2
Convergence in Action
Before turning to the more detailed discussion of institutional frameworks for fostering convergence, this chapter discusses five examples that illustrate why convergence is a fruitful concept and demonstrate its power to yield scientific insights applied to real-world problems.
2.1 A KNOWLEDGE NETWORK WILL
IMPROVE DISEASE TREATMENT
“One of the most exciting illustrations of the possibilities of convergence is in the area of precision medicine. By precision medicine I mean the notion of treatments, diagnostics, approaches to the patient that are increasingly focused on the individual specific needs of that particular patient” (Harvey Fineberg, Workshop on Key Challenges in the Implementation of Convergence, September 16-17, 2013, Washington, DC). Recent advances in biomedical research have created an explosion of data that could be used to develop this concept of precision medicine—a more precise and more accurate classification of disease that could revolutionize diagnosis, therapy, and clinical decisions, leading to more individualized treatments and improved outcomes for patients. For example, as the cost of sequencing genomes continues to fall, the increase in the amount of available genetic data is boosting understanding of the root causes of specific diseases and conditions, such as high cholesterol. Using genomic data, scientists have already found that a significant number of patients
with high cholesterol have a nonfunctional copy of a gene that encodes a low-density lipoprotein receptor. For these individuals, lifestyle interventions such as diet and exercise alone are ineffective at reducing the early onset of cardiovascular disease. Identifying these patients would allow their doctors to prescribe statin drugs at an early age, rather than first attempting to control cholesterol with diet and exercise.
However, it can take years for advances in biomedical research to reach doctors and patients, and in the meantime health care expenditures are incurred for treatments that are only effective in specific subgroups. Although the increasing use of electronic patient records is making it easier for healthcare providers to record clinical data, researchers frequently do not have access to this base of information. In order to find groups of patients for study, researchers often use informal referral networks to identify physicians working with patients with diseases of interest. Regulations that govern patient privacy, such as the Health Insurance Portability and Accountability Act (HIPAA) also create obstacles to linking genomic and clinical data.
Convergence among the biomedical, technological, clinical, and regulatory fields could help create a knowledge network for precision medicine that integrates these multiple sources of information (Figure 2-1). Molecular data, medical histories, information on social and physical environments, and health outcomes could be continuously updated and made accessible to the research community, health care providers, and the public. Analyzing connections between information sets—for example, information on patients’ genomes and environmental exposures—would help scientists to formulate and test disease mechanisms and clinicians to develop new treatments based on unique features of a disease and tailored to each patient.
The knowledge network created by this vision is a tool. One example of the type of impact it might eventually enable would be to improve treatment and enhance disease survival rates among multiple demographic and socioeconomic groups, an outcome that would require life, medical, behavioral, social, and systems science contributions. The recognition of social factors in contributing to health disparities, increasing compliance with medical recommendations, and improving the organization and delivery of health care provides an additional opportunity to engage the expertise of many fields (Pescosolido et al. 2012).
2.2 THREE DIMENSIONAL PRINTING WILL
BRING NEW HEALTHCARE OPTIONS
After years of debilitating pain caused by arthritis, patient Brook Hayes was in urgent need of a hip replacement. But her orthopedic
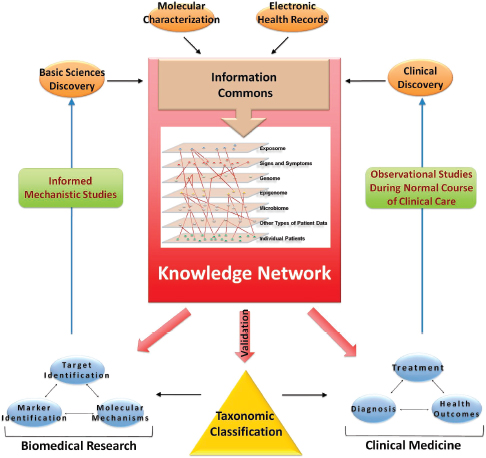
Figure 2-1 Building a biomedical knowledge network for basic discovery and medicine. At the center of a comprehensive biomedical information network is an information commons that contains disease information linked to individual patients and is continuously updated by a wide set of new data emerging though observational studies during the course of normal health care.
