John Leslie King
-University of Michigan-
As the chair of the committee that is organizing this two-day event on “The Future of Scientific Knowledge Discovery in Open Networked Environments,” I would like to thank primarily the National Science Foundation, but also the Library of Congress and the Department of Energy, for sponsoring this project. The event was organized by two boards of the National Academies, the Board on Research Data and Information in collaboration with the Computer Sciences Telecommunications Board. In my opening remarks, I would like to address two points: one on the motivation for this event and the other one on the means.
The motivation lies in the fact that we are undergoing a relatively rapid change in capability, because of technology and improved knowledge. What is possible today was not even thinkable some years ago for data generation, storage, management, and sharing. Developments in these areas are having a rather profound impact on the nature of science.
This is not new historically, however. When the technologies of optics were developed and the microscope and telescope were created, these technologies changed what we could do, but they did not immediately change what we did do. Over time, there was an evolution of social conventions, of data record keeping, and so forth. Take for example the classical story of Tycho Brahe, who spent years looking through telescopes and taking detailed measurements, and later his data were instrumental in Johannes Kepler’s proof that the orbits of the plants were elliptical rather than circular.
That kind of practice has been repeated many times in the history of science, and it is being repeated now. Some of my colleagues at the University of Michigan are working on the ATLAS Project at the Large Hadron Collider in CERN, the European Center for Nuclear Research. ATLAS is an interesting operation. It has nearly 3,000 principal investigators, and many of the papers produced through this project have more pages dedicated to the listing of authors than to the paper itself. Obviously they have had to develop new conventions to deal with this situation. If we ask them about how they know who did what in a project or a paper, they will say that they just know. That is probably more tacit knowledge than it is explicit knowledge.
The ATLAS detector is capable of generating a petabyte of data per second; most of these data are thrown away immediately as not relevant for the work. The amount of data left is still huge, however, and CERN distributes large volumes of data globally through different networks. Such big and important operations have to be planned. We cannot just expect that information technology people will take care of that simply because nobody has ever done it before. It took years to plan the data distribution plan for CERN. This, of course, did not involve the details of how the data get used, how credit gets assigned, and so forth. The point I am trying to make is that this is an emerging and very challenging frontier, and that we are still
at the infant stage of it. The means are to address four issues: (1) the opportunity–benefit space; (2) the techniques that we know now and those that we think are going to emerge; (3) the barriers we have to overcome; and (4) the options that we can use to move forward.
Opening Remarks by Project Sponsors
Alan Blatecky
-National Science Foundation-
Thinking about the cyberinfrastructure and the impact of data, we need to understand that it is an ecosystem with multiple components that are dependent upon each other. Figure 1-1 presents a very complex and interconnected world of cyberinfrastructure. Although data is a key component, there are other components that impact data dramatically; this includes networking, software, computational resources, expertise, and so on.
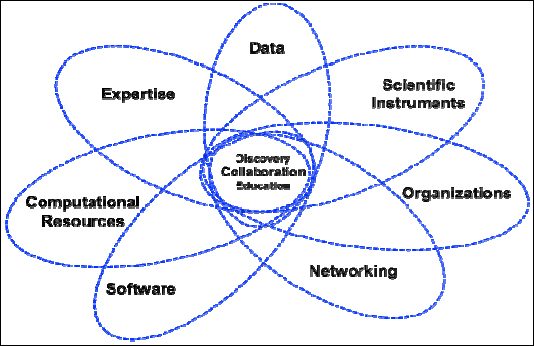
FIGURE 1-1 Cyberinfrastructure.
SOURCE: Alan Blatecky
Furthermore, Figure 1-1 makes two points. First, data cannot be considered as a separate entity. Second, as data capabilities grow, this has an impact on networking requirements, on organizations, on expertise, and so forth—and also vice versa. Advances in one component affect other areas. As mentioned with the ATLAS Project example in the opening presentation, as advanced scientific instruments are developed, they enable tremendous new capabilities. The challenge is how to accommodate and take advantage of these new capabilities, and how does this impact the cyberinfrastructure environment?
