Details in Support of the Risk Exemplar in Chapter 6
In this appendix, the committee details the data and assumptions used in the risk exemplar, described in Chapter 6.
PATHOGENS
The exemplar includes four enteric pathogens: adenovirus, norovirus, Salmonella, and Cryptosporidium. In the following discussion, each organism is briefly described and an estimated density in secondary effluent for use in the exemplar is provided. Modifications in those densities are then estimated that correspond to each of the scenarios in the exemplar. Finally, the densities are adjusted so that they are in the same form as those used in dose-response testing, and a risk of illness is estimated using quantitative microbial risk estimation methodology (Haas et al., 1999).
Pathogen Occurrence in Secondary Effluents
The information and assumptions used to estimate pathogen occurrence in undisinfected secondary wastewater effluent as a starting point for the risk exemplar is discussed in this section and summarized in Table A-1. Pathogen reduction from subsequent disinfection and treatment steps is discussed in the next section.
Adenovirus
Adenovirus is a waterborne pathogen that has been associated with recreation-related outbreaks in the United States. It causes a large spectrum of human diseases from diarrhea to eye and throat infections (Jiang, 2006; Mena and Gerba, 2008). Quantitative data on adenovirus occurrence in water and wastewater are available in the current literature, because their occurrence is often used as a marker for human viral contamination in waters. The dose-response model for this virus has also been developed previously based on epidemiological studies (Haas et al., 1999); thus, it is an organism for which quantitative risk assessment is possible.
TABLE A-1 Estimated Pathogen Densities in Secondary Effluent
| Organism | Concentration |
| Adenovirus | 5,000 gc/L |
| Norovirus | 10,000 gc/L |
| Salmonella | 500 cfu/L |
| Cryptosporidium | 17 oocysts/L |
Human adenovirus occurrence data in the exemplar were collected from peer-reviewed literature, which used molecular biology–based genome quantification methods (He and Jiang, 2005; Albinana-Gimenez et al., 2006; Bofill-Mas et al., 2006; Haramoto et al., 2007; Katayama et al., 2008; Fong et al., 2010; Schlindwein et al., 2010). Reported densities vary over a wide range, between 1 and 105 genome copies/liter (gc/L). A density of 5 × 103 gc/L, which falls in the most frequently reported range, was chosen by the committee as a typical concentration in secondary effluent.
Although the genome-based method is sensitive at detecting viral presence, it does not provide information on viral infectivity; thus the presence of a genome
is not synonymous with the presence of an infectious unit (IU). Dose-response studies were conducted using tissue culture assays for quantification of IU. There is limited quantitative information on the side-by-side data for IUs and genome copies although it is generally known that infectivity decays more rapidly than does the density of genome copies (R.A. Rodriguez et al., 2009). Based on a single report (He and Jiang, 2005), where three side-by-side polymerase chain reaction (PCR) and tissue culture assays were performed on adenovirus isolated from secondary effluent, it is estimated that the ratio between genome copies and infectious units is approximately 1,000:1. Thus, genome count densities estimated for adenovirus for each scenario were reduced by three orders of magnitude to convert to IUs during the risk estimation process.
Norovirus
Norovirus is one of the most important enteric viruses for both waterborne and foodborne outbreaks in the United States. Several recent studies have focused on occurrence of this virus in water and wastewater (Pusch et al., 2005; Haramoto et al., 2006; Katayama et al., 2008; Nordgren et al., 2009; Victoria et al., 2010). In these studies, the density of the norovirus genome varies over a wide range with densities as high as 107 gc/L reported in raw sewage. Based on the published literature, a density of 104 gc/L is estimated to be the median occurrence in secondary effluent. Once again, although the genome-based method is sensitive at detecting the presence of copies of the genome of the virus, it does not provide information on viral infectivity. Norovirus has not been successfully cultivated using conventional tissue culture methods, and so no work is available to establish the ratio between genome density and IU density.
A dose-response model for norovirus was used based on the study by Teunis et al. (2008), using the estimate for single unaggregated virus. Because norovirus has not been successfully cultivated in vitro, these studies were conducted using fresh virus and the genome count quantified by PCR. Published work has shown that the fraction of genome copies that are infectious drops rapidly in the environment (R.A. Rodriguez et al., 2009). Thus, for the purposes of this exemplar, the same 1,000:1 was applied before risk estimation.
Salmonella
Salmonella has long been a well-studied waterborne enteric pathogen. The concentration of this microorganism in raw sewage ranges between 102 and 104 cfu/100 mL (Asano et al., 2007). Taking the average of these two and assuming the same 2-log reduction during primary and secondary treatment that normally occurs for Escherichia coli produces an estimate of 5 × 102 cfu/L in secondary effluent for the exemplar. Again, the dose-response model for this organism has been developed previously based on epidemiological studies (Haas et al., 1999).
Cryptosporidium
Cryptosporidium is associated with both drinking water and recreational water outbreaks in the United States. The health significance of this organism has motivated a number of studies to understand its occurrence and persistence in the water environment (Rose et al., 1996; Gennaccaro et al., 2003; Robertson et al., 2006; Lim et al., 2007; Castro-Hermida et al., 2008; Chalmers et al., 2010; Fu et al., 2010). The peer reviewed literature reports a range of Cryptosporidium densities in secondary treated effluents varying with season and geographical location. Studying this literature, a density of 50 oocysts/L is estimated as typical for secondary effluents. However, most of the data on oocyst concentration were determined using the indirect fluorescent-antibody assay (IFA), which also does not directly measure IUs. A study comparing oocyst densities as determined by IFA with IU densities as determined by a focus-detection-method most-probable-number technique in cell culture (Slifko et al., 1999) found a ratio of approximately 3:1 in 18 samples of secondary effluent (Gennaccaro et al., 2003). Using this ratio, a density of 50 oocysts/L produces an estimate of 17 IUs/L in secondary effluent for the exemplar. More than one dose-response model has been developed for this organism (Haas et al., 1999).
Assumptions Concerning Fate, Transport, and Removal
The following is a brief discussion of assumptions made regarding fate, transport, and removal for the pathogens in the exemplar.
Scenario 1—de Facto Reuse
As discussed in Chapter 6, Scenario 1 represents a conventional water supply drawn from a surface water source with a 5 percent contribution from treated wastewater. For this scenario a nonnitrified secondary effluent is assumed to be disinfected with chlorine prior to discharge to bring fecal coliforms from 105/100 mL to 200/100 mL, a 2.7-log reduction (99.8 percent). The exemplar assumes combined chlorine is the active disinfectant. According to Butterfield and Wattie (1946), E. coli, the principal target of the fecal coliform measurement, are generally as or more resistant to combined chlorine than Salmonella spp. (S. dysenteriae). Accordingly, the same 2.7-log reduction was assumed for Salmonella spp. For adenovirus and norovirus, removal was assumed to follow the removal credit for viruses in the surface water treatment rule, which was judged to be negligible. It is also assumed that this limited disinfection has no impact in the viability of Cryptosporidium.
