International Harmonization of Laboratory Animals
Hideki Katoh
Institute for Experimental Animals, Hamamatsu University School of Medicine, Hamamatsu, Japan
and
Central Institute for Experimental Animals, Kawasaki, Japan
STRAINS AND COLONIES USED IN ANIMAL EXPERIMENTATION
Laboratory animals may be divided into three major genetic types: inbred animals, hybrid animals, and closed colonies. Each type is used in animal experimentation in ways that maximize the application of its characteristics.
Inbred Strains
Almost no genetic differences can be found between any two animals within a particular inbred strain. Therefore, use of inbred animals generates better stability and reproducibility of results than closed colony animals in all types of animal experiments. Experiments also typically require fewer numbers of these animals, which is an important advantage with respect to animal welfare. However, because there are major genetic differences from one inbred strain to another (for example, in responses to drugs) there may be completely different results (such as a high response level in one strain and a low level in another). If an animal experiment is performed using inbred animals, it is necessary to perform the experiment first with several different strains to select the most appropriate strain.
Many types of research are also performed by utilizing strain differences in responses such as sensitivity and resistance. Examples of this research include biochemical studies on substances that cause strain differences such as proteins and enzymes and genetic studies on strain differences.
Hybrid Animals
In laboratory animal science, hybrid animals are usually obtained by mating among different inbred strains. They include the following four types: F1 hybrid, F2 hybrid, three-way cross, and four-way cross.
As explained subsequently in Genetic Test Method for Genetic Composition, genetic control of hybrids is easy, and it is possible to produce hybrid colonies with a high degree of reproducibility. Hybrids are considered appropriate for animal experiments because they generally show excellent reproductivity and good health, which compensate for the defects of their inbred parent strains such as low productivity due to inbreeding degeneration and various physiologic and biochemical defects caused by mutant genes. Historically, however, there have been few examples of the widespread use of hybrids in animal experimentation.
Closed Colonies
Closed colonies of rats and mice have long been used as representative species in experiments such as toxicity tests. Gene polymorphism is maintained in closed colonies, and the genotypes of individual animals are known to differ based on genetic testing (Katoh and others 1998). In this respect, closed colonies correspond to human populations; however, it is evident from an understanding of the origins of closed colonies that they cannot always be considered representative of species such as mice and rats. The main reason is that a single population (colony) does not possess all of the genes or gene polymorphisms of the species. There is also a strong possibility that closed colonies will lose their genetic stability because of artificial (human) control. Extreme phenomena (the bottleneck effect) concerning the number of members of colonies associated with microbiologic cleaning, in particular, are likely to occur during cesarean section, and we have experienced several actual examples of this.
SAFETY STUDIES AND LABORATORY ANIMALS
Studies in Which Animal Species Present a Problem
Mice and rats are widely used in new drug development, especially in toxicity (acute, subacute, and chronic) tests. In these studies, the responses are strong as long as the genetic differences in the same species are negligible. The doses are high, and individual differences or strain differences are not likely to appear. Although they are not performed at this time, studies formerly used the LD50 (50% lethal dose; the amount or concentration that causes death of 50% of the animals when a drug is administered) as the parameter. Therefore, primarily closed colonies, rather than inbred animals, have been used historically in this
type of study. In many cases, for example Wistar and Sprague-Dawley rats and ICR and Swiss mice have been used. However, these closed colonies show clear differences in spontaneous malformation rates, spontaneous cancer rates, types of cancer, body weight, and life span.
Many new drugs have been developed using closed colonies, and a large amount of data has been accumulated. When the same type of studies are performed in the future, specific consideration should be given to selection of strains based on materials used in former studies. It is natural to conclude that it is desirable to use the same strain as that used before and the time before that. Even if the strain name is the same, the name of the breeder should be reported.
These precautions are of even greater interest in high-precision research in fields such as immunology, which involves individual genes and molecular genetics. In the past, however, when the causal relation between the response and the related substance (DNA or protein) was vague or unclear, especially in typical animal experiments such as toxicity tests, almost no consideration was paid to selection of the strain.
Studies Showing Individual and Strain Differences
Animal experiments such as those performed in new drug development at present include not only those using mice and rats as typical species such as LD50 studies but also studies in which physiologically or metabolically active enzymes such as P450 or the p53 or H-ras genes are used. In these studies, it is essential to be aware of the possibility that various factors such as strain selection may affect the response because relatively weak responses are measured. The effect of the genetic background on these particular genes is unknown. In the worst case, when the genetic background is inbred, it may not be possible to perform the animal experiment. Unfortunately, however, we have had no time to evaluate the effects of genetic backgrounds of inbred animals on various genes, and we do not know whether such evaluations are worthwhile at this time.
