Summary of Speaker Presentations
TIME AND ENERGY SCALES IN BIOLOGY
Everything is driven by electron transfer. In biology, in our own systems, every system is rooted, built upon electron transfer.
– Leslie Dutton
Leslie Dutton, University of Pennsylvania, has spent his research career trying to understand fundamental oxidation-reduction reactions and coupled events in biological systems. He noted that 26 percent of natural enzymes—proteins that catalyze chemical reactions—are engaged in oxidation-reduction reactions. Through the capture of sunlight or chemicals, these reactions are the basis of life on Earth. Enzymes are involved in producing oxygen and food, and are also involved in breaking food down to produce energy and cellular material in humans, animals, and plants. Dutton’s work has revealed common machinery in enzymes driven by electron transfer (Moser et al., 1992). He has developed a set of engineering guidelines for how the machinery works and how synthetic proteins that might be able to carry out programmed enzyme functions can be constructed in laboratories (Page et al., 1999).
For example, Dutton has identified common machinery in natural oxidoreductases, which are involved in physiological processes such as photosynthesis. In these enzymes, electrons tunnel one at a time through the insulating protein medium that separates one redox center from another. Dutton has found that almost all (greater than 97 percent) physi-
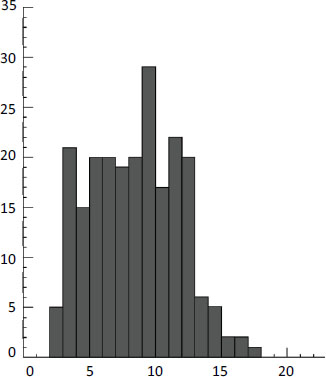
(Distance Å)
FIGURE 2-1 Distribution of physiologically productive single-electron tunneling distances (in angstroms or Å) in electron transfer and oxidoreductase proteins. More than 97 percent of the proteins have tunneling distances of less than 14 Å.
SOURCE: Adapted from Moser et al. (2010, Fig. 1A).
ologically productive electron reactions in these enzymes have tunneling distances within 4 to 14 Å (Figure 2-1; Moser et al., 2010). These distances appear to be ideal for burying and protecting redox centers so that they can maintain catalytic rates (on the millisecond timescale) found in most physiological enzymatic reactions.
Natural energy systems have evolved a general and simple engineering for electron transfer that can be used for understanding uncharacterized, naturally occurring oxidoreductases, and in constructing synthetic enzymes. Dutton talked about his ultimate goal to design artificial proteins, or what he calls “maquettes,” from scratch. He starts with a simple artificial protein scaffold, and then introduces function by progressive, iterative, and entirely reversible design steps. He tries to avoid mimicry of natural enzymes (examples shown in comparison to a maquette in
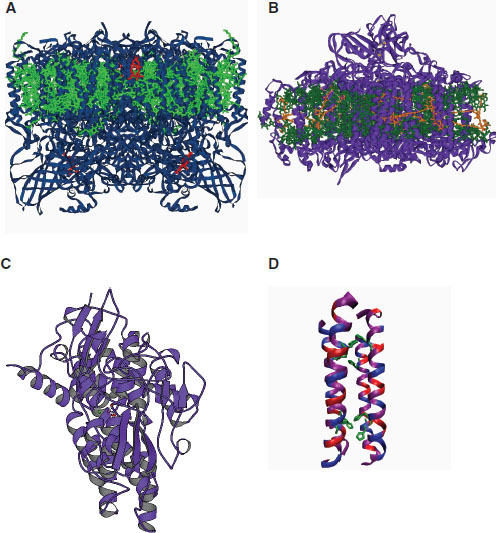
FIGURE 2-2 Comparison of natural enzymatic systems and Dutton’s synthetic “maquette”: (A) photosystem II, (B) photosystem I, (C) natural hydrogenase, (D) maquette.
SOURCE: Leslie Dutton, presentation slides.
Figure 2-2) and stays independent of natural selection. Dutton hopes to develop general-utility “omnibus” maquettes with closely related structures and diverse functions. He said these synthetic enzymes could be applied to helping meet energy needs, or in medical and clinical settings to replace systems in the human body that are not functioning properly. For example, such a maquette could provide a photocatalytic function for solar energy applications.
References
Moser, C. C., J. M. Keske, K. Warncke, R. S. Farid, and P. L. Dutton. 1992. Nature of biological electron transfer. Nature 355(6363):796-802.
Moser, C. C., J. L. Anderson, and P. L. Dutton. 2010. Guidelines for tunneling in enzymes. Biochem. Biophysics. Acta 1797(9):1573-1586.
Page, C. C., C. C. Moser, X. Chen, and P. L. Dutton. 1999. Natural engineering principles of electron tunneling in biological oxidation-reduction. Nature 402(6757):47-52.
Energy is something that biology has been doing very successfully for many billions of years.
– Penelope Boston
Penelope Boston, New Mexico Institute of Mining and Technology, conducts research on cave geomicrobiology, microbial life in highly mineralized environments, unique or characteristic biominerals, and biosignature detection (Boston et al., 2001). Although her work is not specifically focused on bioinspired energy, Boston’s work explores extreme environments, specifically subsurface earth. The biological systems she has encountered serve as inspiration for discovering new approaches to energy tranformations (Figure 2-3). Extreme environments contain biological organisms that are usually novel and unique in many ways. The chemical processes that microbes carry out in subsurface earth is of great interest to researchers seeking to relate biology to usable energy for civilization. Researchers ask questions such as “What are the natural microbial processes that occur in exotic organisms?” and “How can we use what we learn [about the microbes] in industrial applications?”
