (PCP), in real time and directly measure the rate of the enzymatic reaction (usually referred to as the turnover rate in a catalytic process). The turnover rate is 100 or so per second.5 Similar measurements are now being conducted on PCP to determine the turnover rate of PCP directly.
As important as that observation was, there is more to the story. When Xie and co-workers placed the dechlorinating enzymes in their native state in the agarose gel, they expected to see a steady fluorescence. Instead, they found that the spectral mean of the fluorescence flucuates with time and on the same time scale as the dechlorination process. This phenomenon is hidden in ensemble-averaged measurements. Xie and co-workers believe that the spectral variations are due to conformational changes in the enzyme and that these results confirm the earlier observations of Xue and Yueng6 at Iowa State University. Using a single-molecule technique, those authors showed that the activity of identical copies of enzymes varied by over a factor of 4. They speculated that that might be due to conformational changes in the enzyme and that each conformation might have a different reactivity. If verified in real-time observations, this will be a major advance in our understanding of the detailed mechanism of enzymatic reactions. Clearly, single-molecule microscopy will be an important technology as we try to understand the bioremediation of recalcitrant environmental contaminants.
NMR SPECTROSCOPY AND THE STRUCTURE OF REPAIR ENZYMES
The EMSL has made a major investment in NMR spectroscopy. There are many applications of NMR spectroscopy, but I will discuss only 1 here—structural biology. In our studies of the biodegradation of contaminants, it is often critical to know the structure's key enzymes to be able to understand the mechanism of the degradative process. Similarly, the structures of the biomolecular species involved in the repair of DNA damaged by exposure to hazardous chemicals are critical to understanding the detailed molecular processes involved in the sequence of events that lead from exposure to uptake to damage to cancer (or other adverse health effect). Fourth-generation NMR spectroscopy provides the resolution equivalent to 2-2.5 Å X-ray spectroscopy. In addition, it does not require the protein to be crystallized, an often-daunting problem itself, and the species can be studied in aqueous solution, a far more natural setting than the crystalline environment.
The initial focus of the research effort in the EMSL is on understanding of the molecular processes involved in nucleotide excision repair (NER) of damaged DNA. NER is the process most commonly associated with damage resulting from exposure to carcinogenic chemicals. It involves several steps, such as unraveling of the DNA, location of the damage, excision of the altered nucleotides, and resynthesis of the correct DNA sequence, and each step can involve proteins with well-defined roles. The logical place to begin is with an understanding of the underlying structural basis of the initial recognition event and with the first structural signal to the proteins involved in the DNA-repair process. In NER, recognition of DNA damage is the responsibility of the protein XPAC (xeroderma pigmentosum group A complement). XPAC then recruits RPA (replication protein A) to the damaged site, and both are involved in recruiting other proteins: XPA recruits a heterodimer of ERCC1 (excision-repair cross-complementing protein 1) and xerodoma pigmentosum group F protein, which acts as a 5' endonuclease, and RPA recruits xerodoma pigmentosum group G protein, which serves as a 3' endonuclease. Together, the 2 endonucleases excise the damaged DNA patch in preparation for replacement with newly synthesized DNA.
To begin, Buchko and Kennedy determined the structure of a synthetic 40-residue peptide corresponding to the zinc-binding core of XPAC. 7 They have now completely assigned the backbone of the ''minimal (damaged-DNA) binding domain,'' referred to as MBD,8 and completely determined its secondary structure, including its topology. The residues in the zinc-binding domain of XPAC and GATA-1 (an erythroid chick transcription factor) make contact with the major groove in DNA. GATA-1 contains several hydrophobic side-chain amino acids that allow it to recognize a thymine-rich DNA sequence and bind tightly to it through the formation of several hydrophobic patches. XPAC, on the other hand, contains mostly hydrophilic residues. Kennedy and co-workers speculate that the zinc-binding domain of XPAC retains its ability to interact with double-stranded DNA but that its distribution of amino acids is such that it does not bind tightly to any specific DNA sequence—which would be a fatal flaw in an enzyme responsible for detecting DNA damage.
Although Kennedy and co-workers are studying XPAC, their goal is to determine the structure of XPAC-MBD bound to damaged DNA. Determining the full structure of the MBD of XPAC is certainly within range of the 750-MHz NMR spectrometer now in operation in the EMSL. However, determination of the structure of the complete XPAC could well require the 900-MHz NMR spectrometer. Studies of the complexes of XPAC with DNA or other proteins will almost certainly require the increased sensitivity and resolution of the 900-MHz instruments.
