Proceedings of a Workshop
September 2016 |
IN BRIEF |
Refining the Concept of Scientific Inference When Working with Big Data
Proceedings of a Workshop—in Brief
Big Data—broadly considered as datasets whose size, complexity, and heterogeneity preclude conventional approaches to storage and analysis—continues to generate interest across many scientific domains in both the public and private sectors. However, analyses of large heterogeneous datasets can suffer from unidentified bias, misleading correlations, and increased risk of false positives. In order for the proliferation of data to produce new scientific discoveries, it is essential that the statistical models used for analysis support reliable, reproducible inference. The Committee on Applied and Theoretical Statistics (CATS) of the National Academies of Sciences, Engineering, and Medicine convened a workshop to discuss how scientific inference should be applied when working with large, complex datasets.
CREATION OF NEW OPPORTUNITIES, RISKS, AND CHALLENGES
On June 8-9, 2016, participants described many promising applications of Big Data analysis—for example, to provide insights about the mechanisms of complex biological processes and to support development of personalized medical treatments. Yet, as Michael Daniels (University of Texas at Austin) explained, creating decision-relevant knowledge from this tremendous amount of information hinges on making reliable inferences. Causal inference is particularly challenging when drawing from large, diverse datasets that reflect many phenomena across a wide range of scales. Daniels emphasized the importance of moving beyond point estimation by formally quantifying uncertainty, although tools and methods for doing so are currently lacking. Thus, he cautioned researchers not to publish strong conclusions without an understanding of uncertainty, particularly given the risk of false discoveries and the growing concern regarding irreproducible research.
Alfred Hero (University of Michigan) reminded participants that most Big Data is collected opportunistically, as opposed to intentionally for the analysis at hand, which can yield a variety of problems related to lack of controls, unidentified bias, missing and irregular data, and an abundance of confounding factors. For these reasons, bigger data does not always yield better inferences or predictions, he said. During a discussion of comparative effectiveness research using electronic health records, Sebastien Haneuse (Harvard University) encouraged researchers to compare opportunistic data to that which would result from a dedicated study designed to answer the question at hand and to use this comparison as a guidepost in assessing the suitability of available data. Even when available data do contain the appropriate information, Genevera Allen (Rice University and Baylor College of Medicine) explained, analysts face a host of data curation and statistical modeling
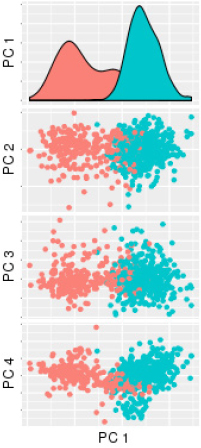
![]()
challenges. In one example, she illustrated confounding batch effects that can arise from seemingly small differences in data collection and processing protocols within different labs, across measurement platforms, or simply from instrument maintenance, as shown in Figure 1. Roderick Little (University of Michigan), commenting from the audience, noted that data preprocessing can affect the reliability of the downstream inference, and he encouraged statisticians to carefully study and consider upstream processing of data as part of their analyses.
Often the information needed to answer a specific question is not contained in the available data, and Robert Kass (Carnegie Mellon University) suggested analysts closely evaluate the suitability of available data to support the inference task prior to analysis. In one example, Dylan Small (University of Pennsylvania) identified natural experiments within large electronic health records data. Audience member Mitchell Gail (National Institutes of Health) suggested that the statistics community work to define a taxonomy of questions and corresponding scientific goals that may be addressed through analysis of large, heterogeneous datasets—for example, based on the level of mechanistic understanding of underlying phenomena—to clearly distinguish targeted studies from hypothesis-generating activities.
Genevera Allen also emphasized the difference between exploratory and confirmatory analyses, and she suggested that the community develop better tools to interpret and communicate the significance and uncertainty of findings from exploratory analyses. Traditional measures of significance such as p-values may be misleading with large, complex datasets, where the large number of data points can create an illusion of significance, said Cosma Shalizi (Carnegie Mellon University). While this may be well understood within the statistics community, he encouraged the development and dissemination of new diagnostic tools that can be used easily by those without extensive training in statistics such as bench scientists and policy analysts. Elizabeth Stuart (Johns Hopkins University) called for formalizing the sensitivity of analyses to underlying assumptions, particularly in the context of drawing inference from Big Data, as a necessary step for understanding the significance and robustness of results.
Beyond methodological and analytical challenges, practical difficulties such as version control, data storage, data sharing, and computational complexity are exacerbated in the analysis of large, complex datasets, said Andrew Nobel (University of North Carolina at Chapel Hill). He described how increasing computational demands could lead to long run times which in turn might reduce investigators’ willingness to replicate analyses. Nobel called for more transparent and accessible documentation concerning the versioning and evolution of datasets and statistical models to account for multiple testing (“cherry picking”) in analysis of Big Data. Nobel said, “These are the kind of haystacks that we need to count” to evaluate the significance of any “needles” revealed in the analysis.