Boston’s work has recently focused on lava tubes, a byproduct of volcanic activity. Lava tubes are found throughout the Earth, and they tend to be heavily colonized by unusual bacteria, fungi, and archaea. Boston noted that the physical conditions to which surface-living organisms are commonly exposed are radically different from those of the interior (i.e., subsurface) conditions of this planet. The biggest difference between the surface level and the interior of the planet is that there is no sunlight in the subsurface, and thus no photosynthesis beyond the entrance zone or twilight zone of the cave. Some of the advantages of studying in these sites are that the temperature is virtually constant throughout the system, the microbes are low-nutrient and mineral-rich, and these sites do not have surface weather, and so they remain isolated and untouched. Part of Boston’s work focuses on subsurface biominerals, that is, minerals associated with some biological process, such as sulfur (Boston et al.,
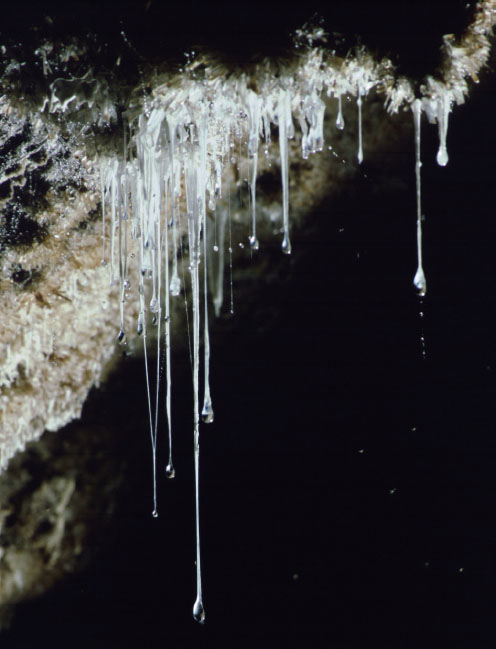

FIGURE 2-3 Slimy strings of bacteria often called “snottites” are found in caves Boston has explored. This cluster was found in Cueva de Villa Luz, Tabasco, Mexico.
SOURCE: Photo copyright Kenneth Ingham. Used by permission.
2006). The minerals found in these environments are clues to what type of metabolism sustains the microbial communities (Melim et al., 2009).
Boston believes that the subsurface rock habitats are the sites for novel metabolism shopping; they are underexploited in terms of interesting characteristics, because very few people have studied these habitats. During the presentation, Boston highlighted her work and that of others, including work in Lechuguilla Cave in New Mexico. The main message Boston made is that if one is in search of novel microbes and novel metabolism, the subsurface is a great place to look.
References
Boston, P. J., M. N. Spilde, D. E. Northup, L. A. Melim, D. S. Soroka, L. G. Kleina, K. H. Lavoie, L. D. Hose, L. M. Mallory, C. N. Dahm, L. J. Crossey, and R. T. Schelble. 2001. Cave biosignature suites: Microbes, minerals, and Mars. Astrobiology 1(1):25-55.
Boston, P. J., M. N. Spilde, D. E. Northup, L. Rosales-Lagarde, and K. Stafford. 2006. Subsurface sulfur systems: Production and preservation of distinctive biogenic signatures in sulfur, iron, manganese, and carbonate cave systems. Geochim. Cosmochim. Acta 70(18S):A60.
Melim, L. A., R. Liescheidt, D. E. Northup, M. N. Spilde, P. J. Boston, and J. M. Queen. 2009. A biosignature suite from cave pool precipitates, Cottonwood Cave, New Mexico. Astrobiology 9(9):907-917.
If you understand electron transfer, you ought to be able to build your own electron transfer stuff.
–Steve Benner
Steven Benner, Distinguished Fellow at the Foundation for Applied Molecular Evolution, is interested in chemical genetics, synthetic biology, paleogenetics, planetary biology, systems biology, and the connection of natural history to the physical sciences. Benner introduced four approaches to understanding life: prebiotic chemistry, searching the cosmos, paleogenetics, and synthetic biology. However, the focus of his presentation was his current work on synthetic biology (Benner and Sismour, 2005), which he defined as an “artificial self-sustaining chemical system capable of Darwinian evolution.” As a field, the term synthetic biology was created in 1986 by Waclaw Szybalski (1974) . As discussed by Benner, different risks are involved and need to be considered when studying synthetic biology. The goal of his laboratory is to develop a system that is self-sustaining and capable of evolving, using a six-letter artificial genetic alphabet based on natural nucleic acids, which they refer to as an artificially expanded genetic information system (AEGIS; Yang, et al., 2010).
Benner explained the relationship of the artificial bases to the naturally occurring Watson-Crick nucleic acid base pairs adenine (A) and thymine (T), and guanine (G) and cytosine (C), shown in the top frame of Figure 2-4. He explained that the two principles working in the Watson-Crick base pairing are the size complementary principle and the hydrogen-bonding complementary principle. If the Watson-Crick–based pairing is shifted, to hydrogen-donor-and-acceptor groups within the context of size complementary, eight additional bases can be created, forming four additional base pairs. The artificial base pairs have been named K and X, Z and P, V and J, and iso-C and iso-G.
Although the following results have not been published, according to Benner, he and his collaborators have successfully created synthetic DNA that replicates in artificial cells and is capable of Darwinian evolution. He explained that such synthetic biological systems are ideal and desirable because they do not carry around the billion years of “baggage” like natural systems (i.e., components that are useful for the natural organism, but not necessarily for the function of the desired application).
In conclusion, Benner continues to work toward his goal to develop a system that is self-sustaining and capable of evolving.
References
Benner, S. A., and A. M. Sismour. 2005. Synthetic biology. Nat. Rev. Genet. 6:533-543.