INTERNATIONAL HARMONIZATION OF LABORATORY ANIMALS
At present, international harmonization of data from animal experiments is being promoted by the International Conference on Harmonization (ICH) of Technical Requirements for Registration of Pharmaceuticals for Human Use. The most important items for animal experiments are reflected in Good Laboratory Practice. As discussed previously, the harmonization issues that have caused problems in several testing facilities have been related to laboratory animals (strains or colonies), food, and experimental methodology.
In the case of food, objective (scientific) standards can be established by identifying the contents, for example, in percentages of protein, and for experimental methods and techniques, by preparing records using photographs or imag
ing results. For laboratory animals, however, the only objective information we currently have is that of strain names. Discussions have only recently begun to address the question of whether there is scientific evidence for a discussion on the international standardization of laboratory animals.
The need for international genetic and microbiologic standardization of laboratory animals such as mice and rats was recognized in the early 1980s. Dr. Nomura, among others, recognized that standards should be based on high-quality industrial products as well as strict methods for evaluating this quality. It was agreed that participation in the international market should be contingent on fulfillment of such standards. The results of these initial efforts toward standardization appear in the ICLAS Manual for Genetic Monitoring of Inbred Mice (Nomura and others 1984).
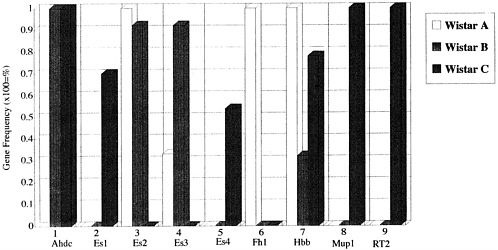
Using genetic testing before genetic monitoring, it is possible to determine the genotypes of individual inbred strains with accuracy. It is also possible to differentiate one strain from another based on whether all animals in a strain have particular genes at the gene loci or whether genotype information for several gene loci is present. This method applies not only to inbred animals but also to closed colonies. An example of this differentiation is shown in Figure 1, in which a gene of the Es3 locus is present in 30% of colony A, 90% of colony B, and 0% of colony C.
An Ideal Global Standard
The facts and examples described above may be used as standards for a closed colony of laboratory animals. Colonies that meet these standards may be considered global standard colonies.
Items to be determined
It is possible to differentiate certain closed colonies from other colonies by the presence of particular genes in the colony or differences in the frequencies of these genes. For these reasons, the types of genes present at a certain time and their frequencies should be determined. In Table 1, the gene frequencies of 21 biochemical markers in closed colonies of the rat are shown. As is evident, Acon1a can be detected in both SD colonies in the frequencies of 0.958 and 0.567, respectively, but cannot be detected in Wistar colonies and Donryu colony.
Prohibited items
Individual animals must not be introduced into another colony either from colonies with the same name or from colonies with different names.
Recommended items for compliance
The following methods are recommended: maintenance method (the rotation system or a system based on it is recommended to assure genetic stability);
TABLE 1 Gene Frequencies of 21 Biochemical Markers in Six Closed Colonies
|
Locus |
Allele |
Jcl:Wistar |
Crj:Wistar |
Iar:Wistar |
Jcl:SD |
Crj:SD |
Donryu |
|
|
1 |
Acon1 |
a |
0.0000 |
0.0000 |
0.0000 |
0.9580 |
0.5670 |
0.0000 |
|
b |
1.0000 |
1.0000 |
1.0000 |
0.0420 |
0.4330 |
1.0000 |
||
|
2 |
Ahd2 |
b |
0.1830 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
|
c |
0.8170 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
||
|
3 |
Ahdc |
a |
0.0000 |
1.0000 |
1.0000 |
1.0000 |
0.9020 |
1.0000 |
|
b |
1.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0980 |