The water treatment plant has been modified to be compliant with the requirements of the Long-Term-2 Enhanced Surface Water Treatment Rule (LT2ESWTR; EPA, 2006a). Assuming no diminishment during transport in the river, the Cryptosporidium contribution from upstream wastewater plants in the exemplar puts the density of oocysts in the water plant’s source water at approximately 0.85 oocyst/L. This classifies the supply in “Bin 2” according to LT2ESWTR, which corresponds to a requirement of 1 log of removal for Cryptosporidium beyond the performance of conventional treatment. Hence, additional treatment to achieve 1- and 2-log removal is required for Cryptosporidium and viruses, respectively.1
For the exemplar it is assumed that the drinking water treatment plant uses free chlorine for primary disinfection and that it has been modified to obtain 1 log of additional inactivation of Cryptosporidium using UV light (required dose, 2.5 mJ/cm2). Under the LT2ESWTR, the inactivation credit for UV at a dose of 2.5 mJ/cm2 is 1 log Cryptosporidium and negligible for viruses. Thus the 2-log virus inactivation requirement must be met by free chlorine. At a low temperature of 5 °C (a conservative surface water temperature), this corresponds to a C·t of 8 mg-min/L. So the process train is conventional water treatment (coagulation, flocculation, filtration) followed by UV (3 mJ/cm2) and chlorination (8 mg-min/L) and this train will get the full 4-log removal/inactivation credit for both Cryptosporidium and viruses, required by the LT2ESWTR.
In the exemplar, excluding dilution, the overall reduction in Cryptosporidium is assumed to correspond to the 4-log removal required by EPA, and the reductions in adenovirus and norovirus are also assumed to correspond to EPA’s assumptions for 2 logs of physical removal in conventional treatment and an additional 2 logs of inactivation via chlorination (totaling 4-log removal). EPA’s LT2ESWTR does not provide direct guidance on Salmonella spp., and so an independent analysis is required. Salmonella spp. are understood to be more sensitive to free chorine than are E. coli (Butterfield et al., 1943). According to Figure 13-5 in Crittenden et al. (2005), a C·t of approximately 1 mg-min/L is required for 2-log removal of E. coli at 25 °C; thus, a C·t of 8 mg-min/L will achieve a 16-log reduction of E. coli. For the effect of chlorine on Salmonella spp., the exemplar discounts this to an inactivation credit of 4 logs to account for temperature. Exposure to low levels of UV light also affects Salmonella spp. and to some degree adenovirus and norovirus. According to data in a recent Dutch review (Hijnen et al., 2005a), a low-pressure UV dose of 2.5 mJ/cm2 should result in a 1.5-log inactivation of Salmonella spp., a 0.1-log reduction in adenovirus, and a 0.3-log reduction in norovirus. In the exemplar, the effect of UV on the Salmonella spp. is included, and the impact of UV on these viruses is neglected. Thus, the overall water treatment plant removal is 4 logs for Cryptosporidium, 5.5 logs for Salmonella spp., and 4 logs for adenovirus and norovirus. A summary of removal for microorganisms and their resulting densities is given in Table A-2.
_____________
1 Actually, the LT2ESWTR gives conventional drinking water treatment (without disinfection) credit for the physical removal of 2 logs of Cryptosporidium and viruses, respectively. Where Cryptosporidium is concerned, this is a bit confusing because the regulation requires 4-log removal of Cryptosporidium for any alternative process in Bin 2, but requires only one additional log removal for conventional treatment. It appears that the 2-log credit is actually a holdover from the earlier interim enhanced surface water treatment rule, which established the 2-log credit and that EPA expects 3-log removal of Cryptosporidium. For the exemplar it is assumed that the drinking water treatment plant achieves 3-log Cryptosporidium removal and requires UV disinfection to achieve one additional log. An actual plant might make other choices from the microbial treatment toolbox to accomplish similar results.
TABLE A-2 Summary of Log (and %) Removals of Pathogens in Various Steps of Scenario 1
| Process | Adenovirus | Norovirus | Salmonella | Cryptosporidium |
| Disinfection at wastewater treatment plant | 0 (0%) | 0 (0%) | 2.7 (99.8%) | 0 (0%) |
| Dilution in stream | 1.3 (95%) | 1.3 (95%) | 1.3 (95%) | 1.3 (95%) |
| Removal in water treatment | 4.0 (99.99%) | 4.0 (99.99%) | 4.0 (99.99%) | 3.0 (99.9%) |
| Removal by UV | 0 (0%) | 0 (0%) | 1.5 (96.8%) | 1.0 (90%) |
Scenario 2—Soil Aquifer Treatment and Groundwater Recharge
As described in Chapter 7, in Scenario 2, a nitrified and partially denitrified secondary effluent, which has been subjected to granular media filtration, is applied to surface spreading basins with subsequent soil aquifer treatment (SAT). The effluent is not disinfected. It is assumed that the water will remain in the subsurface for 6 months with no dilution from native groundwater. While the assumption of no dilution is in contrast to hydrogeological characteristics of subsurface systems, this condition was selected to assign removal credits only to physicochemical and biological attenuation processes occurring during SAT. Subsequently, the water is abstracted from a deep well, disinfected at the wellhead, and chlorinated prior to consumption, assuming no blending occurs with other source waters in the distribution system. These assumptions describe a scenario where drinking water is consumed that originates 100 percent from reclaimed water after additional treatment using SAT.
Effect of SAT on Virus Removal. During percolation through porous media or groundwater recharge, the removal of pathogens from infiltrating reclaimed water depends primarily on three attenuation mechanisms: straining, inactivation, and attachment to aquifer grains (McDowell-Boyer et al., 1986). Subsurface systems, such as riverbank filtration and SAT have been reported as efficient treatment systems for the removal of microbial contaminants. With respect to virus removal, the field experiments conducted by Schijven et al. (1998, 1999, 2000) are considered a benchmark for removal under relatively homogeneous and steady-state conditions in a saturated sand aquifer. During dune recharge using water that was spiked with bacteriophages (MS2 and PRD1), Schijven et al. (1999) reported a virus reduction of 3 logs within the first 2.4 m and another linear 5-log removal within the following 27 m of transport in the subsurface. Spiking tests with bacteriophages conducted by Fox et al. (2001) under field conditions suggested a 7-log removal over a distance of 100 m. During a deep-well (~300 m below surface) injection study, Schijven et al. (2000) spiked pretreated surface water with bacteriophages (MS2 and PRD1) and observed a 6-log removal within the first 8 m of travel followed by an additional 2-log removal during the subsequent 30 m of travel. These values are well within the range of virus inactivation values reported by others (Dizer et al., 1984; Yates et al., 1985; Powelson et al., 1990). Findings from these field studies demonstrated that infiltration into a relatively homogeneous sandy aquifer can achieve up to 8-log virus removal over a distance of 30 m in about 25 days. Loveland et al. (1996) revealed some of the conditions that favor removal of viruses in the subsurface and concluded that precipitated ferric, manganese, and aluminum oxyhydroxides form positively charged patches on the soil grains. These patches provide favorable attachment sites for negatively charged viruses. Powelson and Gerba (1994) also reported that virus inactivation is more efficient under unsaturated than saturated infiltration conditions. In addition, some studies reported that virus inactivation may be enhanced by microbial activity (Quanrud et al., 2003; Gupta et al., 2009) resulting in the expression of enzymes that are detrimental to other microorganisms (Yates et al., 1987). Considering that these conditions (i.e., biological activity, sequence of unsaturated to saturated conditions, presence of metal oxyhydroxides) are commonly observed in SAT systems and the retention time in the potable reuse case study of the exemplar using groundwater recharge via SAT is 6 months, a conservative removal of 6-log was assumed during SAT for both adenovirus and norovirus.