Figure 1-2 addresses four data challenges.
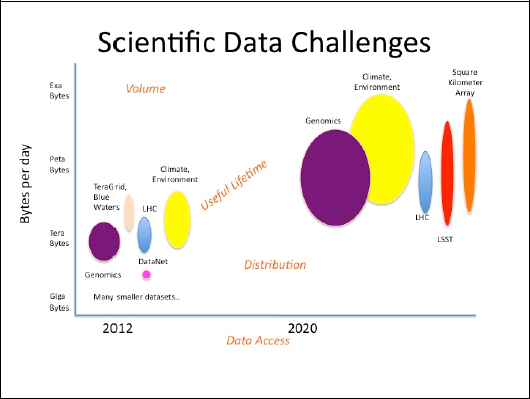
FIGURE 1-2 Scientific data challenges.
SOURCE: Alan Blatecky
The first point is the growth and useful lifetime in data from today to what is expected in 2020. While terabytes and petabytes of data and information are currently being generated, in less than 10 years we are going to have volumes in petabytes and exabytes. In short, there is going to be a tremendous growth in the amount of scientific data being generated, and there can be a long lifetime associated with the data. Therefore, issues of curation, sustainability, preservation, and metadata approaches need to be addressed.
The second challenge is the volume of data by discipline, instrument, or observatory. In Figure 1-2, the vertical dimension of the ellipses is intended to illustrate the volume of data being generated by that area. As Prof. King pointed out, the Large Hadron Collider (LHC) will be generating a gigabyte per second—multiple terabytes a day, in essence—and the figure illustrates how much volume that is.
The third challenge is distribution. In Figure 1-2, the horizontal dimension of the ellipse illustrates the amount of distribution for that data. For example, the data from the LHC or Large Synoptic Survey Telescope will be stored in a few locations; hence, the ellipse is fairly narrow. Genomics data or climate data, on the other hand, will be located at literally hundreds, if not thousands, of locations. The point here is that distribution is a very different challenge from volume or lifetime. Genomic data are being generated at incredible rates at many universities and medical centers, and managing that data across hundreds of sites is a significant challenge.
The fourth challenge deals with data access. How will the data be accessed and what policies are required for access (e.g., privacy, security, integrity)? These are some of the challenges we need to address.
The point of all of this is exactly what Prof. King suggested in his opening presentation—that science, technology, and society will be transformed by data. Modern science will depend on it. We need to be focusing on multidisciplinary activities and collaborations. In this new Age of Observation, there is a whole new world opening up for everyone; this is similar to what happened with the development of telescopes, only now data will be the “instrument” that will change the way we understand the world.
The last point I will make is that the National Science Foundation (NSF) Advisory Committee on Cyberinfrastructure established six task forces that worked for almost 2 years to develop a series of recommendations on cyberinfrastructure. The data task force addressed these issues in depth, and as the task force concluded, there are infrastructure components to deal with, as well as culture changes and economic sustainability. It is necessary to determine how to manage data and deal with such diverse issues as ethics, intellectual property rights, and economic sustainability. In this regard, the NSF should address the following issues:
![]() Infrastructure: Recognize data infrastructure and services (including visualization) as essential research assets fundamental to today’s science and as long-term investments in national prosperity.
Infrastructure: Recognize data infrastructure and services (including visualization) as essential research assets fundamental to today’s science and as long-term investments in national prosperity.
![]() Culture Change: Reinforce expectations for data sharing; support the establishment of new citation models in which data and software tool providers and developers are credited with their contributions.
Culture Change: Reinforce expectations for data sharing; support the establishment of new citation models in which data and software tool providers and developers are credited with their contributions.
![]() Economic Sustainability: Develop and publish realistic cost models to underpin institutional and national business plans for research repositories and data services.
Economic Sustainability: Develop and publish realistic cost models to underpin institutional and national business plans for research repositories and data services.
![]() Data Management Guidelines: Identify and share best practices for the critical areas of data management.
Data Management Guidelines: Identify and share best practices for the critical areas of data management.
![]() Ethics and Internet Protocol: Train researchers in privacy-preserving data access.