COMPUTATIONAL MOLECULAR SCIENCE AND MATERIALS FOR SEPARATING RADIONUCLIDES
The production of plutonium during the Cold War generated about 10 × 103 gal of high-level radioactive waste, which is currently stored in large underground tanks at the Hanford and Savannah River sites. The high-level wastes are complex mixtures of highly radioactive elements, such as cesium-137 and strontium-90; long-lived radionuclides, such as americium-241, plutonium-239, and technetium-99; and RCRA-listed chemical wastes, such as heavy metals, solvents, chelating agents, and nitrates. Cleanup costs could be sharply reduced if the small fraction of radioactive material (about 10-100 g per metric ton of waste) could be separated from the enormous volume of low-level waste, inasmuch as the former is destined for storage in an expensive underground repository where it must be immobilized over geologic spans of time (10,000 yr or more).
A number of metal-separation processes (such as solvent extraction, supercritical fluid extraction, liquid membranes, and ion exchange) involve complexing of the targeted metals with organic or inorganic ligands. For optimal performance, the ligand must satisfy a number of constraints, each of which is challenging but all of which together present a daunting exercise in materials design. When I joined the EMSL project in late 1989, Dave Feller and I examined the use of computational modeling techniques to help design a new generation of separation materials that would improve on the crown ethers that were being considered for extraction of cesium and strontium from tank wastes. We concluded that, with the computational resources available at the time, it was not feasible to address even the simplest—predicting the preference of 18-crown-6 for the potassium ion over the other alkali ions (lithium, sodium, rubidium, cesium, and francium). In fact, attempts to carry out such calculations met with failure.
Fortunately, advances in computer technology (processor speed, memory density, and disk capacity) continued unabated during the early 1990s, and by 1994 Glendening and co-workers9 were able to carry out first-principles quantum-mechanical calculations on a realistic model of the exchange reaction:
M+ refers to any other alkali ion. The calculations, which took months to do on the computers available at DOE (a Cray C-90 supercomputer at the National Energy Research Supercomputing Center), predicted that the reaction was uphill (endothermic) for all ions ![]() , roughly reproducing the experimental data on endothermicity. More important, they showed that the preference for K+ over the other alkali ions was the result of a subtle interplay between the solvation of the ion by water molecules and its "solvation" by the crown ether. The calculations also provided a wealth of important new information on the structure and binding energies of the gas-phase M+ (18-crown-6) complexes.
, roughly reproducing the experimental data on endothermicity. More important, they showed that the preference for K+ over the other alkali ions was the result of a subtle interplay between the solvation of the ion by water molecules and its "solvation" by the crown ether. The calculations also provided a wealth of important new information on the structure and binding energies of the gas-phase M+ (18-crown-6) complexes.
This experience, as well as others, convinced us of the need to develop a new generation of molecular-modeling software that could take advantage of massively parallel computer architectures. Parallel computing is the most feasible way to obtain the substantial increases in computing capability needed to tackle the more-complex molecules relevant to the separation of radionuclides, but it has posed a substantial challenge. In the early 1990s, many fundamental issues related to the design of massively parallel computers had yet to be settled. In addition, the explicit handling of communication required for massively parallel computers forced the scientific-software developer to manage the data used and generated in the calculation explicitly—a daunting task when dealing with the million lines of FORTRAN code typical of state-of-the-art molecular-modeling packages.
To tackle this problem, we assembled a team of theoretical chemists (Robert Harrison, Ricky Kendall, Martyn Guest, Jeff Nichols, Tjerk Straatsma, Michel Dupuis, and Tony Hess), computer scientists (Jarek Nieplocha and
Matt Rosing), and applied mathematicians (Rik Littlefield and George Fann). The group first designed the overall architecture for the molecular-modeling software, which is now known as NWChem.10 The software was designed to be highly modular, with many of the "services" needed by the molecular-modeling applications, such as the computation of integrals over basic functions, being provided through application programmer interfaces (APIs). The extensive use of APIs is critical: It allows new techniques to be introduced into NWChem with minimal effort. In addition, NWChem uses a hardware-abstraction layer—the software-development toolkit consisting of global arrays, CHEMIO, memory allocator, and run-time library—to provide isolation from the computer hardware; only the software-development toolkit must be ported from one type of machine to the next. The net result is a molecular-modeling package that provides high performance, is scalable to hundreds if not thousands of processors, and is portable over a wide range of computer architectures (IBM SP, Cray T3D and T3E, Kendall Square, Intel Paragon, and even a collection of work stations).