Effect of SAT on Bacteria Removal. For subsurface treatment, such as SAT and riverbank filtration, several studies have reported efficient inactivation of coliform
bacteria. Havelaar et al. (1995) reported removal in excess of 5 logs for total coliform during transport of impaired river water over a 30-m distance from the Rhine River and over a 25-m distance from the Meuse River to a well. During a deep-well (~300 m below surface) injection study, Schijven et al. (2000) spiked pretreated surface water with E. coli and observed a 7.5-log removal within the first 8 m of travel in the subsurface. During SAT in the Dan Region Project, Israel, Icekson-Tal et al. (2003) measured 5.3-log removal of total coliform and 4.5-log removal of fecal coliform bacteria. Total coliforms were rarely detected in riverbank-filtered waters, with 5.5- and 6.1-log reductions in average concentrations in wells relative to river water (Weiss et al., 2005). The efficient removal of fecal and total coliform bacteria during subsurface treatment and essentially their absence in groundwater abstraction wells after SAT or riverbank filtration was confirmed by various other studies (Fox et al., 2001; Hijnen et al., 2005b; Levantesi et al., 2010). Considering these field and controlled laboratory studies as well as a retention time of 6 months in the subsurface for the surface spreading groundwater recharge case of the exemplar, 6 logs of removal was assumed for bacteria (Salmonella) through SAT treatment in the exemplar.
Effect of SAT on Cryptosoridium. Under the LT2ESWTR (EPA, 2006a), immobilization of Cryptosporidium within granular media, often accomplished by sand or riverbank filtration can result in cost-effective removal of protozoa and other pathogens (Ray et al., 2002a,b; Tufenkji et al., 2002). By meeting certain design standards (i.e., unconsolidated, predominantly sandy aquifer with 25- or 50-ft setback from the river), EPA assigns 0.5-log or 1.0-log removal credits for Cryptosporidium, respectively. Log removal calculations require counts per volume of the same organism in the initial water source (e.g., reclaimed water) and groundwater wells. Given the usually low counts of Cryptosporidium in impaired source waters, log removal studies under ambient conditions are not practical. Bacterial spores, anaerobic clostridia spores, and aerobic endospores are resistant to inactivation in the subsurface, similar in shape to Cryptosporidium but smaller and sufficiently ubiquitous in both impaired surface water and groundwater that log removal can be calculated. Findings from studies investigating the fate of bacterial spores in gravel aquifers suggest a high mobility and similar removal of Cryptosporidium, making bacterial spores adequate surrogate measures.
Findings from various field studies suggest that large removal of anaerobic and aerobic spores occurs during passage across the surface water–groundwater interface, and lesser removal is observed during groundwater transport away from this interface. Havelaar et al. (1995) reported 3.1-log removal of anaerobic spores during transport over a 30-m distance from the Rhine River to a well and 3.6-log removal over a 25-m distance from the Meuse River to a well. Schijven et al. (1998) measured 1.9-log removal over a 2-m distance from a canal. This finding is consistent with field monitoring results from a riverbank filtration site in Wyoming, where Gollnitz et al. (2005) observed a 2-log removal of Cryptosporidium in groundwater wells characterized by flow paths between 6 and 300 m. At a riverbank filtration site at the Great Miami River, Gollnitz et al. (2003) reported a 5-log removal of aerobic spores in a production well located 30 m off the river. Wang et al. (2002) reported 1.7-log removal of aerobic spores over the first 0.6-m distance and 3.8-log removal over a distance of 15.2 m at a riverbank filtration facility at the Ohio River. Less efficient removal of approximately 0.6 logs over a distance of 12 m was reported for transport solely within groundwater (Medema et al., 2000). For an injection experiment in a sandy aquifer at distances relatively far from an injection well, Schijven et al. (1998) observed negligible removal of anaerobic spores over a 30-m distance. Besides straining, inactivation might be important for the attenuation of Cryptosporidium during subsurface treatment. For two Cryptosporidium strains examined, NRC (2000) assumed a 1-log inactivation over 100 days and 180 days (corresponding to an inactivation rate coefficient of 0.023/d and 0.013/d, respectively). Considering these field and controlled laboratory studies as well as a retention time of 6 months in the surface spreading groundwater recharge case of the exemplar (much longer than is the case for any of the preceding citations), a removal credit of 6 log for Cryptosporidium was assumed for SAT treatment.
Effect of Wellhead Chlorination. The exemplar assumes that chlorination is provided at the wellhead in order to achieve a 4-log virus removal credit, and so this
is the removal assigned to adenovirus and norovirus. At 15 °C (an approximate groundwater temperature), this would require a C·t of 4 mg-min/L. Salmonella removal is estimated using the equation in Table 13-3 in Crittenden et al. (2005), and adjusting the log inactivation by a factor of 2 for every 10 °C, results in a removal of 6 logs. No removal is assumed in the exemplar for Cryptosporidium via chlorine. The removals are summarized in Table A-3.
Scenario 3—Reverse Osmosis, Advanced Oxidation, and Deep-Well Injection.
Scenario 3, as discussed in Chapter 6, represents a water supply drawn from a deep well in an aquifer fed by injection of reclaimed water that received secondary treatment by chloramination, microfiltration, reverse osmosis, and high-output low-pressure ultraviolet (UV) light supplemented with hydrogen peroxide (also called advanced oxidation).
Effect of Microfiltration. Olivieri et al. (1999) showed median coliphage removals of 2 logs for microfiltration and 3 logs for ultrafiltration, but for microfiltration, removals as low as 0.1 log were observed on one occasion and removals below 1 log were observed 30 percent of the time. Consequently no virus removal was assumed in the exemplar for microfiltration. There is a great deal of literature on the removal of bacteria and protozoa via membrane filtration. This literature shows virtually complete rejection so long as the membranes remain intact (Jacangelo et al., 1997). Methods used were able to demonstrate between 4 and 5 logs for Cryptosporidium and 7 and 8 logs for bacteria. For the purposes of the exemplar, 99.99 percent removal is assumed for both Salmonella and Cryptosporidium. It should be cautioned that, for specific projects, these removals must be demonstrated for each membrane type and, even then, they cannot be ensured unless monitoring demonstrates that the membranes continue to perform.
TABLE A-3 Summary of Logs (and %) Pathogen Removal Assumed for Processes in Scenario 2
| Process | Adenovirus | Norovirus | Salmonella | Cryptosporidium |
| SAT + 6 mo | 6 (99.9999%) | 6 (99.9999%) | 6 (99.9999%) | 6 (99.9999%) |
| Chlorination at wellhead | 4 (99.99%) | 4 (99.99%) | 6 (99.9999%) | 0 (0%) |
Effect of Reverse Osmosis. In principle, reverse osmosis, which is designed to remove individual ions from water, should completely reject all microorganisms. On the other hand, testing has demonstrated that these organisms can pass through these installations unless special quality control practices, beyond those normally exercised in the desalination community, are undertaken (Trussell et al., 2000). This is particularly true where viruses are concerned because these organisms have been shown to pass through flaws in the membranes themselves (Adham et al, 1998). More limited quality control on the installation of the membranes and O-rings has been shown adequate to manage the rejection of bacteria and protozoa. As a result, a removal credit of 99.99 percent is assumed for both bacteria and Cryptosporidium but a credit of only 97 percent is assumed for viruses because this roughly corresponds to the removal of conductivity through reverse osmosis.2
Effect of UV/H2O2. UV/H2O2 installations in existing projects are designed using low-pressure UV to provide 1.2-log removal of NDMA. It has been demonstrated that this corresponds to a delivered UV dose of approximately 1,200 mJ/cm2 (Sharpless and Linden, 2003). Low doses of peroxide and chloramines (both 3 to 5 mg/L) are also present and these absorb some of the UV; nevertheless, the remaining effective UV dose is nearly 10-fold above the dose specified by EPA for 4-log removal of adenovirus or Cryptosporidium in the LT2ESWTR. Evidence is that Salmonella and norovirus are more easily removed than adenovirus (Hijnen et al., 2005a). Consequently a removal of 6 logs (99.9999 percent) is assumed for all these organisms, and this is thought to be very conservative.