Ethics and Internet Protocol: Train researchers in privacy-preserving data access.
Sylvia Spengler
National Science Foundation
I am pleased to have this meeting organized, because of the opportunity for the Board on Research Data and Information and the Computer Sciences Telecommunications Board to come together and begin to explore where the policy, legal, social, and computer science challenges intersect in this area. I am also pleased that we were able to bring together individuals who represent different communities and fields. This is a good opportunity to sustain conversations that have been scattered around the globe for the past 15 years.
From a personal perspective, I am gratified to see that some of the issues that I have long worried about in the fields of data and analytics have finally come to the attention of the people who are going to use the data. We have at the table not only scientists from a single scientific domain, but scientists from many different kinds of domains. Science is inherently multidisciplinary and will require different kinds of expertise, different communities, and new kinds of social environments within science. This is an opportunity to begin to think about that.
Keynote Address: An Overview of the State of the Art
Tony Hey
-Microsoft Research-
I will provide a summary of where I think we are in the data-intensive knowledge discovery area and offer a personal view of some of the issues that we are facing. As mentioned earlier, we are in the midst of a revolution, but we have been in this revolution before. I like the example of Kepler, because Tycho Brahe’s measurements were sufficiently accurate that Kepler could not fit the planetary orbits to circles and was forced to consider ellipses, which led eventually to Newton’s Laws. However, it was actually the data that Tycho Brahe took without a telescope that led Kepler to his conclusion.
Some of my slides are based on Dr. Jim Gray’s last talk in 2007. Dr. Gray was a computer scientist at Microsoft until he disappeared at sea in January 2007 while on his sailboat. Dr. Gray made the argument that we are moving into a new area in which people will need different skills.
Obviously the skills for experimental and theoretical science are not going to disappear—we still need and use them—but computational science requires people to know about, for example, numerical methods, algorithms, computer architectures, and parallel programming. This is a set of specialized skills for doing work in areas such as climate prediction and modeling galaxy formation. Today data-intensive science essentially requires a new set of skills for handling and manipulating large amounts of data, visualizing them, and so on. When I ran the U.K.’s e-Science Program, the agenda was about tools and infrastructure to support data-intensive science. It was distributed, it was data intensive, and it involved a new way of thinking about what we are doing. That is how I distinguish between e-science and data-intensive science.
Figure 1-3 is from Jim Gray’s talk.
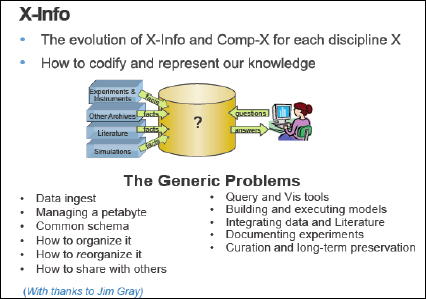
FIGURE 1-3 X-Info.
SOURCE: Microsoft Research
Figure 1-3 shows field X evolving into two separate subdisciplines. There is computational biology, and there is bioinformatics; computational chemistry, and then chemoinformatics. Almost every field (e.g., astroinformatics, archeoinformatics) today is being pervaded by informatics and the need to deal with data. It is no longer just science.
Dr. Gray saw that in responding to the problems of getting and manipulating the data there is a need to teach people new skills. Managing a petabyte of data is not easy, because, for example, it is difficult to send a petabyte easily over the Internet. We need to think about organizing and reorganizing the data in case a wrong assumption was made. Other issues to think about include sharing the data, query and visualization, integrating data and literature, and so on. As we can see, there are many new challenges.
Dr. Gray had this vision of moving from literature to computation and then back to literature through a process of linking and combining different sources of data to produce new data and knowledge. He believed that this was the future global digital library for science and that it could lead to a huge increase in scientific productivity, because the research process is fairly inefficient at present, as described in Figure 1-4.