Harrison and co-workers have carried out calculations on a derivative of 18-crown-6 synthesized by Li and Still.11 The selectivity of this crown ether can be "tuned" by changing various functional groups in the molecule, but at the price of a far-more-complex molecular structure. It is not feasible to do high-level calculations on this molecule with a conventional supercomputer, and even the calculations reported here, whose aim was to quantify the scalability of NWChem, rather than to obtain highly accurate results, would pose a substantial challenge. For a single processor of the IBM Scalable POWER Parallel System (POWER2 processors), nearly 400 h of computer time is required for the solution of the matrix Hartree-Fock (HF) equations (899 basis functions using the aug-ccpVDZ basis set12). With 180 processors, less than 1 h is required; that is, the use of 180 processors speeds up the calculation by a factor of 400! This phenomenon is called superlinear speedup and is caused by increase in the total amount of memory and local disk storage associated with the increase in the number of processors. For solving the HF equations, NWChem uses an adaptive algorithm that takes advantage of the additional disk storage, becoming more and more effective as the amount of available local disk storage increases. With the next-generation IBM computer (POWER2 SUPER processors) just installed in the EMSL, it is estimated that the same calculation can be done in 15 min. As can be seen even in this rather simple example, the developers of NWChem pioneered many innovations to take advantage of the power offered by massively parallel computer systems. NWChem greatly expands the range and accuracy of modem molecular-modeling approaches and will have a large influence, not only in environmental molecular science, but also in chemistry, materials science, and molecular biology.
CONCLUSIONS
DOE has made a commitment to remediation of its sites—a commitment that recognizes the effort as a formidable, but achievable, endeavor. By constructing and equipping the EMSL at the PNNL, DOE has dramatically enhanced its existing technical base, supporting development of the new knowledge required to clean up its sites and providing a powerful new resource to assist the scientific community in meeting the nation's environmental challenges. The resulting effort will lead to the development of technologies that span the full range of our environmental needs—from remediation of existing hazards to prevention of future environmental insults.
Research in the EMSL is now supported not only by the Office of Energy Research, but also by the Office of Environmental Management, primarily through the Environmental Management Science Program. The major focus of this work is on the molecular processes involved in the processing and storage of high-level wastes, the cleanup of contaminated soils and groundwater, and the health effects arising from exposure to hazardous chemicals. In the short term, this combination of basic, directed basic, and applied research can be expected to greatly enhance our understanding of the issues associated with the remediation of DOE's sites. In the long term, it can be expected to lead to the development of the new technologies that will be needed to achieve DOE's oft-stated goal of "better, faster, cleaner."
The EMSL will also help to forge the emerging scientific discipline of environmental molecular science, a multidisciplinary science that is still in its infancy. By drawing scientists from the chemical, materials-science, geoscience, structural-biology, and molecular-biology communities to the molecular-level issues that underlie environmental remediation, waste processing and storage, and human-health effects, it should be possible to
develop a sound strategy for addressing the nation's environmental problems. Research in the molecular sciences will also have far-ranging effects on many other DOE missions, including energy efficiency and national defense. Also, the new approach to collaborative research embodied in the EMSL—which makes full use of the latest advances in computing and communication technologies—will serve as a model for focusing the country's scientific resources on other vital national issues.
ACKNOWLEDGMENTS
I thank Sunney Xie, Michael Kennedy, David Feller, and Robert Harrison for providing the research material discussed here. I dedicate this paper to the memory of William R. Wiley, for his vision and steadfastness, and to Michael L. Knotek, for teaching us how to identify and cope with the myriad issues in establishing the EMSL. Without them, there would not have been an EMSL.
"Ultimately a hero is a man who would argue with the gods, and so awakens the devils to contest his vision."
(special preface to the 1st Berkeley Edition of The Presidential Papers [1976]). I also dedicate this article to the staff of the William R. Wiley Environmental Molecular Sciences Laboratory, in whose hands the future of the EMSL rests.