Effect of Deep-Well Injection on Pathogen Removal. The lack of microbial activity and the potential absence of metal oxyhydroxides in deep aquifers recharged with reverse osmosis–treated reclaimed water will provide less favorable conditions for virus removal
_____________
2 Based on data from the first 2 years of operation of the Orange County Water District’s Advanced Water Purification Facility (B. Dunivan, OCWD, personal communication, 2011).
and/or inactivation. Thus, in the exemplar, no removal credit for viruses was considered for the reuse scenario using direct injection into a potable aquifer. Likewise, given the lack of a surface water–groundwater interface in direct injection projects and a rather low inactivation rate in aquifers, no removal credits for Salmonella or Cryptosporidium were assigned for the direct injection process and groundwater travel time. It is noteworthy that these are conservative assumptions, because pathogen inactivation could occur in deep aquifers receiving reverse osmosis permeate.
Effect of Wellhead Chlorination. As described under Scenario 2, wellhead chlorination was assigned a 4-log virus removal credit, and a 6-log removal for Salmonella. No removal is assumed in the exemplar for Cryptosporidium via chlorine. The removals are summarized in Table A-4.
Summary of Results on Pathogen Densities
Using the assumptions and results summarized earlier, calculations were conducted to produce an estimate of the densities of each of the four pathogens studied in the drinking water produced in each of the three scenarios. The results of these calculations are summarized in Table A-5.
Quantitative Microbial Risk Assessment
The pathogen densities shown in Table A-5 can be translated into risk of illness using the methodologies for quantitative risk assessment summarized in Chapter 5. Table A-6 summarizes the coefficients derived from the literature in order to facilitate those calculations, as well as the pertinent dose-response model equations.
TABLE A-4 Summary of Logs (and %) Pathogen Removal Assumed for Processes in Scenario 3
| Process | Adenovirus | Norovirus | Salmonella | Cryptosporidium |
| Microfiltration (MF) | 0 (0%) | 0 (0%) | 4 (99.99%) | 4 (99.99%) |
| Reverse osmosis (RO) | 1.5 (97%) | 1.5 (97%) | 4 (99.99%) | 4 (99.99%) |
| UV/H2O2 | 6 (99.9999%) | 6 (99.9999%) | 6 (99.9999%) | 6 (99.9999%) |
| Deep-well Injection + 6 mo | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) |
| Chlorination at wellhead | 4 (99.99%) | 4 (99.99%) | 6 (99.9999%) | 0 (0%) |
Table A-7 summarizes the quantitative microbial risk assessment in three parts for the three scenarios. Table A-7A details the pathogen densities at the point of exposure (i.e., the tap). The virus densities in Table A-7A were used to compute the daily risk (based on a daily consumption of 1 L) using equations (1) or (2) as appropriate for the organism being considered. Table A-7B shows the estimated levels of excess illness that result from the drinking water from a single exposure (1-L consumption). A consumption of 1 L/d is used for consumption of unboiled water as contrasted with the consumption of 2 L/d used for total consumption (Roseberry and Burmaster, 1992).
TRACE ORGANIC CHEMICALS
For potable reuse projects, there is growing concern among stakeholders and the public about potential adverse health effects associated with the presence of trace organic chemicals in reclaimed water. Reclaimed water can contain thousands of chemicals originating from consumer products (e.g., household chemicals, personal care products, pharmaceutical residues), human waste (e.g., natural hormones), industrial and commercial discharges (e.g., solvents, heavy metals), or chemicals that are generated during water treatment (e.g., disinfection byproducts) (see Chapter 3). For the risk exemplar, 24 chemicals were selected that represent different classes of contaminants (i.e., nitrosamines, disinfection by-products, hormones, pharmaceuticals, antimicrobials, flame retardants, and perfluorochemicals).
Chemical Occurrence in Secondary Effluents
For disinfection byproducts in secondary effluents, data were obtained from Krasner et al. (2008), which reported occurrence of unregulated and regulated disinfection byproducts for secondary wastewater treatment processes with various disinfection practices for a range of different geographical regions of the United States. These datasets were validated and augmented with results from field monitoring efforts reported by Snyder et al. (2010a) and Dickenson et al. (2011). Hormones and pharmaceutical occurrence data were adopted
TABLE A-5 Summary of Exemplar Calculations to Establish Pathogen Levels in Drinking Water for the Three Scenarios
| Scenario 1 De facto Reuse: Secondary effluent, disinfected with chlorine, diluted 95%, conventional water treatment | ||||||||||
| 2° Effluent Concentration |
Removal through Disinfection |
Discharge Concentration |
95% dilution |
Concentration in River |
Die-off & Predation |
WTP Influent Concentration |
Removal in Conventional WTP |
Removal by UV |
Drinking Water Concentration |
|
| Norovirus | 10,000 gc/L | 0% | 10,000 gc/1 | 95% | 500 gc/L | 0% | 500 gc/L | 99.99% | 0.0% | 0.050 gc/L |
| Adenovirus | 5,000 gc/L | 0% | 5,000 gc/L | 95% | 250 gc/L | 0% | 250 gc/L | 99.99% | 0.0% | 0.025 gc/L |
| Salmonella | 500 CFU/L | 99.80% | 1.0 CFU/L | 95% | 0.1 CFU/L | 0% | 0.1 CFU/L | 99.99% | 96.8% | 1.6E-07 CFU/L |
| Cryptosporidium | 17 oocyst/L | 0% | 17 oocyst/L | 95% | 0.85 oocyst/L | 0% | 0.85 oocyst/L | 99.9% | 90% | 8.5E-05 oocysts/L |
| Scenario 2 Secondary effluent, no disinfection, followed by SAT, 6 mo retention, no dilution, free chlorine disinfection | |||||
| 2° Effluent Concentration | SAT Removal | Concentration at Wellhead | Removal by Chlorination | Drinking Water Concentration | |
| Norovirus | 10,000gc/L | 99.9999% | 1.0E-02 gc/L | 99.99% | 1.0E-06 gc/L |
| Adenovirus | 5,000 gc/L | 99.9999% | 5.0E-03 gc/L | 99.99% | 5.0E-07 gc/L |
| Salmonella | 500 CFU/L | 99.9999% | 5.0E-04 CFU/L | 99.9999% | 5.0E-10 CFU/L |
| Cryptospoidium | 17 oocyst/L | 99.9999% | 1.7E-05 oocysts/L | 0.00% | 1.7E-05 oocysts/L |
| Scenario 3 Secondary effluent, MF, RO, UV/H202, groundwater injection, free chlorine disinfection | ||||||||||
| 2° Effluent Concentration | Removal through MFl | Removal through RO | Removal through UV/H2O2 | AWT Effluent Concentration | Removal through Groundwater Injection | Wellhead Concentration | Removal by Free Chlorine | Drinking Water Concentration | ||
| Norovirus | 10,000gc/L | 0% | 97% | 99.9999% | 3.0E-04 gc/L | 0% | 3.0E-04 gc/L | 99.99% | 3.0E-08 gc/L | |
| Adenovirus | 5,000 gc/L | 0% | 97% | 99.9999% | 1.5E-04 gc/L | 0% | 1.5E-04 gc/L | 99.99% | 1.5E-08 gc/L | |
| Salmonella | 500 CFU/L | 99.99% | 99.99% | 99.9999% | 5.0E-12 CFU/L | 0% | 5.0E-12 CFU/L | 99.9999% | 5.0E-18 CFU/L | |
| Cryptospoidium | 17 oocyst/L | 99.99% | 99.99% | 99.9999% | 1.7E-13 oocysts/L | 0% | 1.7E-13 oocysts/L | 0% | 1.70E-13 CFU/L | |
TABLE A-6 Dose-Response Parameters for Quantitative Microbial Risk Assessment
| Exponential k | Beta Poisson α | Beta Raisson N50 | Beta Poissonβ | Dose-Response Modelsa | |
| Norovirusb | 0.04 | 0.055 | |||
| Adenovirusc | 0.4172 | ||||
| Salmonelladd | 0.3126 | 23600 | 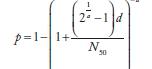 |
||
| Gryptodporidume | 0.0042 | ||||
aIn these equations, p and d are the single exposure risk and dose, respectively. As discussed previously, when the drinking water concentration is measured in genome count densities, the concentration is divided by 1000 to convert to infectious units.