Szybalski, W. 1974. In vivo and in vitro initiation of transcription. In A. Kohn and A. Shatkay (Eds.), Control of Gene Expression, pp. 23-24, and Discussion pp. 404-405 (Szybalski’s concept of Synthetic Biology), 411-412, 415-417. New York: Plenum Press.
Yang, Z. Y., F. Chen, S. G. Chamberlin, and S. A. Benner. 2010. Expanded genetic alphabets in the polymerase chain reaction. Angew. Chem. Int. Ed. Engl. 49(1):177-180.
BACTERIORHODOPSIN: A MODEL PROTON ION PUMP
We should take advantage of what was learned by biology, and take this knowledge and transfer it for our own uses.
–Janos Lanyi
Janos K. Lanyi, from the University of California, Irvine, studies the physiology, biochemistry, and molecular mechanisms of ion translocation across biological membranes. One system he has studied for many years is the light-driven electrogenic transmembrane proton ion pump in the cytoplasmic membrane of Halobacterium salinarum, called bacteriorhodopsin (BR), which he spoke about in detail during the workshop.
Lanyi provided an overview of how biological systems work and discussed key principles needed to construct useful energy collection
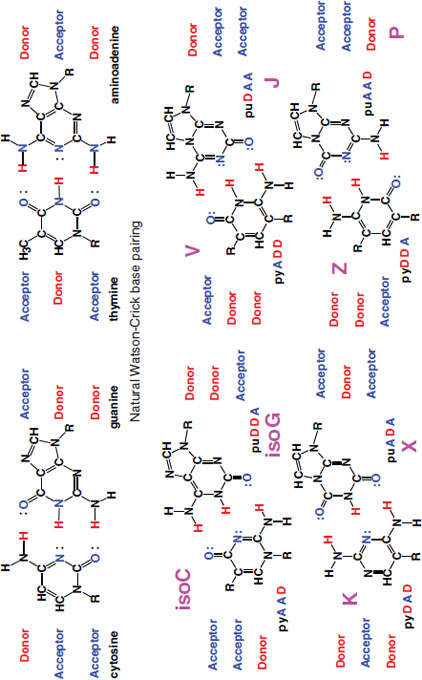
FIGURE 2-4 (Top) Natural Watson-Crick nucleic acid base pairs adenine (A), cysteine (C), guanine (G), and thymine (T); (Bottom) Benner’s artificial orthogonal DNA base pairs K, X, ZP, V, J, iso-C, and iso-G, created by shuffling donors/acceptors.
SOURCE: Stephen Benner, presentation slide.
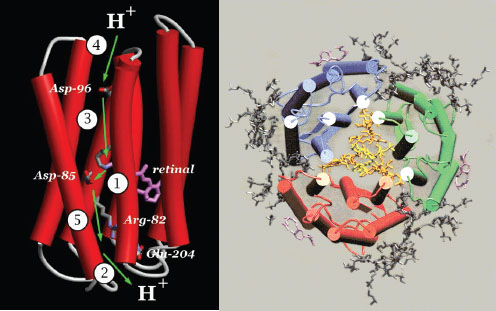
FIGURE 2-5 (Left) Ribbon diagram side view of bacteriorhodopsin showing movement of the proton at each step of the photocycle. (Right) Cartoon representation of top view of BR as it exists in nature, with three BR protein monomer units surrounded by lipid molecules. The three BR monomers are shown in blue, red, and green.
SOURCE: (Left) Adapted from Lanyi (2004). (Right) Baudry et al. (2001).
systems and/or biologically inspired technologies. He then focused on BR (Lanyi, 2004), which is a small membrane protein with a single type of subunit (homooligomer as depicted in Figure 2-5) that contains a single retinal chromophore. Lanyi and others study BR to discover new information about the mechanism of proton transport. BR is an ideal system to study, because of its simplicity compared with other kinds of proton pumps (Table 2-1).
As described by Lanyi, in a retinal-based ion pump, retinal isomerization after photon absorption initiates molecular rearrangements and results in ion uptake on one side of the membrane, translocation, and then release on the other side—the extracellular side of halophilic archaea. Then, again, another proton is taken up from the cytoplasmic side in a later reaction, which results in a net electrogenic translocation of a proton. The specific plumbing part of this pump is known, as is the proton part. Lanyi said that one of the important questions about the BR system is “How does the isomerization of the retinal chromophore in the reaction drive all the steps in this larger protein?” Lanyi said that the principle of this ion pump is that the strain in the distorted photoisomerized retinal
TABLE 2-1 Comparison of Several Types of Protein Pumps
| Protein | Cofactor, Substrate, etc. | MW | Subunits |
| Mitochondrial cytochrome oxidase | Hemes, Cu, Fe | 130,000 | 13 |
| Mitochondrial cytochrome bc1 | Hemes, FeS, ubiquinone | 225,000 | 11 |
| Mitochondrial ATPase | ATP/ADP | 350,000 | >20 |
| Mammalian NaK pump | ATP/ADP | 280,000 | 8 |
| Mammalian Ca pump | ATP/ADP | 290,000 | 2 |
| Bacteriorhodopsin | Retinal | 24,000 | 1 |
SOURCE: Janos Lanyi, presentation slide.
gradually relaxes as the binding site accommodates its changed shape. Inherently, the relaxation includes deprotonation of the Schiff base. The cascade of conformational changes that ensues allows the release of a proton at one surface and uptake at the other.
In conclusion, Lanyi said BR serves as an important and useful model for better understanding of the mechanism of photoactivity and proton transport in biological systems that could help in developing future energy systems.
References
Baudry, J., E. Tajkhorshid, F. Molnar, J. Phillips, and K. Schulten. 2001. Molecular dynamics study of bacteriorhodopsin and the purple membrane. J. Phys. Chem. 105:905-918.