SOURCE: NRC 2011a, p. 52.
surgeon explained that Brook’s small stature and severe hip deformities meant that standard hip replacement surgery would not work. She needed customized hip implants—and to build them, her medical team turned to three-dimensional (3D) printing technology (Mayo Clinic 2013). Used industrially for the past several decades, 3D printers work by following instructions from a computer to deposit thin layers of material into structures. Because there is no human operator or hard-coded machine instructions, and no molds to design or special tools needed, 3D printers can build a wide variety of objects from scratch in hours. The technology
allows engineers to experiment with new ideas and numerous design iterations quickly and cheaply, and is often used to develop intricate parts used in architecture, industrial design, automotive and aerospace engineering, the military, and civil engineering (Gross et al. 2014).
Convergence between the engineering and biotechnology worlds is now bringing 3D printing to medicine. Customized joint implants and other medical devices such as dental implants or hearing aids can be designed using computerized axial tomography scans of the patient’s body and constructed on demand. Not only are the finished devices the right size and shape for the patient, but the quickly fabricated models also give surgeons an opportunity to practice difficult procedures before entering the operating theater.
Although most 3D printers make objects from plastics, researchers around the world are working on ways to 3D print with living cells to construct organs and tissues for research and for use in transplantation. Unlike traditional tissue engineering techniques, which involve seeding living cells on an artificial scaffold to build cartilage, bone, or muscle, biological 3D printing will, in theory, allow researchers to control the placement of cells and other components to more closely mimic natural structures (Figure 2-2) (Fountain 2013; Ozbolat and Yu 2013).
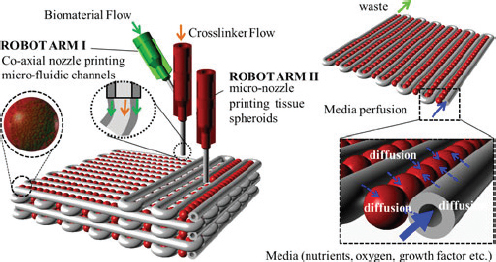
Figure 2-2 Conceptual model of three dimensional printing for organs and tissues. Droplets of cells and biomaterials are deposited layer by layer in 3D assemblies. A separate nozzle creates microfluidic channels that can circulate perfusion media.
SOURCE: Copyright 2013 IEEE. Reprinted, with permission, from Ozbolat and Yu 2013, Fig. 5.
Applying 3D printing to biomedical uses requires integrating knowledge from life sciences on how to sustain the viability of cells throughout the printing process, from material science on biocompatible scaffolding to support the cells, and from electrical and mechanical engineering to design and construct the device. Addressing this research frontier by bringing together multiple fields has led to progress that no single discipline could have achieved alone.
Printing a complex, functional organ such as a heart or liver for patient transplantation is still beyond reach, but academic and industrial research communities have made progress toward this goal, for example, by constructing strips of liver cells that can be used to test experimental drugs. Other laboratories have experimented with 3D printing of skin or heart cells that might be used for skin grafts or to repair damage from heart attacks (Chang et al. 2011; Guillotin and Guillemot 2011; Ferris et al. 2013; Koch et al. 2013; Organovo 2014). Further contributions will be needed for novel 3D products, such as printed bones, tissues, and organs, to move through the necessary regulatory channels to be approved for widespread use in clinical settings.