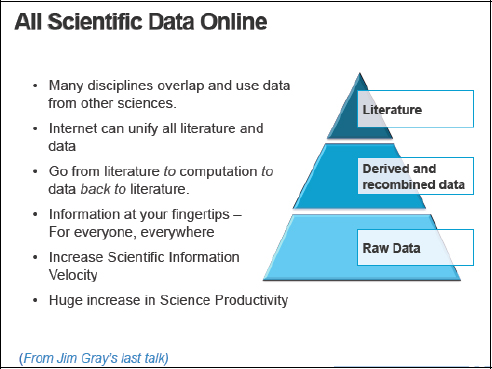
FIGURE 1-4 All scientific data online.
SOURCE: Microsoft Research
The other issue that I would like to emphasize is the reproducibility of science. If a scientist is interested in a paper, accessing the data on which that paper is based is not easy, and it is probably only in the minority of cases that a scientist can do it. So this was Dr. Gray’s vision—a big, global-distributed digital library—and that was why I accepted to cochair a National Science Foundation (NSF) Office of Cyberinfrastructure Taskforce on Data and Visualization. We tried to produce a report quickly, because the data agenda is a very important area for all the agencies, not just NSF2.
__________________________
2 The final version of the report can be found on the NSF Web site http://www.nsf.gov/od/oci/taskforces/.
The report covers areas such as infrastructure, delivery, culture and sociological change, roles and responsibilities, ethics and privacy, economic value and sustainability, and data management guidelines. Also, NSF is now requiring that its grantees have a data management plan.
Another point I would like to make is related to funding. To illustrate this point, I will give an example from the Coastal Ocean Observation Lab at Rutgers University. After a boating or aircraft accident at sea, the U.S. Coast Guard (USCG) historically has relied on current charts and wind gauges to determine where to hunt for survivors. But thanks to data originally collected by Rutgers University oceanographers3 to answer scientific questions about earth-ocean-atmosphere interactions, the USCG has a new resource that promises to literally save lives. This is a powerful example that large datasets can drive myriad new and unexpected opportunities, and it is an argument for funding and building robust systems to manage and store the data.
However, one of the group’s frustrations today, unfortunately, is the lack of funding to design and support long-term preservation of data. A large fraction of the data the Rutgers team collects has to be thrown out, because there is no room to store it and no support within existing research projects to better curate and manage the data. “I can get funding to put equipment into the ocean, but not to analyze that data on the back end,” says Schofield.
Given the political climate that we are in, there is a need to convince the taxpayers who fund this kind research that it is worthwhile. Thus, we need engagement with the citizens.
Let me provide another example. The Galaxy Zoo uses the Sloan Digital Sky Survey data, which contains around 300 million galaxies. Since there are only about 10,000 professional astronomers, several Oxford University astronomers enlisted public support to help classify them in their Galaxy Zoo project. They have had more than 200,000 individuals participating who have helped classify galaxies and have found various interesting discoveries, such as a blue galaxy called Hanny’s Voorwerp. Hanny is a Dutch school teacher. Another example is a new classification of galaxies called “green pea galaxies.” Hence, engaging the public is a good strategy.
Then there is the issue of return on investment. Why do scientists want to save the data? To address this issue, I will use this interesting collaboration called the Supernova Factory. Because data curation and management were considered a priority in this project, today the Supernova Factory is a shining example of a significant return on investment, both in financial resources and in scientific productivity. It reduced labor, it reduced false supernova identification by 40 percent, it improved scanning and vetting times by 70 percent, and it reduced labor for search and scanning from six to eight people for 4 hours to one person for 1 hour. Not only did the system pay for itself operationally within one-and-a-half years, but it enabled new science discovery. The important factor in this case is that data curation and management were considered a priority from the very beginning, so the developers built a system that embodied that.
__________________________
3 At Rutgers University’s Coastal Ocean Observation Lab, scientists have been collecting high-frequency radar data that can remotely measure ocean surface waves and currents. The data are generated from antennae located along the eastern seaboard from Massachusetts to Chesapeake Bay.
In his talk, Dr. Gray also listed some calls for action. One of them is related to funding and support of software tools. NSF and other funding agencies are working on developments in support of software tools, but this is always going to be a problem. We can buy some tools off the shelf, but there are always going to be gaps that the commercial software industry does not cover.