Lanyi, J. K. 2004. Bacteriorhodopsin. Annu. Rev. Physiol. 66:665-688.
NOVEL MECHANISMS OF ANAEROBIC METHANE OXIDATION
If, in nature… a principle was discovered [three times], then it’s a good sign that that is the solution to the problem.
–Rudolf Thauer
The research of Rudolf K. Thauer, Max Planck Institute for Terrestrial Microbiology, focuses on the biochemistry of methanogens—microorganisms that make methane and other microorganisms that oxidize methane to CO2—primarily from a methane seep area in the Western Black Sea (Figure 2-6).
Thauer discussed anaerobic oxidation of methane with sulfate as an electron acceptor. This is of great interest because previously the anaerobic oxidation of methane was thought to be impossible in biological systems,
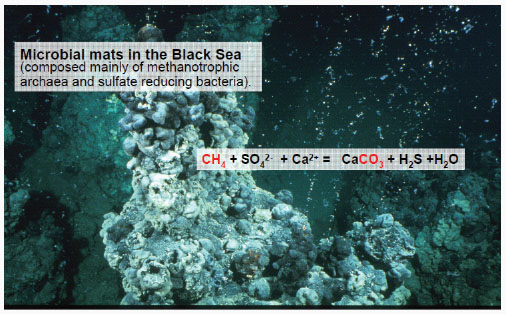

FIGURE 2-6 Microbial mats collected at cold methane seeps in the Black Sea carry out anaerobic oxidation of methane to carbon dioxide using sulfate as the electron acceptor. These mats, which predominantly consist of sulfate-reducing bacteria and archaea of the ANME-1 and ANME-2 types, contain large amounts of proteins very similar to methyl-coenzyme M reductase from methanogenic archaea.
SOURCE: Mayr et al. (2008).
because of the strength of the CH bond in methane (Thauer, 2010). To homolytically cleave the CH bond of methane, almost 440 kilojoules per mole of energy is needed. The only other bond that is on that order is the OH bond of water. One can attack methane with an OH radical, a scenario that happens in the atmosphere, but it appeared to be almost impossible to do that with a sulfur radical, or so it was thought. However, there was early evidence from Reeburgh in the 1970s that contradicts this. Now, Thauer said, there is no doubt that methane can be used by anaerobes to fuel their energy metabolism using either sulfate, MnIV, FeIII, or nitrite as the terminal electron acceptor (Figure 2-7). However, the biochemistry of this process is still largely unknown.
Thauer studies microbes from the Western Black Sea that use a nickel catalyst to oxidize methane. It has now been shown that the nickel enzyme purified from a methanogen can catalyze the oxidation of methane to methyl-coenzyme M (∆Go´ = 30 ± 10 kJ/mol) with apparent Km and Vmax consistent with the values estimated for cultures catalyzing anaerobic oxidation of methane (AOM) with sulfate (Scheller et al., 2010). The crystal structure of the MCR homologs purified from Black Sea microbial mats
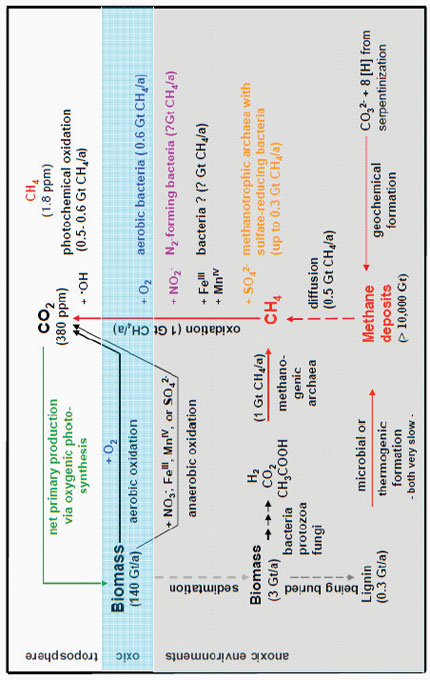
FIGURE 2-7 Schematic of the global methane cycle, which highlights the different microorganisms involved in catalyzing the anaerobic oxidation of methane with nitrite (magenta) and sulfate (orange).
SOURCE: Adapted from Thauer (2010).
catalyzing AOM with sulfate revealed that the enzyme from methanotrophic archaea uses the same coenzymes as methanogenic archaea, the best-studied system. It is therefore very likely that methane is oxidized to methyl-coenzyme M in methanotrophic archaea via the same mechanism as it is formed from methyl-coenzyme M in methanogenic archaea.
The key idea Thauer presented during his talk was that the mechanism used by microorganisms could inspire chemists to build catalysts that might be used to oxidize methane. This would be a major breakthrough because methane is the largest hydrocarbon source on Earth and storing it is problematic. However, it would also be beneficial if methane could be catalyzed to form methanol, because methanol can be easily stored and transported.
References
Mayr, S., C. Latkoczy, M. Krüger, D. Günther, S. Shima, R. K. Thauer, F. Widdel, and B. Jaun. 2008. Structure of an F430 variant from archaea associated with anaerobic oxidation of methane. J. Am. Chem. Soc. 130(32):10758-10767.
Scheller, S., M. Goenrich, R. Boecher, R. K. Thauer, and B. Jaun. 2010. The key nickel enzyme of methanogenesis catalyses the anaerobic oxidation of methane. Nature 465: 606–608.
Thauer, R. K. 2010. Functionalization of methane in anaerobic microorganisms. Angew. Chem. Int. Ed. Engl. 49:6712-6713.