0.0000 |
||
|
4 |
Akp1 |
a |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
|
b |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
||
|
5 |
Alp1 |
a |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
|
b |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
||
|
6 |
Amy1 |
a |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
0.6500 |
1.0000 |
|
b |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.3500 |
0.0000 |
||
|
7 |
Es1 |
a |
0.0000 |
0.0000 |
0.7000 |
0.1920 |
0.3750 |
1.0000 |
|
b |
1.0000 |
0.9670 |
0.3000 |
0.0000 |
0.1670 |
0.0000 |
||
|
c |
0.0000 |
0.0330 |
0.0000 |
0.8080 |
0.4580 |
0.0000 |
||
|
8 |
Es2 |
a |
1.0000 |
0.9170 |
0.0000 |
0.7670 |
0.3080 |
0.0000 |
|
b |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
||
|
c |
0.0000 |
0.0000 |
0.5420 |
0.1670 |
0.0000 |
0.5920 |
||
|
d |
0.0000 |
0.0830 |
0.4580 |
0.0670 |
0.6920 |
0.4080 |
||
|
9 |
Es3 |
a |
0.3330 |
0.9170 |
0.0000 |
0.2920 |
0.3920 |
0.9500 |
|
b |
0.0000 |
0.0000 |
0.4250 |
0.0000 |
0.0000 |
0.0000 |
||
|
c |
0.0000 |
0.0000 |
0.2330 |
0.4670 |
0.0000 |
0.0000 |
||
|
d |
0.6670 |
0.0830 |
0.3420 |
0.2420 |
0.6080 |
0.0500 |
||
|
10 |
Es4 |
a |
0.0000 |
0.0000 |
0.5420 |
0.1580 |
0.0000 |
0.0000 |
|
b |
1.0000 |
1.0000 |
0.4580 |
0.8420 |
1.0000 |
0.1420 |
||
|
c |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.8580 |
||
|
11 |
Es6 |
a |
1.0000 |
0.5520 |
1.0000 |
0.4330 |
0.6670 |
1.0000 |
|
b |
0.0000 |
0.4480 |
0.0000 |
0.5670 |
0.3330 |
0.0000 |
||
|
12 |
Es7 |
a |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
|
b |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
||
|
13 |
Es8 |
a |
0.0000 |
0.9170 |
0.4330 |
0.2330 |
0.7170 |
0.1330 |
|
b |
1.0000 |
0.0830 |
0.5670 |
0.7670 |
0.2830 |
0.8670 |
||
|
14 |
Es9 |
a |
1.0000 |
0.8330 |
0.5670 |
0.7650 |
0.2830 |
0.9150 |
|
c |
0.0000 |
0.1670 |
0.4330 |
0.2350 |
0.7170 |
0.0850 |
||
|
15 |
Es10 |
a |
1.0000 |
0.9170 |
0.5670 |
0.7830 |
0.2850 |
0.9170 |
|
b |
0.0000 |
0.0830 |
0.4330 |
0.2170 |
0.7150 |
0.0830 |
||
|
16 |
Es14 |
a |
1.0000 |
1.0000 |
0.7330 |
0.0000 |
0.5830 |
1.0000 |
|
b |
0.0000 |
0.0000 |
0.2670 |
1.0000 |
0.4170 |
0.0000 |
||
|
17 |
Fh1 |
a |
1.0000 |
0.0000 |
0.0000 |
0.0250 |
0.0000 |
1.0000 |
|
b |
0.0000 |
1.0000 |
1.0000 |
0.9750 |
1.0000 |
0.0000 |
||
|
18 |
Gc |
a |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
|
b |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
||
|
19 |
Hbb |
a |
1.0000 |
0.3170 |
0.7830 |
0.4250 |
0.0000 |
1.0000 |
|
b |
0.0000 |
0.6830 |
0.2170 |
0.5750 |
1.0000 |
0.0000 |
||
|
20 |
Mup1 |
a |
0.0000 |
0.0000 |
1.0000 |
0.0000 |
0.0350 |
0.9420 |
|
b |
1.0000 |
1.0000 |
0.0000 |
1.0000 |
0.9650 |
0.0580 |
||
|
21 |
Svp1 |
a |
1.0000 |
0.5670 |
1.0000 |
0.4000 |
0.7170 |
1.0000 |
|
b |
0.0000 |
0.4330 |
0.0000 |
0.6000 |
0.2830 |
0.0000 |
long-term maintenance method (cryopreservation of embryos should be used); and measurement method of genetic variations (the gene frequency is used as an index).
Genetic test method for genetic composition
Genetic testing may be performed using the methods applied by the ICLAS Monitoring Center. These methods are recommended because a comparison between colonies is necessary.
Standard Candidates to Replace Closed Colonies
If the major problems characteristic of closed colonies are not solved, it is necessary to consider other genetic populations. Candidates are F1 and F2 hybrids as well as three- and four-way cross hybrids.
F1 hybrids
These colonies are obtained by mating between two strains. When the gene loci of A strain and B strain are expressed as A and B, A/B (heterotype) appears at all gene loci in F1 and there is no genetic difference among individuals. Because physiologic or reproductive heterosis appears, the animals show excellent traits such as active behavior or great reproductivity. However, when a genetic evaluation is performed, these colonies are considered less suitable for replacement of closed colonies with their high level of diversity.