bTeunis et al. (2008, Table III—pooled response for infection).
cFrom Haas et al. (1999, Table 9-15).
dFrom Haas et al. (1999, Table 9-3, Pooled Salmonella strains)
eOriginal Iowa strain data for Cryptosporidium (Haas et al., 1996).
from studies comparing the chemical composition of reclaimed and conventional waters at seven field sites in the United States (Snyder et al., 2010a; Dickenson et al. 2011), with some additional data from select pharmaceuticals adopted from Krasner et al. (2008). Other chemicals of interest, such as antimicrobials, chlorinated flame retardants, and perfluorochemicals, were adopted from field monitoring efforts using secondary treated effluents reported by Snyder et al. (2010a), Laws et al. (2011), and Drewes et al. (2003). It has been demonstrated that disinfection processes used in the treatment of wastewater and drinking water are effective in removing a significant number of hormones and pharmaceutical compounds (Snyder et al., 2007), but disinfection processes also introduce disinfectant byproducts, and for these reasons, previously cited measurements are used in the exemplar as opposed to the model-based estimates used for microbials. Table A-8 lists the concentrations of the 24 chemicals in disinfected secondary effluent, and Table A-9 shows the concentrations for undisinfected secondary effluent.
TABLE A-7 Summary of Quantitative Microbial Risk Assessment of Risk Exemplar
| Organism | Scenario 1 De Facto Reuse | Scenario 2 SAT | Scenario 3 MF/RO/UV |
| A. Pathogen Densities | |||
| Norovirus | 0.050 gc/L | 1.0E-06 gc/L | 3.0E-08 gc/L |
| Adenovirus | 0.025 gc/L | 5.0E-07 gc/L | 1.5E-08 gc/L |
| Salmonella | 1.6E-07 CFU/L | 5.0E-10 CFU/L | 5.0E-18 CFU/L |
| Cryptosporidium | 8.5E-05 oocysts/L | 1.7E-05 oocysts/L | 1.7E-13 oocysts/L |
| B. Risk of Illness (illness/(capita*d)) | |||
| Norovirus | 3.6E-05 | 7.3E-10 | 2.2E-11 |
| Adenovirus | 1.0E-05 | 2.1E-10 | 6.3E-12 |
| Salmonella | 1.7E-11 | 5.4E-14 | 0 |
| Cryptosporidium | 3.6E-07 | 7.1E-08 | 0 |
| C. Relative Risk | |||
| Norovirus | 1 | 2.0E-05 | 6.0E-07 |
| Adenovirus | 1 | 2.0E-05 | 6.0E-07 |
| Salmonella | 1 | 3.2E-03 | 0 |
| Cryptosporidium | 1 | 0.2 | 0 |
Assumptions Concerning Fate, Transport, and Removal
Scenario 1—De Facto Reuse
For the scenario describing de facto reuse (Scenario 1), it was assumed that the surface water providing dilution of treated wastewater discharge to a drinking water source represents a pristine water quality with respect to trace organic chemical concentrations, as reported by Krasner et al. (2008). The concentration of unregulated and regulated disinfection byproducts after conventional treatment (including coagulation/flocculation, filtration, free chlorine as primary disinfectant, and residual chloramines) is adopted from Krasner et al. (2008). The effectiveness of conventional water treatment for hormones, pharmaceuticals, and other trace organic chemicals was adopted from an investigation of
TABLE A-8 Estimation of Margin of Safety for Scenario 1—Drinking Water from Surface Water Source with 5% Contribution from Wastewater Discharges
| Name of Chemical | Unit | 2° Effluent with Disinfect. |
Surface Watera | Blend 95% SW 5% 2° Effluent |
Drinking Waterb | Risk Based Action Level |
Margin of Safety (unitless) |
| Nitrosaminesc,d,e | |||||||
| N-Nitrosodimethylamine (NDMA) | ng/L | 10 | <2 | <2 | <2 | 0.7 | >0.35 |
| Disinfection Byproductsf,g,h | |||||||
| Bromate | μg/L | N/A | N/A | N/A | N/A | 10 | N/A |
| Bromoform | μg/L | 18 | <0.5 | <1.1i | 3 | 80 | 27 |
| Chloroform | μg/L | 25 | <1 | <1.7i | 5 | 80 | 16 |
| Dibromoacetic acid | μg/L | 10 | <1 | <1i | <1 | 60 | >60 |
| Dibromoacetonitrile | μg/L | 16 | <1 | <1.3i | <1.3 | 70 | >54 |
| Dibromochloromethane | μg/L | <1 | <1 | <1i | <1 | 80 | >80 |
| Dichloroacetic acid | μg/L | 31 | <1 | <2i | 5 | 60 | 12 |
| Dichloroacetonitrile | μg/L | 0.3 | <1 | <1i | <1 | 20 | >20 |
| Haloacetic acid (HAA5) | μg/L | 70 | <1 | <4i | 10 | 60 | 6 |
| Trihalomethanes THMs) | μg/L | 57 | <0.5 | <3.1i | 30 | 80 | 3 |
| Pharmaceuticalsf,g,h | |||||||
| Acetaminophen | ng/L | 1 | <1 | <1i | <1 | 350,000,000 | >350,000,000 |
| Ibuprofen | ng/L | 38 | <1 | <2.4i | <2.4 | 280,000,000 | >120,000,000 |
| Carbamazepine | ng/L | 180 | 10 | 19 | 19 | 186,900,000 | 10,000,000 |
| Gemfibrozil | ng/L | 305 | 1 | 16 | 16 | 140,000,000 | 8,600,000 |
| Sulfamethoxazole | ng/L | 30 | <1 | <2i | <2 | 160,000,000 | >80,000,000 |
| Meprobamate | ng/L | 240 | 5 | 17 | 17 | 280,000,000 | 17,000,000 |
| Primidone | ng/L | 98 | 1 | 6 | 6 | 58,100,000 | 10,000,000 |
| Othersc,f,g,h | |||||||
| Caffeine | ng/L | 210 | 10 | 20 | 20 | 70,000,000 | 3,500,000 |
| 17-β Estradiol | ng/L | 0.15 | <0.1 | <0.1i | <0.1 | 3,500,000 | >35,000,000 |
| Triclosan | ng/L | 2.5 | <1 | <0.6i | <0.6 | 2,100,000 | >3,500,000 |
| TCEP (tris(2-chloroethyl)phosphate) | ng/L | 400 | <10 | <25g | <25 | 2,100,000 | >84,000 |
| PFOS | ng/L | 54 | 10 | 12 | 12 | 200 | 17 |
NOTES: N/A = data not available.
