In CNS disorders that have a neuroinflammatory or immune component, quantification of analytes in the CSF and blood may enable assessment of the peripheral versus central immune response, determine whether these analytes correlate with development or progression of disease, and help show if inflammation in the peripheral compartment reflects what is going on in the brain, said Brian Campbell. Indeed, some of the strongest evidence supporting the role of neuroinflammation in CNS disease has arisen from biomarker discovery programs. For example, Richard Perrin described an AD biomarker discovery program he conducted with colleagues at the Knight Alzheimer’s Disease Research Center at Washington University (Craig-Schapiro et al., 2011). Using a multiplex immunoassay (Luminex) platform applied to 333 paired CSF and plasma samples from cognitively normal to mildly demented individuals, the researchers measured 190 analytes thought to be important in the disease. These analytes included cytokines, chemokines, metabolic markers, growth factors, and other proteins. The studies indicated that a panel of plasma or CSF proteins can be used to predict who has existing brain amyloid or will likely become amyloid positive and who will likely develop dementia (see Figure 6-1). Markers that predicted conversion to amyloid positivity included anti-inflammatory markers in the CSF and both pro- and anti-inflammatory markers in the plasma, indicating that peripheral inflammation is an early event in AD pathogenesis. Once people develop amyloid plaques, anti-inflammatory markers are seen in both the CSF and plasma along with markers of low adiposity and low insulin signaling. Among those who are likely to develop dementia in the next 3 to 4 years, Perrin said there is a robust neuroinflammatory signal, especially in the CSF, as well as evidence of vasculopathy that may be related to blood‒brain barrier (BBB) breakdown and hypothalamic, pituitary, and metabolic changes. Perrin maintained that the variability in levels of inflammatory biomarkers over the course of disease argues against the disease being a continual process, as has been proposed.
However, Campbell said there remains a lack of consensus in the literature about most CSF and plasma markers that have been measured thus far, which stems largely from small sample sizes, insufficient validation of assays, and lack of standardization in terms of analytes measured and collection, handling, and storage of samples. The Alzheimer’s Disease Neuroimaging Initiative (ADNI), a public‒private partnership established by the NIA and the Foundation for the National Institutes of Health (FNIH) in 2004, specifically addressed these issues of standardization
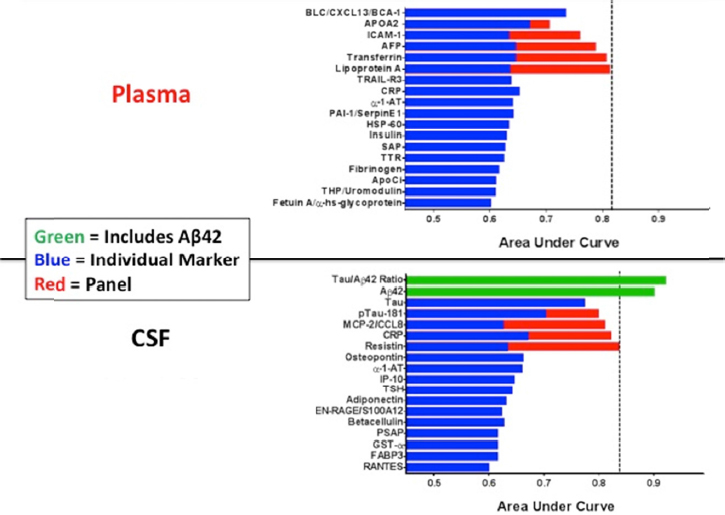
SOURCE: Presentation by Perrin, March 21, 2017.
and validation of AD biomarkers, noted Eliezer Masliah, director of the Division of Neuroscience at NIA. More recently, other consortia have also emerged to build on ADNI’s efforts to standardize and validate assays.
Campbell and Edward Bullmore described two other consortia established over the past 5 years specifically to identify inflammatory biomarkers for CNS diseases: the Wellcome Trust Consortium for Neuroimmunology of Mood Disorders and Alzheimer’s Disease, and the FNIH Biomarkers Consortium’s project on Inflammatory Markers for Early Detection and Subtyping of Neurodegenerative and Mood Disorders. The operations of these two consortia are discussed in more detail in Chapter 7. In both programs, rather than taking the relatively agnostic approach of measuring a large number of analytes across different categories as was
previously described by Perrin, they have focused specifically on a select group of analytes related to inflammation. Perrin said these are complimentary approaches.