I personally came to the conclusion that building complete cyberinfrastructure using open-source software is very difficult. If scientists could buy different pieces of the infrastructure from various vendors and use any relevant open-source software available for them, then they could focus their funding and software development efforts into building the pieces that are not available in the commercial market.
Dr. Gray pointed out that while the biggest projects have a software budget, the small projects generally do not. He felt there was a range of actions that could be done, and that is what he meant by funding generic laboratory information management systems—support going from data capture to publication. His advice was to “let a thousand flowers bloom”—scientific data management research, data analysis, data visualization, and new algorithms and tools.
Figure 1-5 illustrates the data life cycle. It begins with data acquisition and modeling. Because of collaboration, the data are usually distributed. Then the question is how to get the data into a usable form. This would include data analysis, data mining, and visualization. Next is the stage of writing up the data and sharing them in various ways. The final stage in the life cycle is data archiving and preservation, which, for example, can be important for environmental studies where scientists cannot repeat the same measurement. There is no hope of preserving all of the data, though, because this may be too expensive.

FIGURE 1-5 Data life cycle.
SOURCE: Microsoft Research
I will now talk about a tool called ChronoZoom that we have been collaborating on with Prof. Walter Alvarez at the University of California, Berkeley. What Prof. Alvarez is trying to do is to develop a new “Big History” perspective that begins with the origins of the universe, but makes it possible to look at history on various scales. It will help scientists to put not only biological evolution but also geological evolution and historical events in the system,
and then compare them and see what influenced what in these areas, by looking at it from these different scales. The zoom metaphor is what appealed to Prof. Alvarez, so we built him a tool that enables detailed examination and exploration of huge differences in time scales. This example illustrates that working with such data is not just for the sciences but also for the arts and the humanities.
When I arrived at Microsoft in 2005, Bill Gates was giving a talk at the Supercomputing Conference. Figure 1-6 is one of the slides that he used, which illustrates a new era of research reporting.

FIGURE 1-6 Envisioning a new era of research reporting.
SOURCE: Microsoft Research
Mr. Gates envisioned that the documents would be dynamic, so that someone could get the data, interact with the data and images, change the parameters, and run it again. Furthermore, besides it being peer reviewed, there could also be a tool like an Amazon rating, or it could be similar to a social network that someone follows. For example, a scientist might always pay attention to what a certain expert from the Massachusetts Institute of Technology (MIT) recommends to read and follow that expert rather than following others in the field. I thought that was a very far-forward-looking vision for Mr. Gates’s talk in 2005.
Another important issue is being able to identify datasets. DataCite4 is promoting the use of Digital Object Identifiers for datasets. The scientific community needs to have easier access to research data and to be able to make the data citable. The Open Researcher and Contributor ID (ORCID)5 collaboration is trying to solve the issue of contributor name
__________________________
4 DataCite is an international consortium to establish easier access to scientific research data on the Internet, and increase acceptance of research data as legitimate, citable contributions to the scientific record, and to support data archiving that will permit results to be verified and repurposed for future study. (See http://datacite.org/)
5 ORCID aims to solve the author-contributor name ambiguity problem in scholarly communications by creating a central registry of unique identifiers for individual researchers and an open and transparent linking mechanism
ambiguity—to determine how to know, for example, that Anthony J. G. Hey and Tony Hey are the same person. It is a particular problem in Asia. Thus, ORCID and DataCite are important trends.
There is also a revolution being forced upon us in scholarly communication. Dr. Gray called for establishing digital libraries that support other sciences in the same way that the National Library of Medicine does for medicine. He also called for funding the development of new authoring tools and publication models, and for exploring the development of digital data libraries that contain scientific data and not just the metadata. It does not necessarily have to be done within databases, although that is what Dr. Gray was thinking, but we should certainly support integration with the published literature.