aTaken from median conc. from Krasner national occurrence survey (Krasner et al., 2008)
bRemaining after conventional surface water treatment (including coagulation/flocculation; filtration, free chlorine; residual chloramines); no transformation occurred in surface water.
cKrasner et al. (2008).
dSnyder et al. (2010a).
eDickenson et al. (2011)
fBellona et al. (2008).
gM. Wehner, OCWD, personal communication, 2009.
hBellona and Drewes (2007).
iWhen surface water concentrations were below the detection limit, one-half the detection limit was used in the dilution calculations. (In contrast, for Scenarios 2 and 3, the detection limit is used for concentrations below the detection limit to be a more conservative assumption in the relative comparison and because secondary effluent is likely to contain higher levels of contaminants than pristine surface waters.) If the final calculated concentration was below the detection limit, less than the detection limit was reported.
five conventional drinking water plants in the United States by Snyder et al. (2010a). The removal efficiencies assumed were within the same range as reported by Snyder et al. (2008a) for conventional drinking water processes.
Scenario 2—Soil Aquifer Treatment and Groundwater Recharge
For Scenario 2 describing a potable reuse system using surface spreading leading to groundwater recharge, an effluent quality is assumed that mirrors the secondary effluent qualities assumed in Scenario 1, except that Scenario 2 represents a undisinfected, filtered, secondary wastewater effluent. The water quality after 6 months of SAT, assuming no dilution with ambient groundwater, and a final disinfection with free chlorine at the wellhead, is based on findings from field monitoring efforts at SAT and riverbank filtration installations (Drewes et al., 2003; Hoppe-Jones et al., 2010; Snyder et al., 2010a; Laws et al., 2011). The data are augmented by field monitoring results for disinfection
TABLE A-9 Estimation of Margin of Safety for Scenario 2—Drinking Water from Deep-Well Supplied by Spreading of Undisinfected, Filtered, Effluent
| Name of Chemical | Unit | 2° Effluent, No Disinfection | Drinking Water Conc. | Risk-Based Action Level | Margin of Safety (unitless) |
| Nitrosaminesa,b | |||||
| N-Nra-osodimethylamine (NDMA) | ng/L | 2.7 | <2 | 0.7 | >0.35 |
| Disinfection Byproductsa,b,c | |||||
| Bromate | μg/L | N/A | N/A | 10 | N/A |
| Bromoform | μg/L | 2 | 0.5 | 80 | 160 |
| Chloroform | μg/L | 10 | 1 | 80 | 80 |
| Dibromoacetic acid | μg/L | 0.5 | <1 | 60 | >60 |
| Dibromoacetonitrile | μg/L | <1 | <0.5 | 70 | >140 |
| Dibromochloromethane | μg/L | N/A | N/A | 80 | N/A |
| Dichloroacetic acid | μg/L | 1 | <1 | 60 | >60 |
| Dichloroacetonitrile | μg/L | <1 | <1 | 20 | >20 |
| Haloacetic acid (HAA5) | μg/L | 2 | 5 | 60 | 12 |
| Trihalomethanes (THMs) | μg/L | 1 | 5 | 80 | 16 |
| Pharmaceuticalsa,b,c | |||||
| Acetaminophen | ng/L | 10 | <1 | 350,000,000 | >350,000,000 |
| Ibuprofen | ng/L | 50 | 5 | 280,000,000 | 56,000,000 |
| Carbamazepine | ng/L | 200 | 150 | 186,900,000 | 1,200,000 |
| Gemfibrozil | ng/L | 610 | 61 | 140,000,000 | 2,300,000 |
| Sulfamethoxazole | ng/L | 295 | 221 | 160,000,000 | 720,000 |
| Meprobamate | ng/L | 320 | 32 | 280,000,000 | 8,800,000 |
| Primidone | ng/L | 130 | 130 | 58,100,000 | 450,000 |
| Others | |||||
| Caffeine | ng/L | 280 | <1 | 70,000,000 | >70,000,000 |
| 17-Β Estradiol | ng/L | 1.5 | <0.1 | 3,500,000 | >35,000,000 |
| Triclosana,b,c | ng/L | 25 | 2.5 | 2,100,000 | 840,000 |
| TCEP(Tris(2-Chloroethyl)-phosphate)2,a,b,c | ng/L | 400 | 360 | 2,100,000 | 5,800 |
| PFOSa,b,c,d | ng/L | 54 | 54 | 200 | 3.7 |
| PFOAa,b,c,d | ng/L | 21 | 21 | 400 | 19 |
NOTES: N/A = data not available.
aBellona et al. (2008).
bM. Wehner, OCWD, personal communication, 2009.
cBellona and Drewes (2007).
dSnyder et al. (2010a).
byproducts after SAT reported by Krasner et al. (2008) and Dickenson et al. (2011).
Scenario 3—Reverse Osmosis, Advanced Oxidation, and Deep-Well Injection
For the potable reuse scenario via direct injection (Scenario 3), a reclaimed water quality after microfiltration, reverse osmosis, and advanced oxidation (UV/H2O2) is assumed. The concentration of disinfection byproducts in this reclaimed water after advanced treatment is adopted from monitoring at full-scale installations as reported by Wehner (2009), Bellona et al. (2008), and Bellona and Drewes (2007). Hormones, pharmaceuticals, and other trace organic chemicals in this highly treated reclaimed water are adopted from Wehner (2009), Bellona and Drewes (2007), Bellona et al. (2008), and Snyder et al. (2010a). The water quality after 6 months of retention in a potable aquifer, assuming no dilution with ambient groundwater, followed by chlorination at the point of abstraction is based on field monitoring data reported by Wehner (2009) and Snyder et al. (2010a).
The concentration levels of each of the 24 chemicals discussed above are presented in Tables A-8, A-9, and A-10 for the three scenarios for the source waters or the reclaimed water applied to the spreading or direct injection projects. Additionally, the “drinking water” column represents the final water quality delivered to customers at the end of the final treatment processes from the drinking water treatment plant (Scenario 1) or after wellhead disinfection after withdrawal from the environmental buffer (Scenarios 2 and 3). Table A-11 summarizes the concentrations of contaminants at the point of exposure for all three scenarios.