SOLAR ENERGY HARVESTING IN THE
EPICUTICLE OF THE ORIENTAL HORNET
There’s a great importance today in science and industry for structures that can manipulate light. This area is extensively studied in insects and has inspired the development of many nanometric photonic structures.
–Marian Plotkin
Marian Plotkin, from the University of Singapore, discussed the possible ability of the oriental hornet to harvest solar energy (Plotkin et al., 2010). He began his presentation by describing the epicuticle of an oriental hornet (Figure 2-8), the external coating of its exoskeleton.
One of the interesting activities of the oriental hornet is its digging capability. Since the hornet builds its nest underground, it digs soil and small stones and then picks up the stones in its mouth and flies them a short distance to drop the stones from midair. It then returns to its nest to continue this process. The process is interesting because the digging activity is correlated with solar insulation, the intensity of solar radiation. The digging activity of oriental hornets is twice as much at noon as it is in the morning or in the evening. Studies conducted by Plotkin and others

FIGURE 2-8 Adult oriental hornet resting on silk caps.
SOURCE: Plotkin et al. (2010).
in the group show a statistically significant correlation between digging activity and UVB radiation.
Plotkin described the two types of cuticles, the brown and yellow cuticles (Figure 2-9), each having different pigments. The yellow cuticle contains a pigment believed to harvest solar energy for the oriental hornet. The brown cuticle is composed of the epicuticle, which is the outermost part of the cuticle, the exo-endocuticle, and the hypocuticle. Plotkin said that the size of the layers changes. The outermost layer is about 1 to 2 micrometers, while the deepest layer, the hypocuticle, is around 30 nanometers. The cuticle is brown because of a pigment called melanin, which is inside every layer. The yellow cuticle is a bit different from the brown cuticle. It also has the epicuticle, exo-endocuticle, and hypocutical layers. The layer sizes, however, are thinner than those of the brown cuticle. For example, there are about 15 layers at the exo-endocuticle, compared with 30 layers at the brown cuticle. One can see the yellow because it is contained in yellow pigment granules between the exo-endocuticle layer and the hypocuticle layer—xanthopterin.
Studies conducted by Plotkin and others in his group, found
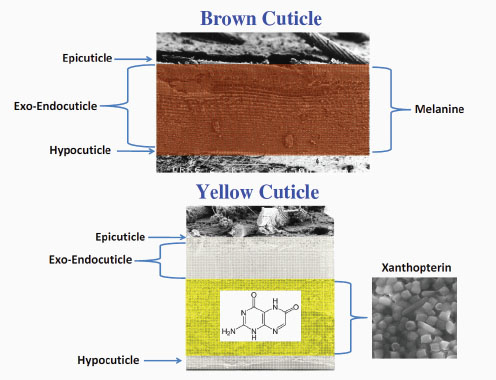
FIGURE 2-9 Cross sections of the brown and the yellow hornet cuticles.
SOURCE: Adapted from Plotkin et al. (2010).
- The oriental hornet cuticle exhibits antireflection and light-trapping properties. The measurements show that the complex layered structure increases the antireflective effectiveness. It is believed that the cuticular multilayered structure may introduce a gradual change in effective refractive index by varying the proportion of chitin nanorods within the protein matrix or by altering the angle of deposition of the nanorods.
— The xanthopterin pigment acts as a light-harvesting molecule.
— The oriental hornet correlates its digging activity with solar insulation.
— The oriental hornet cuticle is a photonic structure capable of harvesting sunlight.
- The pigment xanthopterin within the cuticle transforms sunlight into electrical energy, which may serve the metabolic activity performed within the pigment layer, hence aiding in the digging activity. They concluded this by conducting an experiment using a dye-sensitized solar cell. They inserted the pigment xanthopterin inside the electrode, and as light was absorbed by the pig-
ment and transformed to electricity, a current voltage measurement was taken. Plotkin said that this experiment indicates that xanthopterin is actually a light-harvesting molecule. It is believed by Plotkin that the electrical energy is converted into chemical energy to aid the hornet in its digging activity. However, he does not know how energy is transformed inside the oriental hornet; he believes the key to answering this question is understanding the mechanism at the molecular level.
Plotkin concluded by saying that there is a great need for structures that can manipulate light today in science and industry. This area is extensively studied in insects and has inspired the development of many nanometric photonic structures. Insects are able to construct durable, precise, and complex nanometric structures in their cuticle. Plotkin’s main message was that the oriental hornet cuticle structure and its self-assembly processes during metamorphosis could serve as a model for improving the design of photonic structures.
Reference
Plotkin, M., I. Hod, A. Zaban, S. A. Boden, D. M. Bagnall, D. Galushko, and D. J. Bergman. 2010. Solar energy harvesting in the epicuticle of the oriental hornet (Vespa orientalis). Naturwissenschaften 97(12):1067-1076.
ENERGY PRODUCTION IN BIOFUEL CELLS
We have thousands of species, maybe millions of species of microbes…. They had to have invented wonderful machines to accomplish all the things that we want to do.
–Kenneth Nealson
Ken Nealson, University of Southern California and the J. Craig Venter Institute, specializes in research on microbial physiology and genomics, environmental microbiology, metagenomics of natural populations, and microbial fuel cells. At the workshop, Nealson focused on electromicrobiology, the Shewanella organism, and some of the lessons he and his group learned from studying microbial fuel cells.