REFERENCES
1. Koomanoff FA. Scientific User Facilities, A National Resource. U.S. Department of Energy, Office of Energy Research, Washington, DC, 1994.
2. Riley RG, Zachara JM, Wobber FJ. Chemical Contaminants on DOE Lands and Selection of Contaminant Mixtures for Subsurface Science Research, U.S. Department of Energy, Office of Energy Research, Washington, DC, 1992 (DOE/ER-0547T).
3. Xie XS. Single molecule spectroscopy and dynamics at room temperature. Accounts of Chemical Research 1996; 29:598—606
4. Xun L, Topp E, Oser, CS. Confirmation of Oxidative Dehalogenation of Pentachlorophenol by a Flavobacterium Pentachlorophenol Hydroxylase. Journal of Bacteriol 1992; 174: 5745-5747.
5. Lu HP, Xun L, Xie XS. Manuscript in preparation.
6. Xue QF, Yeung ED. Difference in the Chemical Activity of Individual Molecules of an Enzyme. Nature 1995; 373: 681-683.
7. Buchko GW, Kennedy MA. Human Nucleotide Excision Repair Protein XPA: 1H NMR and CD Solution Studies of a Synthetic Peptide Fragment Corresponding to the Zinc Binding Domain (101-141). Journal of Biomolecular Structure and Dynamics 1997; 14:677-690.
8. Buchko GW, Ni S, Thrall BD, Kennedy MA. Human Nucleotide Excision Repair Protein XPA: Expression and NMR Backbone Assignments of the 14.7 kD Minimal Damaged-DNA Binding Domain (M98-F219). Journal of Biomolecular NMR 1997; in press.
9. Glendening ED, Feller D, Thompson MA. An Ab Initio Investigation of the Structure and Alkali Metal Cation Selectivity of 18-Crown-6. Journal of the American Chemical Society 1994; 116:10657. [See also, Feller D, Thompson MA, Kendall RA. A Theoretical Study of Substituent Effects on the Binding Specificity of Crown Ethers. Journal of Physical Chemistry 1997; in press.]
10. Anchell J, Apra E, Bernholdt D, Borowski P, Clark T, Clerc D, Dachsel H, Deegan M, Dupuis M, Dyall K, Farm G, Fruchtl H, Gutowski M, Harrison R, Hess A, Jaffe J, Kendall R, Kobayashi R, Kutteh R, Lin Z, Littlefield R, Long X, Meng B, Nichols J, Nieplocha J, Rendall A, Stave M, Straatsma T, Taylor H, Thomas G, Wolinski K, Wong A, NWChem, A Computational Chemistry Package for Parallel Computers, Version 3.0. 1997. Pacific Northwest National Laboratory, Richland, Washington 99352-0999, USA.
11. Li G, Still WC. An 18-Crown-6 Derivative with Only One Conformation. Journal of the American Chemical Society 1993; 115: 3804-3805.
12. Kendall RA, Dunning TH Jr., Harrison RJ. Electron affinities of the fast-row atoms revisited. Systematic basis sets and wave functions. Journal of Chemical Physics 1992; 96: 6796-6806.
Discussant
Ronald W. Harvey
US Geological Survey
Boulder, Colorado
The importance of subsurface bacterial transport in the spread of waterborne diseases has long been recognized. Indeed, the first indirect evidence of subsurface bacterial transport occurred in 1854, when it was observed that a cholera epidemic in central London was the result of bacterial contamination of a public well.1 Injection and recovery tests involving the direct addition of bacteria to freshwater aquifers were conducted as early as the 1890s.2,3 The purpose of early additions of bacteria was to provide a convenient tracer for following the movement of groundwater through karstic, or fractured-rock, terrain. Indicator bacteria, such as coliforms, have been added to the subsurface since the 1930s4 to investigate the transport potential of pathogenic bacteria near water-supply wells. Only within the last 10 yr have specific populations of bacteria been added to aquifers to enhance the degradation of subsurface contaminants. However, many current mechanistic studies of subsurface bacterial transport are being carried out to learn more about the feasibility of using nonindigenous or waste-adapted bacteria in the cleanup of subsurface contamination.