TABLE A-10 Estimation of Margin of Safety for Scenario 3—Reuse with MF/RO/UV-H2O2 and Groundwater Injection
| Name of Chemical | Unit | 2° Effluent, with Disinfect | Drinking Water Conc | Risk-Based Action Level | Margin of Safety (unitless) |
| Nitrosaminesa,b | |||||
| N-Nra-osodimethylamine (NDMA) | ng/L | <2 | <2 | 0.7 | > 0.35 |
| Disinfection Byproductsa,b,c | |||||
| Bromate | μg/L | <5 | <5 | 10 | > 2 |
| Bromoform | μg/L | <0.5 | <0.5 | 80 | > 160 |
| Chloroform | μg/L | 20 | 5 | 80 | 16 |
| Dibromoacetic acid | μg/L | <1 | <1 | 60 | >60 |
| Dibromoacetonitrile | μg/L | N/A | N/A | 70 | N/A |
| Dibromochloromethane | μg/L | <0.5 | <0.5 | 80 | >160 |
| Dichloroacetic acid | μg/L | <1 | <1 | 60 | >60 |
| Dichloroacetonitrile | μg/L | N/A | N/A | 20 | N/A |
| Haloacetic acid (HAA5) | μg/L | 13 | 5 | 60 | 12 |
| Trihalomethanes (THMs) | μg/L | 30 | 10 | 80 | 8 |
| Pharmaceuticalsa,b,c | |||||
| Acetaminophen | ng/L | <10 | <10 | 350,000,000 | >350,000,000 |
| Ibuprofen | ng/L | <1 | <1 | 280,000,000 | >280,000,000 |
| Carbamazepine | ng/L | <1 | <1 | 186,900,000 | >190,000,000 |
| Gemfibrozil | ng/L | 3 | <1 | 140,000,000 | 140,000,000 |
| Sulfamethoxazole | ng/L | 2 | <1 | 160,000,000 | 160,000,000 |
| Meprobamate | ng/L | 0.4 | <0.3 | 280,000,000 | >930,000,000 |
| Primidone | ng/L | <1 | <1 | 58,100,000 | >58,000,000 |
| Others | |||||
| Caffeine | ng/L | <3 | <3 | 70,000,000 | >223,000,000 |
| 17-β Estradiol | ng/L | <0.1 | <0.1 | 3,500,000 | >35,000,000 |
| Triclosana,b,c | ng/L | 3 | <1 | 2,100,000 | >2,100,000 |
| TCEP(Tris(2-Chloroethyl)-phosphate)a,b,c,d | ng/L | <10 | <10 | 2,100,000 | >210,000 |
| PFOSMa,b,c,d | ng/L | <1 | <1 | 200 | >200 |
| PFOAa,b,c,d | ng/L | <5 | <5 | 400 | >80 |
NOTES: N/A = data not available.
aBellona et al. (2008).
bM. Wehner, OCWD, personal communication, 2009.
cBellona and Drewes (2007).
dSnyder et al. (2010a).
Quantitative Chemical Risk Assessment
For each of the 24 chemicals identified in the three water treatment scenarios, potential lifetime health risks were assessed by calculating margins of safety (MOSs), or the risk-based action level (RBAL) divided by the concentration of contaminant in water (see Tables A-8 to A-10). RBALs represent benchmark values for risk or existing chemical-specific action levels, such as EPA maximum contaminant levels (MCLs), EPA health advisories, World Health Organization (WHO) drinking water guidelines, or chemical-specific EPA reference doses (RfDs), Agency for Toxic Substances and Disease Registry (ASTDR) minimal risk levels (MRLs), WHO acceptable daily intakes (ADIs), Food and Drug Administration (FDA) maximum recommended therapeutic doses (MRTDs), and National Library of Medicine/National Institute of Health maximum tolerated doses (MTDs) from which a drinking water action level can be derived (see also Chapter 5). Table A-12 shows the source of the values used for each of the 24 chemicals.
These risk-based values have undergone extensive regulatory and/or peer review and incorporate uncertainty factors to account for variability and uncertainty in the hazard database, and for nonpharmaceuticals, the values consider effects on sensitive subpopulations (e.g., children, pregnant women, the elderly). Conversion of an oral reference toxicity dose to a drinking water action level uses assumptions about daily drinking water intake, consumer body weight, and the relative source contribution of water to total human exposure. The
TABLE A-11 Summary of the Levels of the 24 Chemicals in the Drinking Water for Each Scenario
| Name of Chemical | Unit | Scenario 1 | Scenario 2 | Scenario 3 | |
| Nitrosamines | |||||
| N-Nitrosodimethylamine (NDMA) | ng/L | <2 | <2 | <2 | |
| Disinfection Byproducts | |||||
| Bromate | ng/L | N/A | N/A | <5 | |
| Bromoform | μg/L | 3 | 0.5 | <0.5 | |
| Chloroform | μg/L | 5 | 1 | 5 | |
| Dibromoacetic acid | μg/L | <1 | <1 | <1 | |
| Dibromoacetonitrile | μg/L | <1.3 | <0.5 | N A | |
| Dibromochloromethane | μg/L | <1 | N/A | <0.5 | |
| Dichloroacetic acid | μg/L | 5 | <1 | <1 | |
| Dichloroacetonitrile | μg/L | <1 | <1 | N/A | |
| Haloacetic acid (HAA5) | μg/L | 10 | 5 | 5 | |
| Trihalomethanes THMs) | μg/L | 30 | 5 | 10 | |
| Pharmaceuticals | |||||
| Acetominophen (paracetamol) | ng/L | <1 | <1 | <10 | |
| Ibuprofen | ng/L | <2.4 | 5 | <1 | |
| Carbamazepine | ng/L | 19 | 150 | <1 | |
| Gemfibrozil | ng/L | 16 | 61 | <1 | |
| Sulfamethoxazole | ng/L | <2 | 221 | <1 | |
| Meprobamate | ng/L | 17 | 32 | <0.3 | |
| Primidone | ng/L | 6 | 130 | <1 | |
| Others | |||||
| Caffeine | ng/L | 20 | <1 | <3 | |
| 17-β Estradiol | ng/L | <0.1 | <0.1 | <0.1 | |
| Triclosan | ng/L | <0.6 | 2.5 | <1 | |
| TCEP | ng/L | <25 | 360 | <10 | |
| (Tris(2-chloroethyl)phosphate) | |||||
| PFOS | ng/L | 12 | 54 | <1 | |
| PFOA | ng/L | 11 | 21 | <5 | |
dose metric is expressed as concentrations in drinking water. Although numerous contaminants present in the three scenarios have existing drinking water action levels (such as an EPA MCL), a significant number of chemicals have only oral RfDs, ADIs, or are pharmaceuticals with MRTDs, all expressed as milligrams per kilogram of body weight per day. Risk values such as RfDs and ADIs are generally based upon experimental doses from repeat-dose animal studies that have been adjusted with appropriate uncertainty factors to account for animal to human extrapolation and interhuman sensitivity, while MRTDs are generally derived from doses employed in human clinical trials. To derive RBALs for chemicals without existing drinking water action levels, the following formula was used:
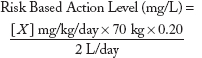
where
| X | = | Oral RfD, ADI, or other reference point such as MRTD; |
| 70 kg | = | Default adult body weight;3 |
| 0.2 | = | Default relative source contribution from drinking water of 20%; |
| 2 L/d | = | Default daily drinking water intake for a 70kg adult. |
| Y | = | Acceptable level in drinking water (i.e., estimated action level) |
_____________
3 WHO drinking water guidelines are based upon a default adult body weight of 60 kg, while a default adult body weight of 70 kg is used by EPA and was used by this NRC committee to estimate RBALs using FDA MRTDs.