2.3 CONVERGENCE OCCURS IN FEDERAL AGENCIES: ARPA-E
Convergence occurs not only in academic settings, but is also embedded in innovative government programs. The Department of Energy’s Advanced Research Projects Agency–Energy (ARPA–E) was authorized under the 2007 America COMPETES Act and established in 2009 with a goal of “translat[ing] science into quantum leaps in energy technologies” (Majumdar 2013). The program has had several successful convergent research efforts during its short history. One such project aimed to bridge the fields of synthetic biology, microbiology, and chemistry to develop electrofuels—a new, potentially more efficient type of renewable fuel (Ritter 2011; ARPA–E 2014).
Currently, most biofuels are produced from plants, which harvest the sun’s energy and convert it into chemical energy via photosynthesis. However, photosynthesis is an inefficient energy-conversion process, and growing sufficient plant material requires large areas of farmland and inputs of resources such as water and fertilizer. In contrast, electrofuels are produced by microorganisms that are able to get energy directly from electricity or from chemicals such as hydrogen or ammonia. The bioreactors that house the microorganisms do not require large amounts of agricultural land and have the potential to be produced more efficiently.
The idea of developing the ARPA–E electrofuels program came out of a 2-day workshop and met with skepticism when it was first proposed (Majumdar 2013). However, within 18 months, the effort had succeeded
in bringing together expertise from partners within industry and academia to create an electrofuel using hydrogen and carbon dioxide as input resources. For example, OPX Biotechnologies, a Colorado-based renewable chemicals company, used microbial extremophiles to produce energy rich, long chain fatty acid molecules for a variety of industrial applications. Meanwhile, synthetic biologists at North Carolina State University developed chemical processes to convert fatty acids into liquid fuels. To create the new electrofuel, researchers isolated a bacterium with the natural ability to use hydrogen and carbon dioxide for growth and used a proprietary technique called EDGE (efficiency-directed genome engineering) to modify the microorganism to divert energy and carbon away from growth and toward the production of fatty acids—making it more efficient for fuel production. Synthetic biologists next used a series of chemical processes to convert the fatty acid precursors into liquid transportation fuels. The project exemplifies how an approach that drew on the complementary expertise of several groups of researchers could optimize electrofuel production.
2.4 CONVERGENCE OCCURS IN INDUSTRY: BIOTECHNOLOGY
Industry often has a naturally interdisciplinary outlook to problem-solving approaches, and is an ally and partner in many convergence efforts.1 The history of convergence in commercial biotechnology provides a view of how organizational cultures that support research and innovation have developed in the past in these settings. The example of the founding of Illumina (see Box 2-1) may also provide insights on the process of fostering convergence that are applicable for institutions beyond the commercial sector.
Commercial innovators both catalyze and deliver the social benefits of research convergence. Their position in the biomedical enterprise often enables them to engage the challenges and embrace the opportunities of convergence. The institutional flexibility of commercial enterprises can lower hurdles to convergence because a dynamic commercial sector ignores boundaries between domains of knowledge and removes barriers in pursuit of new opportunities. Moreover, commercial organizations are active at the interface of convergent research and the broader social context that can accelerate or hinder new developments. The participation of those with expertise in nontechnical areas including finance, economics,
_______________
1Mention of specific companies in the report is made for illustrative purposes and reflects information obtained during the committee’s data gathering or from committee members’ knowledge of the field. Such mention does not imply committee, National Research Council, or sponsor endorsement of any commercial product or service.
Box 2-1 Illumina, Inc.: An Example of Convergence
The founding and growth of Illumina is a story of convergence that required the combined knowledge and talents of scientists, engineers, venture capitalists, and business managers—along with some good luck. Key ideas for Illumina’s BeadArray technology were developed by David Walt and his laboratory at Tufts University from their research on fiber optic sensors and chemical monitoring. Their work, which started out using high-power lasers at Lawrence Livermore National Laboratory and small numbers of painstakingly made fibers, benefitted greatly from the development of cheap and robust optics components by the telecommunications industry. Similarly, independent developments in materials science enabled small reaction wells to be etched into a surface for optical interrogation. Parallel work on nucleic acid synthesis and bead-based combinatorial chemistry inspired the idea of “microarrays” of etched wells each containing a randomly deposited chemical probe bead with optical encoding. The high density, built-in redundancy, reproducibility, and flexibility of the new system made numerous applications possible (David Walt, personal communication).