IMPORTANCE OF SUBSURFACE BACTERIAL TRANSPORT AT DEPARTMENT OF ENERGY SITES
Subsurface bacterial transport is potentially important for engineered remediation and natural attenuation at various contaminated sites at Department of Energy (DOE) facilities, especially those subject to subsurface contamination with organic chemicals. The role of bacterial transport at such sites can be complex and can involve various effects, such as the following:
-
Seeding of contaminated zones with bacteria that are genetically engineered for or have adapted to a particular contaminant.
-
Facilitated transport of hydrophobic or surface-active contaminants by mobile bacteria, which might enhance the spread of these chemicals.
-
Cotransport of bacteria with dissolved organic contaminants, which results in longer contact between the contaminants and the populations that affect their degradation.
-
Increased dissemination of genetic information, particularly if introduced bacteria carry genes that confer more-efficient breakdown of organic contaminants.
Little is known about the last 2 mechanisms, and more information is needed about all 4. More research on the role of bacterial transport in the ecology of subsurface microbial communities that ultimately degrade contaminants is also needed.
FUTURE RESEARCH CHALLENGES
Geohydrologic Complexities
Many of the field experiments in which labeled bacteria were injected directly into an aquifer with a conservative tracer (nonreactive solute) involved rather homogeneous sandy deposits.5-7 In well-sorted sandy deposits, the bacteria injected into the aquifer are assumed to follow the same flow paths as the conservative tracer. Therefore, the conservative tracer can be used to predict what points in the acquirer need to be sampled in order to capture breakthrough of labeled bacteria. Also, the concentration histories of the conservative tracer provide important information for the hydrologic portion of a predictive transport model and allow for a reasonable estimate of bacterial retardation.8
However, most contaminated DOE sites are characterized by substantial heterogeneity that can involve fracture-flow or adjacent strata that vary in hydraulic conductivity by a factor of several powers of 10. In heterogeneous aquifers dominated by preferential flow paths, bacteria can be excluded on the basis of size and porosity from substantial fractions that are accessible to the conservative tracer. Injection and recovery investigations involving fractured-rock aquifers suggest that transport of bacteria in fracture-flow environments can be much faster than that of the conservative tracer.9 That phenomenon has also been demonstrated in the laboratory10 for highly stratified formations in which the major conductive zone is next to a layer characterized by porosity too fine for access by microorganisms.
On the larger scales that are typically involved in natural attenuation or engineered restoration of contaminated aquifers, even the relatively homogeneous sites look heterogeneous, at least for the purposes of predictive modeling of bacterial transport. It is evident that the simple deterministic models that are typically used to describe small-scale (<<10 m) bacterial transport in saturated porous media are not adequate for larger-scale application at most, if not all, contaminated DOE sites. That is, in part, because of the aforementioned geohydrologic complexities that often become manifest at larger scales. Bacterial transport models are needed that can account for physical variability in aquifer structure, perhaps in a manner consistent with the application of stochastic theory to describe mathematically the large-scale (>100 m) movement of conservative solutes in sandy aquifer sediments.11
Reconciling Discrepancies Between Field And Laboratory Results
Much of the detailed information about bacterial transport behavior in saturated granular media derives from flow-through column experiments involving repacked subsurface material. A number of the more-recent studies on this subject were sponsored by DOE.7,12-14 Typically, filtration theory is used to describe the removal of unattached bacteria being advected downgradient through granular media. In the filtration model, a bacterium's affinity for attachment to the grain surfaces that it comes into contact with is quantified with a collision efficiency (a) factor,5 whose value can vary from 0 to 1.15 Recent findings from flow-through column experiments involving bacteria7,12-14 and bacteriophage16 transport through representative granular media under reasonable chemical and hydrologic conditions suggest little microbial mobility and collision efficiencies of 10-2 to 100. In contrast, recent field observations of bacteria5,17 and bacteriophage18 transport through aquifer sediments suggest substantial mobility with corresponding collision efficiencies of 10-4 to 10-2.