“Electrons must flow,” said Nealson. Electron flow is essential to life, and electron donors and electron acceptors define the thermodynamic range over which microbiology happens in our planet. The energy from electron flow is used to drive reactions of life. The geological environment supplies many oxidants and reductants that life has learned to utilize (Figure 2-10). For example, a methanogen does not have a lot of energy; in
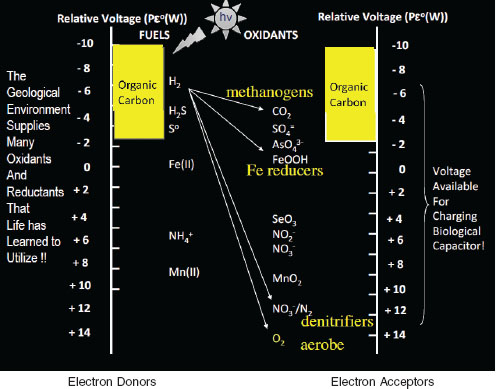
FIGURE 2-10 The relative voltage of electron donors (fuels, left) and electron acceptors (oxidants, right) show the thermodynamic foundation of microbial redox chemistry. A voltage is generated when a donor has a higher potential than an acceptor, provided all other conditions (pH, concentration of reactants, etc.) are acceptable. Life has evolved to use almost every one of the sets of redox couples. Eukaryotic life in general uses organic carbon as electron donor and oxygen as electron acceptor, whereas the prokaryotes (methanogens, FE reducers, and denitrifers) use many other acceptors, taking advantage of abundant and ubiquitous energy sources and oxidants available.
SOURCE: Ken Nealson, presentation slides.
fact it will consume energy to reproduce. However, iron reducers generate more energy than methanogens, but much less than aerobes, because oxygen is a favorite electron acceptor. Thus, the geology of Earth and the mineralogy of Earth and microbiology are strongly connected.
Nealson then spoke about a unique organism that he isolated about 24 years ago in the sediments of Lake Oneida, New York, which he named Shewanella oneidensis (Venkateswaran et al., 1999). Nealson said Shewanella “is a remarkable ‘bug’ that is capable of doing really remarkable environmental chemistry.” He said his laboratory found strange manganese and iron chemistry in a lake in New York that could not be explained by just
chemistry, so they thought it must be due to a microbe capable of reducing iron and manganese solids, which led to the discovery of Shewanella oneidensis.
Nealson showed audience members a short video clip that was taken inside an anaerobic flat capillary. In the video, Shewanella was found to be swimming around and reducing tiny pieces of manganese oxide until all of this terminal electron acceptor was depleted. Under normal conditions, the manganese oxide is stable for about 10,000 years. However, working at low temperature, this biological catalyst was able to transform the very stable mineral about a million-fold faster than under normal weathering conditions.
Nealson described many important facts about Shewanella that his laboratory and others found through experiments over the years. The important facts include
- Genes for metal reduction are multiheme cytochromes.
- Cytochromes are on outer membranes (C. Myers, Medical College of Wisconsin).
- There are ~10,000 of each cytochrome per cell (Lower et al., 2007).
- Complex interactions exist between proteins (Ming Tien, Pennsylvania State University).
- Proteins are synthesized under anaerobic conditions (Daâd Saffarini, University of Wisconsin, Milwaukee).
- Some strains of Shewanella have duplicate genes (induced in biofilms, J. McLean, J. Craig Venter Institute).
- They are capable of electron transport to the anode of microbial fuel cells (Byung Hong Kim, Korean Institute of Science and Technology, and others).
- Under electron acceptor limitation, cells make electrically conductive nanowires (Y. Gorby, University of Southern California; Gorby et al., 2006; El-Naggar et al., 2010).
- Extracellular polymeric substance (EPS) with conductive metals (pyrite) also works.
Nealson and others sequenced about 20 different strains of Shewanella, and because they had the sequence, they made deletion mutants that were nonpolar in particular spots. They found by deleting virtually all the cytochromes in this microbe one at a time that there are three enzymes that are the main working parts for metal reduction. They are MtrA, MtrB, and MtrC (Mtr = metal reduction), as shown in the conceptual model for extracellular electron transfer in Shewanella sp. in Figure 2-11. They found that if the genes that produce these three enzymes are put into Escherichia coli, the bacteria will reduce iron and manganese. A key point Nealson
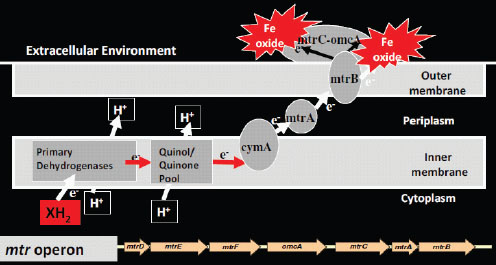
FIGURE 2-11 Conceptual model for extracellular electron transfer in Shewanella sp.
SOURCE: Ken Nealson, presentation slide.
made is that MtrA, B, and C are all decaheme cytochromes. On the outside of the cell is an MtrC complex. Spanning the pariplasm is MtrA, which is also a multiheme cytochrome.
Nealson then spoke about the modes of extracellular transfer, highlighting three types:
- Direct transfer using multiheme cytochromes located on the cell exterior,
- Biogenic environmental soluble mediators, and
- Bacterial nanowires (or conductive EPS).
Lastly, Nealson spoke about how his laboratory has worked on developing Shewanella-based microbial fuel cells, which produce electricity from the metabolism of the microbe (e.g., He et al., 2009). Initially, his group developed fuel cells to take advantage of the electron transfer properties of the bacteria in an anaerobic environment coupled to an abiotic electron acceptor on the other side of the cell. More recently, Nealson and coworkers found the Shewanella metabolism could be engineered to be both an electron donor in an anerobic environment at the anode and an electron acceptor in an aerobic environment at the cathode of the fuel cell, what he described as “stretching out the microbes’ metabolism and putting it to work for humans.” He showed preliminary images of the fuel cell being used to purify polluted water, which indicate the possibility of
using these microbial fuel cells for a low-cost (low-energy) solution for wastewater treatment, especially in the developing world.