I would like to address the journal subscription issue now, and I will use data from the University of Michigan as an example. The University of Michigan libraries are canceling some journal subscriptions because of budget cuts and the increasing costs of the subscriptions (in some cases, 70 percent of the library collection budget). University Librarian Paul Courant said that about 2,500 were canceled in the 2007 fiscal year. The University Library budget has gone up by an average of 3.1 percent per year since 2004, and according to Library Journal magazine, the average subscription price of national arts and humanities journals has increased 6.8 percent per year since 2003. National social science journals increased 9.2 percent and national science journals increased by 8.3 percent. I was trying to get more up-to-date figures, so I contacted a librarian at the university who informed me that there are currently some plans at the university to close some libraries. We should not be closing libraries. We should not spend so much money on subscriptions, especially since the research reported in these journals has already been paid for in some sense.
When I was dean of engineering at the University of Southampton in the United Kingdom, I was supposed to look at the research output of 200 faculty and 500 postdoctoral and graduate students. At the same time, the university library would send me a form every year asking which journals to cancel. If the library cannot afford to subscribe to all the journals in which my staff publish, then how could I start a new field—order some new journals in, for example, nanobioengineering? This is a broken system. Therefore, it seemed absolutely essential to me that a university keep a digital copy of all its research output. As a result, we set up a repository that has now become a university repository.
The university now keeps full-text digital copies of everything that its staff and students produce. It is not just the papers that will eventually appear in Nature or other journals, but it is also the research reports, conference papers, theses, and even software and images. No one is asked to violate the publisher’s copyright, since these are not necessarily the versions that appear in print. About 70 or 80 percent of publishers allow keeping a nonfinal version on the Web. What is important is to get the scholarly products captured at the time of production with the relevant metadata so that it becomes easy for the researchers to access these materials.
__________________________
between ORCID and other current author identification schemes. These identifiers, and the relationships among them, can be linked to the researcher’s output to enhance the scientific discovery process and to improve the efficiency of research funding and collaboration within the research community.
A fellow dean at Virginia Tech told me that he had required electronic theses to be online in 1997. Virginia Tech had 200,000 requests for PDFs that year. Less than 10 years later they had 20 million requests. These are extraordinary figures. It shows that making your data available is important.
I do not advocate Webometrics as a way of ranking departments or universities, but looking at just the Google Scholar ranking, which looks at papers and citations, we get the list shown in Figure 1-7.
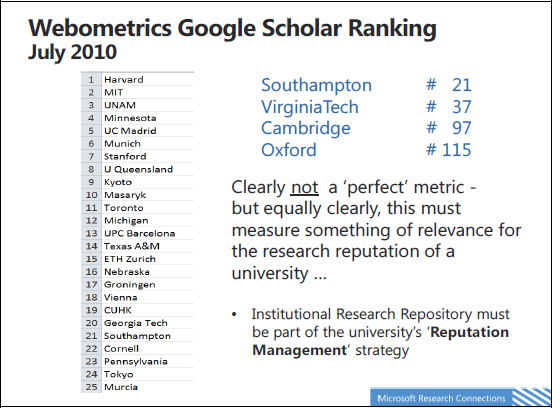

FIGURE 1-7 Webometrics Google Scholar ranking.
SOURCE: Microsoft Research
Harvard University and MIT are at the top. But there is also a large amount of apparent randomness in this ranking. According to citations in Google Scholar, the top university in the United Kingdom is Southampton, at number 21, well above Cambridge and Oxford. This might appear hard to believe, but it is saying something serious about research. If a university is regarded to be a serious research university, it should be showing up on such ranking, because people like such tables. It is a question of reputation management, and I believe that the university research repository is an important piece of that.
As to the future of research repositories, I think they will contain not only full text and the other kinds of scholarly products that I have talked about but also data, images, and software as part of the intellectual output. Some of those items may have an identifier attached that can be used, for example, in a court case. In my view, the university library should be doing that.
Let me offer two examples of how work in this area can be done. There is a large amount of data in databases at the National Library of Medicine so that researchers can have a “walled garden.” Dr. David Lipman and colleagues at the National Institutes of Health have
created a system that allows anyone not only to have access to the text provided through PubMed Central but also to have a range of databases and cross-database searching. That is a centralized repository model.