TRANSFORMATION OF STATISTICS EDUCATION
Underlying many of the challenges associated with inference from Big Data is the need to reform statistics education, remarked Emery Brown (Massachusetts Institute of Technology and Massachusetts General Hospital). He suggested that students be introduced to probability theory, statistics, and data analysis content early: “Give me six weeks out of every year starting in eighth grade … and create a longitudinal course that matches up [across years]. We would change science dramatically.” Brown hopes that teaching statistics in a repetitive and reinforcing manner will help students develop statistical intuition and reasoning, and avoid misconceptions and the impossibility of mastering statistics in a single undergraduate course. Robert Kass suggested that the undergraduate curriculum be organized around fundamental principles of statistics as opposed to introducing students to a sequence of tests based on the available data.
Cosma Shalizi argued that the need to analyze large datasets has not revealed flaws in the concept of statistical inference, but rather has exposed the limitations of “the lies we tell our children” in the form of simplified assumptions used in introductory statistics courses. He described the need for undergraduate statistics curricula to clearly communicate the limitations of such assumptions and provide training in more sophisticated methods, so that inappropriate models are not applied blindly. Andrew Nobel emphasized the importance of teaching statistics students at all levels the basics of statistical computing, database management, and data sharing practices that are increasingly important with Big Data. Jonathan Taylor (Stanford University) agreed but added that good programming skills are not adequately rewarded in statistics programs today and that professors need to lead by example.
As the field of data science continues to grow and new academic programs are created, Joseph Hogan (Brown University) expressed concern that statistics programs will face decreasing enrollment because potential students may not be aware of their fundamental connection to data science. He encouraged funding agencies to articulate the importance of statistics as an essential discipline of data science when issuing funding announcements such as the National Institutes of Health’s Big Data to Knowledge program. Xihong Lin (Harvard University) described how Harvard’s new interdisciplinary health data science program engages students in laboratory rotations in computer science, informatics, statistics, and
domain sciences. Designing curricula for a strong conceptual understanding and programming capabilities alone is insufficient, said Bin Yu (University of California, Berkeley), who emphasized the interdisciplinary nature of statistics and suggested that students receive formal communication training to improve interactions among statisticians, modelers, and disciplinary scientists.
EARLY PARTICIPATION AND COLLABORATION IN EXPERIMENTAL DESIGN
Xihong Lin encouraged statisticians to consider themselves scientists, as opposed to analysts on the periphery of scientific discovery. She pointed to biostatistics as an example of a field with strong collaboration between statisticians and domain scientists, which has led to widespread adoption of best statistical practices and helped lessen instances of common statistical errors by researchers, journals, and funding agencies. Andrew Nobel seconded the call for statisticians to actively engage their collaborators early in the experimental design process. Such engagement, he believed, would provide statisticians with a better understanding of data preprocessing activities and their impact on downstream inference, as well as help to ensure that appropriate data are collected to answer the motivating research questions. Bin Yu described the value of embedding statisticians in experimental labs, which helps them develop a better understanding of the analyses that are feasible and useful based on available data.
Emery Brown called attention to the importance of developing iterative models and engaging in both exploratory and confirmatory analyses with experimental collaborators. Robert Kass noted that this level of engagement among experimental, modeling, and statistical analyses is the exception and not the norm. Michael Kosorok (University of North Carolina at Chapel Hill) described the importance of building trust and strong personal relationships with domain science collaborators so that statisticians are viewed as constructive members of a research team as opposed to nay-sayers brought in at the end of the research process. He argued that as such collaborations lead to significant scientific advances, domain scientists learn why statisticians are cautious and “eventually realize that [a statistician’s] equivocation means a lot more than somebody else’s confidence.” Concern regarding the perception of statisticians as overly negative was shared by Alfred Hero, who urged statisticians not just to challenge the significance of inferences drawn from large, complex data, but also to suggest experimental strategies that would improve confidence in findings under uncertainty.
CRITICAL OPPORTUNITIES FOR FUTURE RESEARCH
During the opening remarks for the workshop, Michelle Dunn (National Institutes of Health), Nandini Kannan (National Science Foundation), and Chaitan Baru (National Science Foundation) requested suggestions for open research questions that, if solved, could advance the biosciences and other scientific domains.