Illumina was founded in 1998 with support from venture capitalists who recognized these attributes and the potential market opportunity in nucleic acid measurement. The company’s continued success required convergence of science and technology depth with business expertise in market analysis and in the development of products and applications. One decision was to get involved in the international HapMap genotyping project cataloguing human genetic variation, which provided an opportunity for Illumina to demonstrate its technology. The acquisition in 2007 of the company Solexa also contributed to Illumina’s ability to extend its technology. Solexa had developed nucleic acid sequencing by synthesis (SBS) derived from the earlier research of Shankar Balasubramanian, David Klenerman, and others at Cambridge University. The technology combined chemistry and informatics to read single bases of DNA as they were incorporated into a growing strand. The combination of Illumina’s core BeadArray and SBS technologies in turn formed the foundation for the recent launch of the HiSeq X high-throughput sequencing platform, which is reportedly capable of generating up to 600 gigabytes of genomic data per day. According to the company, the HiSeqX Ten platform, combining 10 HiSeqX systems and designed for very large scale human genome sequencing, is now capable of reaching the $1,000 genome milestone, which has long been a goal of the life sciences community (Illumina 2014).
public policy, and enterprise management can also serve as a boon to the growth of convergent research and the fruition of its benefits.
To have an idea of how research convergence may play out in the future, it can be instructive to look at examples from the past. Molecular biology is an illustrative case. The need and opportunity to understand cell biological events at a chemical level resulted in the convergence of cell biology and biochemistry in the form of molecular biology. Today molecu-
lar biology is recognized as the unification of the knowledge domains and methods of its foundational disciplines to create a new discipline. What started out as interdisciplinary collaborations became dedicated molecular biology scientific conferences, grant programs, research journals, university departments, degree programs, new technologies and applications, and new industries. A contemporaneous advance in public policy, the Bayh-Dole Act of 1980, decentralized the control of federally funded intellectual property and heralded a proliferation of high-tech companies. Genentech, Amgen, and Genzyme are among the companies that dedicated themselves to the molecular biology convergence.
More recently, structural and functional genomics, mathematics, and computational science have transformed biology into an information science. By converging disciplines—including molecular biology, chemistry, optics, micron-scale manufacturing, combinatorics, and bioinformatics—genomics has catalyzed the development of further genome-scale research in such areas as proteomics and other “omics” fields. Genomics represents an acceleration of the convergence of biology with traditionally nonbiological disciplines like physics and engineering. Market leaders such as Illumina, Agilent, Affymetrix, and their commercial and academic forerunners have helped to advance this convergence, the full benefits of which have yet to be realized.
Systems biology builds further on the genomics-catalyzed transformation of biology into an information science as it incorporates not only engineering in practice, but also engineering design principles and control theory as a means to understand and predict the behavior of complex biological systems. Similarly, branches of mathematics and computer science that address the integration of disparate data types and the analysis and modeling of molecular and cellular networks are key contributors. The past 15 years have seen the creation of systems biology journals, grant programs, conferences, and education initiatives to meet the broader needs of the community engaged in this research, and systems biology has been incorporated into the biotechnology and pharmaceutical sectors. Merrimack Pharmaceuticals, for example, is a company premised on the idea that a systems-level understanding of cancer is important for the development of safe and effective therapeutics.
The ongoing convergence revolution in biology will bring more opportunities that can be translated into commercial and social successes. A likely example is the budding field of synthetic biology, which is in many ways a corollary to systems biology. If one can use engineering theories and design principles to understand and predict the behavior of biological systems, one can use this knowledge to design new biological systems with desired properties for a wide array of applications in health, energy, agriculture, manufacturing, environment, and information and
computation. A critical demand of synthetic biology is the ability to make large numbers of new candidate components (e.g., genes) from which to construct biological entities. Startups like Gen9 and Ginkgo BioWorks are working to meet this convergent need.