Most fields are not going to be fortunate enough to have such a centralized model, because it is expensive; therefore, it is important to develop a more distributed model. One example is WorldWideScience.org. Dr. Walter Warnick and his colleagues at the U.S. Department of Energy are building a consortium whose membership is now up to 65 nations. It offers many publications and databases online, and there is a translation service that allows people to type a query in English, have it translated into Chinese or Russian, and after the search is done, translate the results back into English. The major advantage of this search system over a Google search is that this system can get up to 96 percent unique results that cannot be captured by a simple Google search. This is an example of a distributed system that is getting close to Dr. Gray’s vision.
We all know about Vannevar Bush’s article “As We May Think5.” In a similar vein, the physicist Paul Ginsparg wrote a good article called “As We May Read6.” Prof. Ginsparg started arXiv at Los Alamos National Laboratory for use by high-energy physicists. His idea was to reproduce the way that physicists used to circulate preprints back when papers were typed. They used to make photocopies of papers they were submitting to, for example, Physical Review and send them around to a hundred institutions. Since he had access to a server, he suggested to other physicists that they send digital copies of their papers to him and he would circulate them. That is essentially how the arXiv started. The project is now located at Cornell University.
In the conclusion of his paper, Prof. Ginsparg wrote, “On a one-decade timescale it is likely that more research communities will join some form of global unified archive system without the current partitioning and access restrictions familiar with the paper medium for the simple reason that it is the best way to communicate knowledge and hence to create new knowledge. Ironically, it is also possible that the technology of the 21st century will allow the traditional players, namely the professional societies and institutional libraries, to return to their dominant role in support of the research enterprise.7”
Next, to talk about future research cyberinfrastructure, I thought I would go back in history and look at a slide from 2004. These are the six key elements for a global cyberinfrastructure, as opposed to e-Infrastructure, which was the term used in Europe.
__________________________
5 Atlantic Magazine, July 1945.
6 The Journal of Neuroscience, 20 September 2006, 26(38): 9606-9608; doi: 10.1523/JNEUROSCI.3161-06.2006
7 Ibid.

FIGURE 1-8 Six Key Elements for a Global e-Infrastructure for e-Science.
SOURCE: Microsoft Research
I highlighted the points on technologies and standards for data provenance, curation and preservation, and open access to data and publications by interoperable repositories. I am not sure we have made much progress since then, but it was interesting that we were thinking along those lines 7 years ago. We funded a number of projects. We set up a digital curation center in 2004. It took the people there some time to find their way, but I think they are doing useful work now. They advise people about data management plans, for example. We also set up a software engineering organization called Open Middleware Infrastructure Institute. Its job was to take some of the software we produced in our e-science projects that got the most use and then do testing, documentation, writing specifications, and finally make the software available. The institute worked to make software usable to scientists who had not written it. It brought the level of the open-source software up to a reasonable engineering level. The NSF should be helping its grantees do similar work.
Another example is the National Center for Text Mining (NCTM). My colleagues in the computer sciences area claim that the technology is now mature. I do not think that is the case. I do not think we have proven that intelligent text mining is used by the community yet. The NCTM can be considered as an experiment that is only halfway successful in this area.
I should also mention JISC, which is an NSF-like funding agency in the United Kingdom. JISC funded many pioneering activities in digital libraries and repositories. I was the chair of the JISC Committee for Support and Research. One project that JISC funded was the TARDIS project, which led to the repository at Southampton. Another JISC project was the MALIBU project, which was about hybrid libraries and the invisible Web—a lot of information is locked behind Z39.50 and not accessible to Web browsers, so how could anyone find it? The final example was the CLADDIER project with the British Atmospheric Data Center, which was concerned with linking data to publications, and, again, it was very interesting.