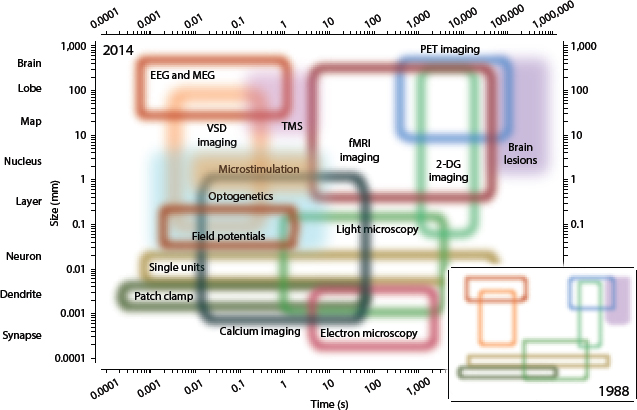
Biomedical research has produced data that describe phenomena across orders of magnitude in spatial and temporal scales, and Alfred Hero called for future research to focus on connecting multiscale data. For example, personalized medicine could be designed more effectively if data describing subcellular processes such as gene and protein expression panels could be mechanistically connected with longitudinal studies such as electronic health records. Providing another example, Jeffrey Morris (MD Anderson Cancer Center) showed the range of temporal and spatial dimensions spanned by currently available neuroimaging techniques (Figure 2) and
said that integrative statistical modeling will be critical to extracting biological knowledge from large, complex data. Connecting these “top-down” and “bottom-up” descriptions of biological processes is more challenging than many other examples of multiscale modeling, said Robert Kass. Andrew Nobel agreed, adding that this is due in part to the inherent complexity of biological processes and the analytical difficulty in tracking multiple processes and mechanisms simultaneously. Integrating top-down and bottom-up descriptions will require sustained collaborations among domain experts, experimentalists, modelers, and statisticians. The breadth of expertise required and the magnitude of this challenge will necessitate significant research investments, concluded Alfred Hero.
Michael Daniels proposed that new methodologies and frameworks for tracking and preprocessing data are required to better understand the causes and types of messiness in Big Data. Genevera Allen agreed, noting that there is likely not one single person who understands all of the data types and preprocessing steps for large datasets such as the Cancer Genome Atlas. In the case of electronic health records, Bin Yu encouraged federal agencies to facilitate data sharing by hospitals and healthcare providers, as many insights could be gained by linking datasets and comparing populations seen at different hospitals or in different regions. A critical bottleneck remains in protecting the privacy of individual patients, and Alfred Hero said he expects privacy concerns to become an increasingly important constraint in the near future.
In presenting methods for network reconstruction, Daniela Witten (University of Washington) pointed to the deep disconnect between statistical theory and practice. She emphasized that models are at best simplified representations of complex biological processes and that their results are subject to numerous assumptions, the validity of which should be scrutinized. Witten said, “Would I tell [biological collaborators] to spend their [entire] career[s] investigating this edge set that I estimated? … The answer to that question is no.”
Looking toward the future, Xihong Lin imagined that computing and data storage in the cloud will become widespread, which will require new approaches to data archiving and sharing. She emphasized the need for models and computational techniques to be scalable and adaptable to different domain sciences. Bin Yu described the possibility that many aspects of statistical analysis could become fully automated in the future, and she said that the community must make sure appropriate techniques are implemented in these emerging analysis packages.
PLANNING COMMITTEE: Michael J. Daniels, University of Texas at Austin, Co-Chair; Alfred Hero III, University of Michigan, Co-Chair; Genevera Allen, Rice University and Baylor College of Medicine; Constantine Gatsonis, Brown University; Geoffrey Ginsburg, Duke University; Michael I. Jordan, University of California, Berkeley; Robert E. Kass, Carnegie Mellon University; Michael Kosorok, University North Carolina at Chapel Hill; Roderick J.A. Little, University of Michigan; Jeffrey S. Morris, MD Anderson Cancer Center; Ronitt Rubinfeld, Massachusetts Institute of Technology; Lance Waller, Emory University. STAFF: Michelle Schwalbe, Committee on Applied and Theoretical Statistics; Rodney Howard, Administrative Assistant; Elizabeth Euller, Senior Program Assistant; Ben A. Wender, Associate Program Officer; Linda Casola, Staff Editor.
DISCLAIMER: This workshop was sponsored by the National Institutes of Health Big Data to Knowledge Initiative and the National Science Foundation. This Proceedings of a Workshop—in Brief was prepared by Ben A. Wender as a factual summary of what occurred at the meeting. The committee’s role was limited to planning the event. The statements made are those of the individual workshop participants and do not necessarily represent the views of all participants, the planning committee, or the Academies. This Proceedings of a Workshop—in Brief was reviewed in draft form by Bin Yu, University of California, Berkeley, and Hal S. Stern, University of California, Irvine, to ensure that it meets institutional standards for quality and objectivity.
SUGGESTED CITATION: National Academies of Sciences, Engineering, and Medicine. 2016. Refining the Concept of Scientific Inference When Working with Big Data: Proceedings of a Workshop—in Brief. Washington, DC: The National Academies Press. doi: 10.17226/23616
Division on Engineering and Physical Sciences

Copyright 2016 by the National Academy of Sciences. All rights reserved.