The FNIH Biomarkers Consortium project selected inflammatory analytes, such as cytokines and chemokines (C-reactive protein, interleukin [IL]-1β, IL-6, TNFα, IL-10, sIL-6R, and IL-1RA), that will be assessed using enzyme-linked immunosorbent assay (ELISA)-based technologies, as well as tryptophan and kynurenine metabolites, which are effector molecules for the immune system that will be assessed using mass spectrometry assays, said Campbell. He said the Consortium hopes to identify a panel of inflammatory biomarkers—a biosignature—that has sufficient power to be used at the individual patient level for diagnosis, subtyping, and monitoring disease progression or response to therapy. Prior results in bipolar disorder studies support this approach, said Campbell (Brietzke et al., 2012; McIntyre et al., 2014). The Wellcome Trust has selected an overlapping set of inflammatory biomarkers to assess. For major depressive disorder, these include CRP, IL-1β, IL-6, TNF-α, IL-10, and tryptophan and kynurenine metabolites, he said.
Proteomic analysis also suggests that fibrinogen, fibrin degradation products, or other products of the coagulation cascade may also be useful as plasma biomarkers in multiple sclerosis and AD (Han et al., 2008; Lee et al., 2007), said Katerina Akassoglou; however, these are not currently included in either of the programs mentioned above.
NOVEL APPROACHES TO IDENTIFYING GENETIC AND MOLECULAR MARKERS OF NEUROINFLAMMATION
Steve McCarroll is pursuing a different approach to identify genetic and molecular markers of neuroinflammation in schizophrenia and other CNS diseases by assessing secreted biomarkers of neuron–microglia interactions. Schizophrenia is a heritable but extremely polygenic illness shaped by genetic variation in at least 100 loci, said McCarroll. The strongest genetic influence appears to come from genes involved in immunity and infection that reside within the major histocompatibility complex locus on chromosome 6, in particular, the complement component 4 (C4) gene, which has been implicated in the pruning of synapses. McCarroll and colleagues have shown that this gene has many structurally different allelic forms that result in different levels of C4A and C4B expression, and that schizophrenia is associated with particular variants
that lead to elevated C4A expression (Sekar et al., 2016). More recent work from the lab of McCarroll’s colleague Michael Carroll has shown that microglia engulf synapses decorated with C4A.
These discoveries, combined with the knowledge that in humans, key cortical regions undergo maturation and pruning during adolescence and early adulthood, which is also when schizophrenia typically presents, suggest that schizophrenia may result or worsen when a normal developmental process is recruited into pathopysiology. Because C4 is a secreted molecule, it also suggested that it might be possible to “listen in” on the conversation between neurons and microglia by assessing C4 levels in the CSF, said McCarroll. Indeed, their recent paper (Sekar et al., 2016) showed that in postmortem brain tissue, C4A RNA expression is about 40 percent higher in patients with schizophrenia than in normal controls. McCarroll added that this effect is much stronger than the 20 percent effect predicted by the genetic relationship to C4 (i.e., the fact that schizophrenia patients on average have high-C4A-expressing alleles). Because there are limits to the value of a measurement in postmortem brain tissue, McCarroll’s lab has been measuring C4 and other potentially relevant analytes in CSF. They have shown that when postmortem tissue and CSF are both available from the same individuals, the protein levels in CSF correlate strongly with levels of RNA in cortical tissue. Because the cortex is the largest source of CSF C4, this suggests that CSF may provide information about C4 expression in brain tissue. Moreover, unlike C4 alleles, C4 protein levels in CSF are a potential biomarker for dynamic processes that are shaped by genes, environment, and development. Plasma levels of C4, however, are unlikely to be informative because C4 does not routinely cross the BBB, said McCarroll.
McCarroll and colleagues have recently initiated an analysis of C4 as well as other proteins and small-molecule metabolites in a set of CSF samples from youths with attenuated psychotic symptoms who are at ultra-high risk for progression to frank psychosis and schizophrenia. They also hope to apply this approach to a large number of samples from healthy individuals to map out the natural history of these analytes and establish normative values.
McCarroll’s lab has also developed a novel technology to enable high-throughput, single-cell analyses of gene expression to study circuitry change during critical periods of development. This technology, called “Drop-seq,” isolates individual cells in millions of tiny droplets, uses beads to deliver different molecular barcodes to each droplet, and then analyzes the messenger RNA (mRNA) transcripts from thousands of
cells simultaneously while remembering the source of each transcript because of the bar code (Macosko et al., 2015). They then use machine-learning approaches to classify cells into groups or types based on their genome-wide transcriptional patterns, allowing them to create atlases of cell types and cell-type-specific gene expression in different tissues. McCarroll’s lab has been using Drop-seq to identify biomarkers of developmental critical periods—the time in development when the synaptic circuitry is changing very quickly—in both neurons and glia. Because many of these mRNAs encode secreted proteins that can be detected in CSF, McCarroll believes the Drop-seq data nominate new CSF biomarkers and aid in the interpretation of CSF biomarker data, making it possible to identify which cells are the source of a particular biomarker and what distinguishes cells that express that biomarker from cells that do not.