The discrepancy between field and laboratory results can be explained, in part, by the destruction of pore structure when subsurface soil is repacked into columns. It has been demonstrated that, when intact subsurface soils are repacked, bacterial transport is greatly diminished because of the destruction of preferential flow paths.19 It has also been shown that in situ transport of bacteria through undisturbed aquifer sediments can be different from that in core material taken from the same location.20 A major advantage of flow-through columns is that they offer a much greater degree of control over experimental conditions. However, the apparent discrepancies between field and laboratory results, and the growing recognition that processes that control subsurface microbial transport behavior can be interrelated and operate on spatial and temporal scales that are not conducive for laboratory study, emphasize the need for more in situ transport studies. Accurate assessment of the potential role of bacterial transport in the restoration of contaminated groundwater at many DOE sites might require injection-and-recovery experiments on scales that are larger than those of previous studies. However, because much of the emphasis over the last 2 decades has been on flow-through column experiments, development of methods for conducting large field-scale investigations of subsurface bacterial transport has lagged.8
Biologic Factors
Most of the previous research on the controls of subsurface bacterial transport has focused on physical and chemical factors. Immobilization at stationary surfaces is a major determinant of the extent to which bacteria move within aquifers. However, many of the other important determinants of a bacterium's fate and transport in aquifers are biologic,21 and the biologic controls of subsurface bacterial transport have been largely ignored. Although protozoa are known to be common features of both shallow22 and deep23 groundwater habitats, their role
in the transport of bacteria through aquifers has not been well studied. These eukaryotic predators of bacteria in the subsurface appear to be particularly abundant ( dry weight) in organically contaminated aquifer sediments.24,25 At least one organically contaminated sandy aquifer yielded evidence that protists might be more efficient at removing unattached bacteria being advected downgradient than are the organic and mineral coatings on the sediment grains.26 Also, growth of bacteria being transported downgradient in organically contaminated aquifers can be substantial and can offset the losses that occur as a result of attachment.21
dry weight) in organically contaminated aquifer sediments.24,25 At least one organically contaminated sandy aquifer yielded evidence that protists might be more efficient at removing unattached bacteria being advected downgradient than are the organic and mineral coatings on the sediment grains.26 Also, growth of bacteria being transported downgradient in organically contaminated aquifers can be substantial and can offset the losses that occur as a result of attachment.21
The dearth of information on the importance of biologic controls on subsurface bacterial transport is due largely to the fact that many aspects of the subsurface ecosystem are so poorly understood that their effects on bacterial transport are not fully appreciated. A number of the biologic controls are interrelated and difficult to describe mathematically; this makes their study in field experiments difficult. However, the effects of several factors, such as microbial competition, predation, parasitism, and growth, on subsurface bacterial transport are ideal candidates for studies sponsored by the Biological and Environmental Research program that could build on the subsurface ecologic information already collected in previous research.
REFERENCES
1. Price M. Introducing groundwater. New York: Chapman & Hall; 1996. 278 p.
2. Abba F, Orlandi B, Rondelli A. Über die filtrationskraft des bodens und die fotschewemmung von bakterien dutch das grundwasser. Z Hyg Infekt Krankh 1898;31:66-84.
3. Pfuhl E. Über die verschleppung von bakterien durch das grundwasser. Z Hyg Infekt Krankh 1897;25:549-54.
4. Caldwell EL, Parr LW. Ground water pollution and the bored-hole latrine. J Infect Dis 1937;61:148-83.
5. Harvey RW, Garabedian SP. Use of colloid filtration theory in modeling movement of bacteria through a contaminated sandy aquifer. Environ Sci Technol 1991; 25:178-85.
6. Bales RC, Li S, Maquire KM, Yahya MT, Gerba CP, Harvey RW. Virus and bacteria transport in a sandy aquifer at Cape Cod, MA. Ground Water 1995;33:653-61.
7. DeFlaun M, Murray CJ, Holben W, Scheibe T, Mills A, Ginn T, Griffin T, Majer E, Wilson JL. Preliminary observations on bacterial transport in a coastal plain aquifer. FEMS Microbiol Rev. [In press]
8. Harvey RW. In situ and laboratory methods to study subsurface microbial transport. In: Hurse CJ, Knudsen CR, McInerney MJ, Stenzenback LD, Walter MV, editors. Manual of environmental microbiology. Washington, DC: ASM Press; 1997. p 586-9.
9. Champ DR, Schroeter J. Bacterial transport in fractured rock: a field-scale tracer test at the Chalk River Nuclear Laboratories. Water Sci Technol 1988;20:81-7.
10. Bales RC, Gerba CP, Grondin GH, Jensen SL. Bacteriophage transport in sandy soil and fractured tuff. Appl Environ Microbiol 1989;55:2061-7.