In summary, Nealson highlighted the importance of exploring unique biological systems such as Shewanella, which he believes have a lot to teach humans about harvesting energy from unconventional sources.
References
El-Naggar, M. Y., G. Wanger, K.-M. Leung, T. D. Yuzvinsky, G. Southam, J. Yang, W.-M. Lau, K. H. Nealson, and Y. A. Gorby. 2010. Electrical transport along bacterial nanowires from Shewanella oneidensis MR-1. Proc. Natl. Acad. Sci. U.S.A. 107(42):18127-18131.
Gorby, Y. A., S. Yanina, J. S. McLean, K. M. Rosso, D. Moyles, A. Dohnalkova, T. J. Beveridge, I.-S. Chang, B.-H. Kim, K.-S. Kim, D. E. Culley, S. B. Reed, M. F. Romine, D. A. Saffarini, E. A. Hill, L. Shi, D. A. Elias, D. W. Kennedy, G. Pinchuk, K. Watanabe, S. Ishii, B. Logan, K. H. Nealson, and J. K. Fredrickson. 2006. Electrically conductive bacterial nanowires produced by Shewanella oneidensis strain MR-1 and other microorganisms. Proc. Natl Acad. Sci. U.S.A. 103:11358-11363.
He, Z., J. Kan, Y. Wang, Y. Huang, F. Mansfeld, and K. H. Nealson. 2009. Electricity production coupled to ammonium in a microbial fuel cell. Environ. Sci. Technol. 43:3391-3397.
Lower, B. H., L. Shi, R. Yongsunthon, T. C. Droubay, D. E. McCready, and S. K. Lower. 2007. Specific bonds between an iron oxide surface and outer membrane cytochromes MtrC and OmcA from Shewanella oneidensis MR-1. J. Bacteriol. 189(13):4944-4952.
Venkateswaran, K., D. P. Moser, M. E. Dollhopf, D. P. Lies, D. A. Saffarini, B. J. MacGregor, D. B. Ringelberg, D. C. White, M. Nishijima, H. Sano, J. Burghardt, E. Stackebrandt, and K. H. Nealson. 1999. Polyphasic taxonomy of the genus Shewanella and description of Shewanella oneidensis sp. nov. Int. J. Syst. Bacteriol. 49(2):705-724.
MICROBES AND THE FOUR BASIC STRATEGIES FOR LIFE ON EARTH
There are these specific solutions [in nature] and yet an inherent flexibility to be able to adapt as those things change.
–Felisa Wolfe-Simon
Felisa Wolfe-Simon, a NASA Astrobiology Research Fellow, highlighted the inherent flexibility of biology. She said life on Earth is metabolically diverse and yet maintains a biochemical unity, and the basis for all life starts with chemical underpinnings. This chemical potential manifests in four metabolic strategies used by life on Earth today, all of which most likely evolved in the distant past; they are photoautotrophic, photoheterotrophic, chemoautotrophic, and chemoheterotrophic. However, a single metabolism is not linked to a unique microbe. Wolfe-Simon said biological systems, which have a 3.5-billion-year history on Earth, are useful models to learn something about energy, because they have had a long time to evolve and learn unique and varied energy solutions. While
there are some very specific energy solutions that have evolved in biological systems, there is also an inherent flexibility in those solutions that has allowed the systems to adapt overtime to a changing environment.
Wolfe-Simon discussed two of the projects she is working on. One of the projects is exploring photosynthesis in cyanobacteria that do not make oxygen (anoxygenic), and how this system can help understand the flexibility of “light-driven energy capture” (Johnston et al., 2009; Chauhan et al., 2011). Cyanobacteria are capable of facultative anoxygenic photosynthesis using a variety of electron donors in addition to water.
The other project Wolfe-Simon spoke about is the recently published research on a microbe, GFAJ-1 (found in high-arsenic soda lakes in California and Nevada, Figure 2-12), that appears to sustain its growth using arsenic instead of phosphorus, for example, in DNA (Wolfe-Simon et al., 2011). This result is important because all life is known to require six critical elements—carbon, hydrogen, nitrogen, oxygen, phosphorus, and sulfur—and these elements are used to make important molecules such as DNA, RNA, proteins, and lipids. However, GFAJ-1 appears to be able to use arsenic when phosphorus is not available. Wolfe-Simon and her coworkers found that arsenic is present in the microbe wherever phosphorus is.
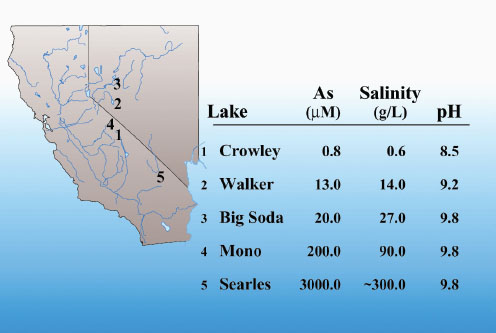
FIGURE 2-12 High-arsenic soda lakes found in the western United States, where arsenic-tolerant cyanobaceria GFAJ-1 exist.
SOURCE: Felisa Wolfe-Simon, presentation slide.
At the time of the workshop, the publication of this work had been met with skepticism and criticism in the scientific research community, which Wolfe-Simon discussed. Since then, much of debate has been captured on the editorial pages of Science (Pennisi, 2011).
Wolfe-Simon concluded by saying that the next steps in learning more about the role of arsenic in the GFAJ-1 bacteria would include synchrotron studies, in-depth physiological characterization, and genomic sequencing. In addition, cultures of GFAJ-1 have been made available to other groups to study independently.