F2 hybrids
These colonies are obtained by mating among F1 animals, and the genotype obtained is A/B×A/B. Mendel's law also applies to all gene loci in F2, namely A/A (A parent type), B/B (B parent type), and A/B (F1 type) at a ratio of 1:1:2. It is evident that the genotype frequency and gene frequency can be expressed quantitatively in F2, and the level of reproducibility of F2 is extremely high.
However, there are many genetic differences among individuals in F2 colonies, and individuals may not have exactly the same genotype. Therefore, F2 colonies have the same genetic diversity among individual animals in the colony as in closed colonies, but the level of genetic diversity is low compared with closed colonies because it is determined by the two parent strains.
Three-way cross hybrids
These colonies are obtained by mating between F1 (A/B) obtained by mating two strains (A and B) and a third C strain. The mated animals have either A/C or
B/C at all gene loci. These colonies have the same excellent reproducibility as F2, and it is possible to produce the colony as long as the parent strains are available. The genotype frequency and gene frequency can be estimated from Mendel's law.
As in F2 colonies, the same high level of genetic diversity among individual animals in the colony is present, and the genetic diversity is even greater than that of F2 because there are three parent strains. However, the extent of genetic diversity is still less than that of closed colonies.
Four-way cross hybrids
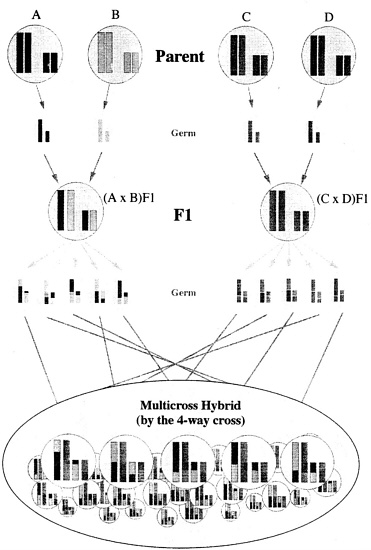
The animals are produced as shown in Figure 2: (1) Two kinds of F1 animals are produced by crossing between inbred strains (that is, line A × line B and line C × line D); (2) different F1 animals are mated, and MCH animals are produced by the four-way cross mating.
Four-way cross hybrid colonies are obtained using four strains (A, B, C and D) by mating between F1 (A/B) obtained by mating strains A and B and F1 (C/D) from mating strains C and D. This hybrid colony also has excellent reproducibility, which may be perpetuated as long as the four parent strains are available. The genotype frequency and gene frequency can also be estimated from Mendel's law.
The genotype of the gene loci of an individual is A/C, A/D, B/C, or B/D. As is evident from so many different genotypes, the genetic diversity among individual animals is greater than that in the F2 or three-way cross colonies, and this colony may be considered artificial, with the highest level of genetic diversity based on the addition of a fourth strain. However, the recessivity is greater than that of closed colonies because the gene source is limited to four strains.
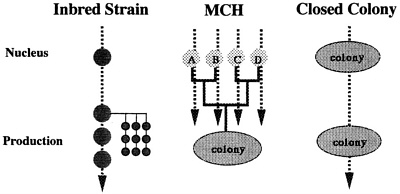
In 1976, development of inbred strains was started by using 40 pairs of closed colony Jcl:ICR. At F9, they were made germ free. Crossing experiments for selection of partner strains were performed using these inbred strains, and four inbred lines were selected. Shown in Table 2 are the genetic profiles of the four inbred strains (IAI, IQI, IPI, and ICT), two F1 hybrids (IAI×IQI and IPI×ICT), and MCH obtained by crossing the F1 hybrids. Pilot production was started in 1983, and various types of basic data concerning the animals produced were collected. According to the results of the experiments performed to date, the Jcl:MCH (ICR) can be used in place of Jcl:ICR. In Figure 3, a comparison of inbred, MCH, and closed colonies is shown. MCH has the following features: (1) MCH is considered an artificial colony with the highest level of genetic diversity; however, its recessivity is greater than that of closed colonies because the gene source is limited to four strains. (2) MCH has excellent genetic reproducibility, which may be perpetuated as long as the four parent strains are available; therefore, genetic characters are balanced and remain unchanged for long periods, which leads to improved reproducibility in animal experiments. (3) Bottlenecks that currently present problems with closed colonies do not develop.