11. Garabedian SP, LeBlanc DR. Large-scale natural gradient tracer test in sand and gravel, Cape Cod, Massachusetts. 2. Analysis of spatial moments for a nonreactive tracer. Water Resour Res 1991;27:911-24.
12. Johnson WP, Martin MJ, Gross MJ, Logan BE. Facilitation of bacterial transport through porous media by changes in solution and surface properties. Colloids Surfaces A. Physicochem Eng Aspects 1966;107:263-71.
13. McCaulou DR, Bales RC, McCarthy JF. Use of short-pulse experiments to study bacteria transport through porous media. J Contam Hydrol 1994;15:1-14.
14. McCaulou DR, Bales RC, Arnold RG. Effect of temperature-controlled motility on transport of bacteria and microspheres through saturated sediment. Water Resources Res 1995;31:271-80.
15. Kinoshita T, Bales RC, Maguire M, Gerba CP. Effect of pH on bacteriophage transport through sandy soils. J Contam Hydrol 1993;14:55-70.
16. Yao K, Habibian MT, O'Melia CR. Water and waste water filtration: concepts and applications. Environ Sci Technol 1971;5:1105-12.
17. Harvey, RW. Microorganisms as tracers in groundwater injection and recovery experiments: a review. FEMS Microbiol Rev. [In press]
18. Pieper AP, Ryan JN, Harvey RW, Amy GL, Illangasekare TH, Metge DW. Transport and recovery of bacteriophage PRD1 in a sand and gravel aquifer: effect of sewage-derived organic matter. Environ Sci Technol 1997;31:1163-70.
19. Smith MS, Thomas GW, White RE, Ritonga D. Transport of Escherichia coli through intact and disturbed soil columns . J Environ Qual 1985;14:87-91.
20. Harvey RW. Transport of bacteria in a contaminated aquifer. In: Mallard GE, Ragone SE, editors. US Geological Survey Water Resources Investigations Report 88-4220.
21. Harvey, RW. Parameters involved in modeling movement of bacteria in groundwater. In: Hurst C J, editor. Modeling the environmental fate of microorganisms. Washington, DC: American Society for Microbiology. 1991. p 89-114.
22. Sinclair JL, Ghiorse WC. Distribution of protozoa in subsurface sidements of a pristine groundwater study site in Oklahoma. Appl Environ Microbiol 1978;53:1157-63.
23. Sinclair JL, Ghiorse WC. Distribution of aerobic bacteria, protozoa, algae, and fungi in deep subsurface sediments. Geomicrobiol J 1989;7:15-31.
24. Sinclair JL, Kampbell DH, Cook ML, Wilson Jr. Protozoa in subsurface sediments from sites contaminated with aviation gasoline or jet fuel. Appl Environ Microbiol 1993;59:467-72.
25. Novarino G, Warren A, Kinner NE, Harvey RW. Protists from a sewage-contaminated aquifer on Cape Cod, Massachusetts, U.S.A. Geomicrobiol J 1994;12:23-36.
26. Kinner NE, Harvey RW, Kazmierkiewicz-Tabaka M. Effects of flagellates on free-living bacterial abundance in an organically-contaminated aquifer. FEMS Microbiol Rev. [In press]
Discussant
James Tiedje
Department of Crop and Soil Sciences
Michigan State University
East Lansing, Michigan
My theme is diversity. Dr. Knotek talked about the hyperthermophiles and about the thermophiles as sort of the center of the tree. But at the opposite end of microbiology are organisms that live in the cold. I will show you a sample site that we work with to look at cold-loving organisms. We worked with a group of Russian scientists who specialized in permafrost soil. There were five sampling sites in a region of northeastern Siberia right off the Arctic Ocean, in the Kalama region near the East Siberian Sea. One of the 2 regions of the world that geologists think has been permanently frozen from the time that the sediments were laid down. In principle, given the depth of these sediments, we can take samples out of the freezer of microbial communities that lived there some 3 million years ago.
We can extract a considerable amount of DNA from these samples. We can see many organisms in these samples varying in size and number in different sites. We stained them with a fluorescent protein stain. They look to be perfectly healthy and are particularly numerous at these sites. We can isolate a few of what we suspect to be there on the basis of the DNA. It is not surprising that if an organism has lived a million years at-12ºC, it will not grow when suddenly put at maybe 10ºC.