TABLE A-12 Summary of Risk-Based Action Values and Sources
| Name of Chemical | Unit | Source of Risk Value | Risk Based Action Level | ||
| Nitrosamines | |||||
| NDMA | ng/L | EPA HA (EPA, 2011) | 0.7 | ||
| Disinfection Byproducts | |||||
| Bromate | μg/L | EPA MCL (EPA, 2011) | 10 | ||
| Bromoform | μg/L | EPA MCL (EPA, 2011) | 80 | ||
| Chloroform | μg/L | EPA MCL (EPA, 2011) | 80 | ||
| DBCA | μg/L | EPA MCL (EPA, 2011) | 60 | ||
| DBAN | μg/L | WHO Drinking Water Guideline Value (WHO, 2008) | 70 | ||
| DBCM | μg/L | EPA MCL (EPA, 2011) | 80 | ||
| DCAA | μg/L | EPA MCL (EPA, 2011) | 60 | ||
| DCAA | μg/L | WHO Drinking Water Guideline Value (WHO, 2008) | 20 | ||
| HAA5a | μg/L | EPA MCL (EPA, 2011) | 60 | ||
| THM | μg/L | EPA MCL (EPA, 2011) | 80 | ||
| Pharmaceuticals | |||||
| Acetominophen | ng/L | FDA MRTD (FDA, 2011) | 350,000,000 | ||
| Ibuprofen | ng/L | FDA MRTD (FDA, 2011) | 280,000,000 | ||
| Carbamazepine | ng/L | FDA MRTD (FDA, 2011) | 190,000,000 | ||
| Gemfibrozil | ng/L | FDA MRTD (FDA, 2011) | 140,000,000 | ||
| Sulfamethoxazole | ng/L | FDA MRTD (FDA, 2011) | 160,000,000 | ||
| Meprobamate | ng/L | FDA MRTD (FDA, 2011) | 280,000,000 | ||
| Primidone | ng/L | FDA MRTD (FDA, 2011) | 58,000,000 | ||
| Other | |||||
| Caffeine | ng/L | FDA MRTD (FDA, 2011) | 70,000,000 | ||
| 17-β Estradiol | ng/L | FDA MRTD (FDA, 2011) | 3,500,000 | ||
| Triclosan | ng/L | EPA RfD (EPA, 2008) | 2,100,000 | ||
| TCEP | ng/L | ASTDR MRL (ASTDR, 2009) | 2,100,000 | ||
| PFOS | ng/L | Provisional EPA HA (EPA, 2011) | 200 | ||
| PFOA | ng/L | Provisional EPA HA (EPA, 2011) | 400 | ||
aHAA5: monochloroacetic acid (MCAA) + dichloroacetic acid (DCAA) + trichloroacetic acid (TCAA) + Monobromoacetic acid (MBAA) + dibromoacetic acid (DBAA).
Ideally, the EPA bases the relative source contribution (RSC) on data regarding exposures that occur from food, air, and other important media such as personal care products or pharmaceutical agents (Donohue and Orme-Zavaleta, 2003). When data allow exposure pathways for other selected media to be quantified, default RSC values of 20, 50, or 80 percent are possible. In the absence of any data, a default RSC of 20 percent is used (Donohue and Orme-Zavaleta, 2003). EPA also assumes a daily drinking water intake of 2 L/d for an adult (EPA, 2004).
MOSs were estimated for each of the 24 contaminants (see summary of results in Table A-13). Where compounds were not detected, the lower limit on the MOS was determined using the level of detection at the concentration in drinking water.
![]()
With the exception of the chemical NDMA, the MOS values are all greater than 1, indicating that there is unlikely to be a significant health risk, even after a lifetime of exposure to these individual chemicals. The analysis does not take into account combined health effects of contaminant mixtures. Simultaneous exposure to multiple chemicals would occur in all three scenarios; thus, a consideration of mixtures would not significantly affect the relative risk comparison for purposes of the risk exemplar. NDMA was not detected in any of the scenarios, but the MOS is less than 1 because the detection limit (2 ng/L) is above EPA’s health advisory level of 0.7 ng/L. The large MOS for pharmaceuticals listed in Table A-13 indicates that potential health risks
TABLE A-13 Margin of Safety for 24 Chemicals for Each Scenario
| Chemical | Scenario 1 | Scenario 2 | Scenario 3 | ||
| Nitrosamines | |||||
| NDMA | >0.4 | >0.4 | >0.4 | ||
| Disinfection Byproducts | |||||
| Bromate | N/A | N/A | > 2 | ||
| Bromoform | 27 | 160 | >160 | ||
| Chloroform | 16 | 80 | 16 | ||
| DBCA | >60 | >60 | >60 | ||
| DBAN | >54 | >140 | N/A | ||
| DBCM | >80 | N/A | >160 | ||
| DCAA | 12 | >60 | >60 | ||
| DCAN | >20 | >20 | N/A | ||
| HAA5 | 6 | 12 | 12 | ||
| THM | 2.7 | 16 | 8 | ||
| Pharmaceuticals | |||||
| Acetaminophen | >350,000,000 | >350,000,000 | >35,000,000 | ||
| Ibuprofen | >120,000,000 | 56,000,000 | >280,000,000 | ||
| Carbamazepine | 10,000,000 | 1,200,000 | >190,000,000 | ||
| Gemfibrozil | 8,600,000 | 2,300,000 | >140,000,000 | ||
| Sulfamethoxazole | >80,000,000 | 720,000 | >160,000,000 | ||
| Meprobamate | 17,000,000 | 8,800,000 | >930,000,000 | ||
| Primidone | 10,000,000 | 450,000 | >58,000,000 | ||
| Others | |||||
| Caffeine | 3,500,000 | >70,000,000 | >23,000,000 | ||
| 17-β Estradiol | >35,000,000 | >35,000,000 | >35,000,000 | ||
| Triclosan | >3,500,000 | 840,000 | >2,100,000 | ||
| TCEP | >84,000 | 5,800 | >210,000 | ||
| PFOS | 17 | 4 | >200 | ||
| PFOA | 36 | 19 | >80 | ||
from exposure to pharmaceuticals in reclaimed water is small. However, RBALs for pharmaceuticals presented in Table A-12 assume that long-term exposure to pharmaceuticals will result in toxicity similar to short-term exposures, which is an admitted area of uncertainty. Additional research to evaluate the effects of long-term, low-level exposure to chemicals in reclaimed water could provide additional insight on whether these areas of uncertainty are biologically significant.
VERIFICATION
The committee performed several levels of verification on this risk exemplar exercise to ensure that the results are sound. In the compilation of the water quality data that provide a basis for the analysis, three committee members worked to gather and/or review the chemical occurrence data used and three additional members gathered and/or reviewed the microbial occurrence data. After the risk analysis calculations were completed and the assumptions documented by the committee members, the chair carefully reviewed the analysis. When the report was in review, Appendix A and the spreadsheet containing the calculations were reviewed in detail by a non-committee member with experience in risk assessment. With no oversight, other than to explain the task, this individual reviewed the values and formulas used in each cell of the spreadsheet and compared them to the information documented in Appendix A. Following this verification, a few minor errors were detected that were discussed with the committee chair and staff and subsequently corrected.
