2.5 CONVERGENCE STIMULATES THE BIOBASED ECONOMY
The final example of convergence in action focuses on its role in stimulating an innovation ecosystem that develops around and in partnership with an academic research center. Many convergence institutes and programs incorporate elements of entrepreneurship. For example, the University of California’s Institute for Quantitative Biosciences (QB3) trains its graduate students and postdoctoral fellows on how to start companies and supports them with a concept called “startup in a box.” Over the past 2 years, this service has reportedly helped students and postdoctoral fellows incorporate 140 companies, 35 of which are functional and rent space from QB3 for a nominal fee in low-cost facilities for small startups. QB3’s
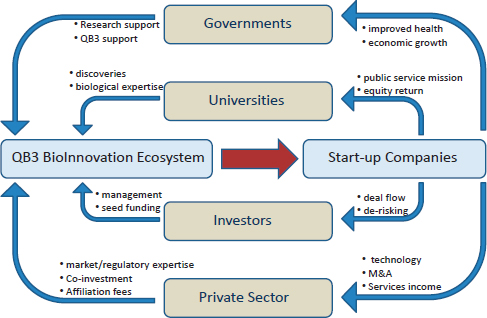
Figure 2-3 The convergence innovation ecosystem at QB3 involves dynamic interactions with government, university, and industry partners.
SOURCE: Courtesy of Regis Kelly, Director, California Institute for Quantitative Biosciences (QB3).
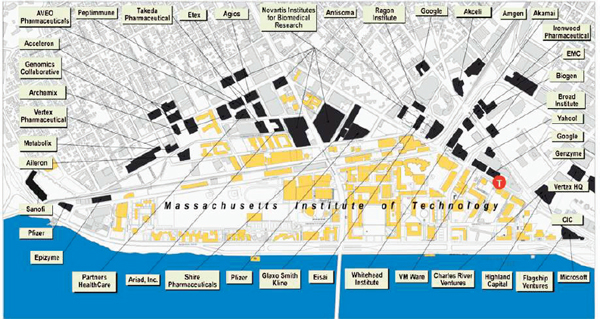
Figure 2-4 Multiple technology companies are located near MIT and form part of the ecosystem created and sustained by convergence.
SOURCE: Courtesy of Phillip A. Sharp, Institute Professor, Massachusetts Institute of Technology
incubator has created over 300 jobs and brought in $150 million from small business grants and angel and venture investments, reportedly for an investment by QB3 of $1 million (Kelly 2013). This interest is a consequence of QB3’s efforts to create a community in which a wide range of stakeholders interact around developing new products and applications, and this ecosystem is helping to foster convergence in new ways (Figure 2-3). The Wyss Institute at Harvard University similarly reports that its researchers are judged not only on typical academic metrics such as journal publications but also on intellectual property, corporate alliances, licensing agreements, and a 5-year technology pipeline, resulting in an institute culture that is less like a traditional academic institution and more like a technology startup (Ingber 2013).
The inclusion in many convergence programs of a focus that spans discovery and application echoes the recommendations made in the ARISE II report on the need for integration across scientific disciplines as well as stakeholders (American Academy of Arts and Sciences 2013). It has been said that innovation moves through people connecting with each other and moving back and forth between a campus and nearby institutions where they can be creative (Sharp 2013). This phenomenon is demonstrated in the cluster of technology companies that have developed in the Boston metropolitan area around institutions such as MIT, Harvard, and Massachusetts General Hospital (Figure 2-4). Similar ecosystems, in which new companies have moved in or sprung up in association with academic centers, are also being formed around convergence hubs such as Stanford University and the University of Michigan North Campus Research Complex.
This page intentionally left blank.