Our interest is in learning more about the physiology of these organisms. How do they adapt to that environment? In particular, how do they live for such a long period at such low temperatures? The point is that the cold-loving organisms are of interest, as well as the high-temperature organisms.
We determined the 16-S ribosomal sequence for the DNA extracted from these samples. There was a gram-positive organism that would be similar to the living organisms that we know in the arthrobacter group. But we also saw some gram-negatives. What are the gram-negatives like? If we determine the sequence and compare it with sequences of other organisms, we can see that they fall into clusters. One cluster includes an organism—recently isolated by some people in our group—that oxidized iron. This represents a primitive lifestyle of iron-oxidizing organisms and might have relevance to the recent Martian discoveries as well. That is one example of new diversity of physiologic types.
What generates the high diversity that we find in nature? We extracted DNA from soils of different depths—surface soils, the unsaturated subsoil, and sediments below the water table. The samples were from 2 sites, in Virginia and in Delaware. This work was done collaboratively with Oak Ridge National Laboratory and partially funded by Office of Biological and Environmental Research.
We amplify the 16S genes, cut them with restriction enzymes, and look for different patterns that indicate different genotypes. When we compare the different patterns, nothing is dominant. We looked at some 700 clones from only 5 g of soil; virtually every organism is different. That is a surprising pattern in biology because organisms that compete invariably show dominance—the dominant organism wins. If we go down to the saturated zone, we see dominance. Organisms are winning the competition. But in the saturated zone, organisms can move, as Ron Harvey described, and substrates are equally mobile.
The interesting pattern, because it is unusual, is the top pattern. The unusual soil diversity profile occurs particularly in the top one which we would call a noncompetitive diversity profile; there is a lack of dominance.
In ecology, the hypothesis that explains a noncompetitive profile would include a superabundance of resources, which is unlikely in these soil communities. Resource heterogeneity is a possibility. But modeling studies suggest that the explanation lies in spatial isolation. In these nonsaturated zones, the organisms are spatially isolated. They are living in their own world and therefore are not competing. That has important practical implications if the spatial-isolation hypothesis is supported. It is easier for an alien population to colonize but more difficult for it or any population to become dominant. That indicates something about bioremediation and the expected rates of bioremediation. Overall, biodiversity should be readily preserved in such environments because there are infinite protected niches for those organisms.
Soils should and seem to harbor a huge amount of microbial diversity. The estimates suggest some 10,000 species of bacteria per gram of soil.
Finally, microbial endemism should be the rule, that is, everything is not everywhere. In other words, you have uniquely, locally adapted species, which has never been thought to be the model to apply to microbiology. The spatial-isolation hypothesis has a lot to do with our understanding of biodiversity in microbial communities.
To conclude with an application of biodiversity, I note a case in which nature has not made an organism to do a job that we would like done—to grow on polychlorinated biphenyls (PCBs). Nature has not combined the complement of genes to allow organisms to grow on PCBs. If such an organism did exist in nature, it would try to metabolize PCBs, producing an acyl hyaline compound, which would be extremely toxic and kill the organism immediately. If there were an organism that could grow on PCBs, it would be suicidal.
We have tried to remove chlorines by dehalogenation to avoid ending up with an intermediate that forms a toxic product and then to look to nature for dehalogenase genes that might be useful. We have 3. One is the PCB pathway, which leads to the potentially toxic intermediate. Of the other 2, 1 is a hydrolytic gene, and the other is a reductive dechlorination gene. An oxidative gene, if added to the projected intermediates coming from PCB metabolism, would produce nonchlorinated products, and the organism should grow. And, in fact, we can get organisms to grow on some PCBs.
In one example, we introduced 2 genes into an organism to begin to co-oxidize PCBs. One organism grew on 2-chlorobiphenyl, which is one of the most important congeners after reductive dechlorination of PCBs. And one grew on 2, 4-dichlorylbiphenyl, another important congener from reductive dechlorination. This shows how we can use diversity in nature—in this case, different genes that we put together in an organism—to achieve growth.
How might we implement this? One of our collaborators, Walt Weber, at the University of Michigan, is working with slowly mixed auger systems. The soil remains in situ and is not constantly mixed, but only stirred occasionally. You can move over a site, mix the soil down the stem of the auger, put in organisms or nutrients, and let the organisms do their job. This is an example of the use of diversity in bioremediation.



