Table 2 Genetic Profiles of the Four Inbred strains Derived from Jcl:ICR, F1 Hybrids, and MCH (F1 × F1)
|
Marker Loci |
|||||||||||||||||||
|
Strain name |
Idh1 |
Pep3 |
Akp1 |
Hc |
Car2 |
Mup1 |
Gpd1 |
Pgm1 |
Ldr1 |
Gpil |
Hbb |
Es1 |
Es2 |
Thy1 |
Mod1 |
Trf |
Es3 |
H2k |
H2D |
|
IAI |
a |
b |
b |
1 |
b |
a |
b |
b |
a |
b |
s |
b |
b |
b |
b |
b |
c |
q |
q |
|
IQI |
a |
b |
b |
1 |
b |
a |
b |
a |
b |
b |
d |
b |
b |
a |
b |
b |
c |
q |
b |
|
(IAI×IQI)F1 |
a |
b |
b |
1 |
b |
a |
b |
ab |
ab |
b |
sd |
b |
b |
ab |
b |
b |
c |
q |
qb |
|
IPI ICT |
a |
b |
b |
1 |
b |
a |
b |
b |
b |
a |
d |
b |
b |
b |
b |
b |
c |
q |
b |
|
a |
b |
b |
1 |
b |
a |
b |
a |
a |
a |
s |
b |
b |
b |
b |
b |
c |
q |
b |
|
|
(IPI×ICT)F1 |
a |
b |
b |
1 |
b |
a |
b |
ab |
ab |
a |
sd |
b |
b |
b |
b |
b |
c |
q |
b |
|
MCH(AQ×PC) |
a |
b |
b |
1 |
b |
a |
b |
a |
a |
ab |
s |
b |
b |
ab |
b |
b |
c |
q |
qb |
|
ab |
ab |
sd |
b |
b |
|||||||||||||||
|
b |
b |
d |
|||||||||||||||||
|
Production methods |
Maintenance by sib mating |
Continuous production from 4 inbred strains |
Reproduction only from a colony with no animals introduced from other colonies |
|
Reproduction |
Poor - Good |
Excellent |
Good |
|
Genetic uniformity |
Clear |
Clear |
Unclear |
|
Genetic characters |
Not suitable for toxicity tests Limited range of characters |
Suitable for toxicity tests Fixed range of characters |
Suitable for toxicity tests Irregular variations in range of characters |
|
Genetic reproducibility |
Very good |
Very good |
Good |
|
Microbiological control |
Easy |
Easy |
Difficult |
FIGURE 3 Characteristics of inbred strains, MCH, and closed colony.
CONCLUSION
The problems of closed colonies have recently become evident in connection with ICH. I have attempted to explain that these problems cannot be solved by looking only at closed colonies and that it is necessary to reflect on the laboratory animals currently used in animal experiments. Finally, from an overall evaluation of closed colony problems, it is clear that these problems are not problems of the animals themselves but problems related to utilization and production, that is, human problems. When these problems have been solved, it should be possible to select appropriate laboratory animals for each animal experiment.
I propose the following recommendations for solving problems related to animal experimentation. For researchers or users: Understanding (1) the limitations of closed colonies in animal experimentation (what types of experiments are appropriate and what types are inappropriate), and (2) the effects of strain differ
ences in some experiments. For breeders: Understanding (1) the reasons quality is required for laboratory animals, (2) the necessity of using objective scientific evidence to evaluate colonies, and (3) which procedures to use to confirm the genetic quality of laboratory animals.
Finally, in animal experiments (especially safety studies) using closed colonies, I recommend collecting and preserving DNA or biochemical marker data for tests on animals (mice and rats) used in safety studies in the development process of a new drug. If the results obtained in a safety study show significant differences from those obtained in a previous study or from other institutions, the causes can be narrowed down to the following: techniques (such as method of administration), environment (such as food, including temperature and humidity), and/or animals (genetic differences between the colonies used in the studies). Techniques and environment are specified in the GLP, as mentioned; but for animals used in the studies, only the strain names remain. It is important to know the genetic composition of the individuals that comprise the colonies used in the studies if the data obtained are to be utilized effectively. I therefore recommend that the genetic test data obtained for the animals used in such studies be preserved for future reference.
REFERENCES
Katoh, H., S. Wakana, M. Ebukuro, and T. Nomura. 1998. Existence of outbred substocks demonstrated using genetic monitoring system. Rat Genome 4:120-125.
Nomura, T., K. Esaki, and T. Tomita, eds. 1984. ICLAS Manual for Genetic Monitoring of Inbred Mice. Tokyo: University of Tokyo Press.