Let me now focus on where we are going in the future. At present, about two papers are being deposited in the National Library of Medicine every minute. No one can keep up with this rate, so how do we deal with that deluge of publications? That does not even include the data generated for those papers. I think that semantic computing will help. Currently computers are great tools for storing, computing, managing, and indexing huge amounts of data. In the future, we will need computers to help with automatic acquisition, aggregation, correlation, interpretation, discovery, organization, analysis, and inference from all of the world’s data. If the system knows that a scientist is looking for information about a star, for example, it should be able to identify which papers contain data about that star and get those papers for the searcher without being asked. It should “understand” what we are looking for. The computer should be computing on your behalf and understanding your intent. These are the important things that we need to do.
I do think that we are moving to a world where all the data are linked, and that at some point a scientist will be informed that paper X is about star Y without even searching for this paper. This is what semantic computing will be able to do. This is coming and is an exciting challenge for the information technology industry and for computer science to see if they can build systems that are useful.
I also think that, in the future, researchers will keep some data and tools locally, but they will also use some services from the cloud. We already know about blogs and social networking in the cloud as well as identity services. Amazon and Microsoft offer storage and computing services. Thus, depending on the area or application, scientists may want to use some kind of cloud service. For example, a scientist may use the cloud to replicate the data to a safe place to store them or to share them for collaboration purposes. Semantic computing is also coming, and I think that we are going to realize Dr. Gray’s big digital global library that contains data and publications.
DISCUSSANT: There is a scientist at Rutgers University named Christopher Rose who is known for, among other things, his paper about why the search for extraterrestrial intelligence is not worthwhile. His point is that the energy required to fill the universe with so much energy that somebody in another galaxy can pick it up is absolutely unbelievable. He also said that the only thing that makes any sense is that if someone throws an object toward a place outside his or her galaxy, it means that this person has to produce half the energy, and the recipient has to produce the other half in the form of a radar system that detects it. When I hear someone saying that we are going to fill the world with knowledge in the hope that you pick up some of it as it passes your head, I wonder if there is not something to be said for the traditional system of trying to put the person who is producing a bit of chemical knowledge in a room with the people who might use chemical knowledge and see if they recognize the intersection.
DR. HEY: It is a vision, and visions can be wrong. I think that we can do much better. We spend a lot of money in the biological sciences, for example, with everybody repeating similar work.
Also, there was a medical study recently released that found that almost 400 medical research projects did not cite obviously relevant papers on the same subject. I think we can do better. I do not think we will fix all the problems, but I do think we are all fortunate enough to have a National Library of Medicine.
DISCUSSANT: Dr. Hey, if you look at where the scientific community is going with publications and data linkage, arguably this is already being done in the real world. If you go to the New York Times Web site, for example, or other publications’ sites, you see that the private sector has provided the engine to have the kinds of searchable ranked articles that we would like to have in the scientific community. What do you think it will take to close the gap? And what is the next level of developments that we should be aiming at today?
DR. HEY: If there are commercial systems that work, then we should use them, and we should not try to reproduce tools and open-source software with our limited research funds. It is absolutely important to have standards and interoperability, but there are experiences that we can learn from the commercial world.
Where we are going? Supercomputer centers produce huge amounts of data, and it will be difficult to manage and transfer the petabytes of data that they produce, so I think that we will have supercomputer centers with data centers. I think there will probably be some regional data centers. There are currently attempts to put some of this infrastructure in place, and I think there is an opportunity for the United States to be leading in this exercise. I am pleased to see that NSF is galvanized to do something in this area.
DISCUSSANT: Much of what was shown in the presentation, including the slide from Bill Gates’s talk in 2005, now exists. Tools like Mendeley for sharing publications and fancy XML documents with objects embedded inside of documents with different formats do exist. But in astronomy, maybe 1 percent of the astronomers use them and maybe 5 percent know that they exist. What did we learn about this culture shift that is needed to let us use tools and technologies more efficiently?
DR. HEY: I absolutely agree. We used to produce useful tools and give them to the users, but we found that they were not using them 6 months later. Even though we would show them how useful they were and they had all agreed, they did not use them. There is a real problem in engaging scientists. We have to produce tools that are as close as possible to the way that scientists are working now, but that at the same time give them an improvement. Advocates are needed, and it will be important to have some people showing how scientists can do great research with these tools.


