References
Chauhan, D., I. M. Folea, C. C. Jolley, R. Kouril, C. E. Lubner, S. Lin, D. Kolber, F. Wolfe-Simon, J. H. Golbeck, E. J. Boekema, and P. Fromme. 2011. A novel photosynthetic strategy for adaptation to low iron aquatic environments. Biochemistry 50(5):686-692.
Johnston, D. T., F. Wolfe-Simon, A. Pearson, and A. H. Knoll. 2009. Anoxygenic photosynthesis modulated Proterozoic oxygen and sustained Earth’s middle age. Proc. Natl. Acad. Sci. U.S.A. 106(40):16925-16929.
Pennisi, E. 2001. Concerns about arsenic-laden bacterium aired. Science 332(6034):1136-1137.
Wolfe-Simon, F., J. S. Blum, T. R. Kulp, G. W. Gordon, S. E. Hoeft, J. Pett-Ridge, J. F. Stolz, S. M. Webb, P. K. Weber, P. C. W. Davies, A. D. Anbar, and R. S. Oremland. 2011. A bacterium that can grow by using arsenic instead of phosphorus. Science 332(6034):1163-1166.
Many biological systems actually work isothermally; that means you are not heating things up to vast temperatures and using a lot of energy there.
–Nadrian C. Seeman
Nadrian C. Seeman, from New York University, discussed his work on using DNA’s chemical information for bottom-up nanoscale control of the structure of matter and taking natural material and making an unnatural object from it. More biomimetic than bioinspired, Seeman described how the chemical information contained in DNA can be used to direct methods of organizing matter and making nanoscale mechanical devices such as assembly lines.
Seeman opened his remarks by reviewing what DNA is—the genetic material of people, plants, and animals. Seeman focused the rest of his presentation on highlighting DNA, the molecule, and what can be done with DNA outside of its biological context.
DNA, in its natural form, is linear topologically because it is unbranched. In its natural state, Seeman said it is not useful for construction of nanoscale mechanical devices, but it does contain genetic information making it an excellent tool for biomimetic synthesis. To produce

FIGURE 2-13 Reciprocal exhange of DNA, an idea Seeman “stole” from nature to create new DNA motifs. The blue and red lines represent two different sequences of DNA. After reciprocal exchange, the blue strand and red strand bedcome red and blue strands.
SOURCE: Nadrian Seeman, presentation slides.
structures more diverse than the natural DNA double helix, Seeman says he stole the idea of DNA and reciprocal exchange from nature (Seeman, 2001). He added “some people use the term bioinspired; I use the term biokleptic.” Figure 2-13 shows a simplistic illustration of reciprocal exchange, where two different DNA sequences combine to produce new structures.
Branched species can then be connected to one another using the same interactions that genetic engineers use to produce constructs, that is, cohesion by molecules tailed in complementary single-stranded overhangs known as “sticky ends.” Such sticky-ended cohesion is used to produce N-connected objects, crystalline lattices, and complex nanomechanical devices such as assembly lines.
The objectives and applications in Seeman’s laboratory include architectural control and scaffolding, producing nanomechanical devices such as nanorobotics and nanofabrication, and self-replicable systems (Gu et al., 2010). Seeman said his lab focuses on DNA because “nucleic acid sequences can be programmed and synthesized, leading to information-based structural, dynamic, and catalytic chemistry.”
As illustrated in Figure 2-14, the intellectual goal of structural DNA nanotechnology is to control the structure of matter in three dimensions (3D) to the highest extent possible and to understand the connection between the molecular and macroscopic scales (Zheng et al., 2009). Seeman said that the ability to control matter in this way has the potential to open new functionality of relevance to energy and other applications. For example, as discussed during the workshop, the 3D lattice structures Seeman created could have applications for separations or storage of small proteins or other molecules.
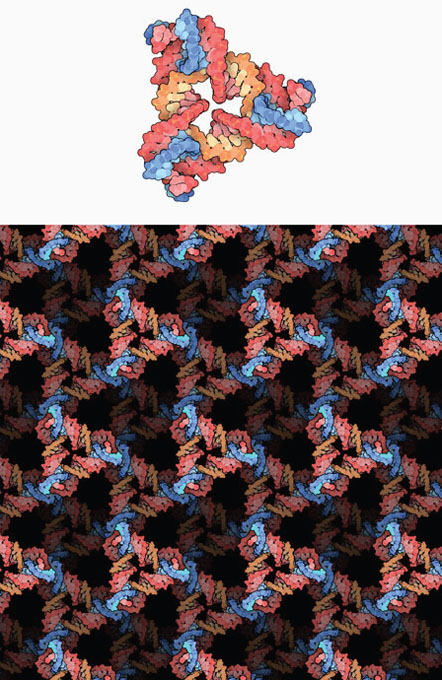
FIGURE 2-14 Illustrations of DNA structures. (Top) Branched DNA unit, with blue representing the connecting sticky ends. (Bottom) Three-dimensional array of engineered DNA units created by Seeman and coworkers.
SOURCE: Illustrations by David Goodsell, based on the RCSB Protein Data Bank entry 3gbi. Used by permission.
References
Gu, H., J. Chao, S. J. Xiao, and N. C. Seeman. 2010. A proximity-based programmable DNA nanoscale assembly line. Nature 465(7295):202-205.
Seeman, N. C. 2001. DNA nicks and nodes and nanotechnology. NanoLetters 1(1):22-26.
Zheng, J., J. J. Birktoft, Y. Chen, T. Wang, R. Sha, P. E. Constantinou, S. L. Ginell, C. Mao, and N. C. Seeman. 2009. From molecular to macroscopic via the rational design of the self-assembled 3D DNA crystal. Nature 461(7260):74-77.
This page intentionally left blank.


























