B
Methodologic Details of Analyses to Evaluate Feasibility of Class Approach and to Define Subclasses
In Chapter 3, the committee presented a reproducible, multifaceted approach for developing subclasses of nonpolymeric, additive organohalogen flame retardants (OFRs) that considered function, chemical structure, and predicted biological activity. The main steps used by the committee can be summarized as (1) identifying and characterizing a “seed” set of chemicals as a working inventory of the class; (2) generating an “expanded” set of chemical analogues to the seed set on the basis of combined functional, structural, and predicted bioactivity information; (3) evaluating the cheminformatic similarity of the seed set to the analogues to evaluate whether the OFRs are distinguishable as a single class; and (4) defining subclasses for hazard evaluation. The goal of that modular approach was to assess the empirical justification for treating a given set of chemicals as a “class” to be evaluated en masse or as a set of subclasses to be evaluated in groups. The committee followed those steps to develop a scientifically sound approach that, although based on the collected seed set of OFRs, is independent of the number of initial structures and thus repeatable and can be applied to future lists of chemicals that might be used as OFRs. This appendix describes the methods used to develop the committee’s classification scheme and is organized according to steps described above.
STEP 1. IDENTIFY AND CHARACTERIZE A “SEED” SET OF CHEMICALS
A list of chemicals that have been used as OFRs was compiled from various documents that list OFRs (Table B-1). The sources were Eastmond (2015), the Environmental Protection Agency of Denmark (Danish EPA 2016), the Environment Agency of the United Kingdom (2003), the International Programme on Chemical Safety (IPCS 1997), the European Food Safety Authority (EFSA 2010, 2011a,b,c, 2012a,b), the Consumer Product Safety Commission (TERA 2016), and the US Environmental Protection Agency (EPA 2015). In the present analysis, 161 chemicals that have been used as “flame retardants” were identified. Despite the fair amount of overlap between sources, only a few (<20) chemicals were listed in all sources; this suggests substantial heterogeneity in the chemical space identified by various groups. The committee emphasizes that this inventory is not necessarily comprehensive, and additional OFRs in commerce might exist.
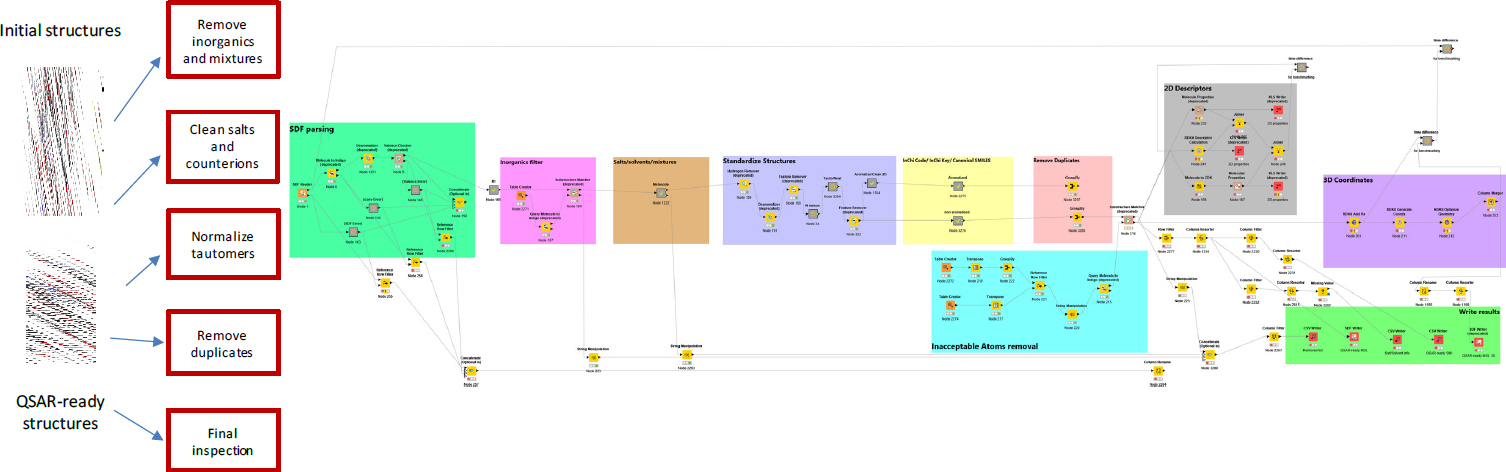
To define the chemical space of OFRs (known OFRs and structural analogues), the committee initially examined and curated the structures of the chemicals identified in the collected inventory of OFRs. First, the names, CAS numbers, and structures in the initial list of 161 OFRs were verified and checked for consistency by using the US Environmental Protection Agency (EPA) Dashboard and other sources. Then, the chemical structures were normalized and deduplicated to generate quantitative structure–activity relationship (QSAR)-ready structures by using a standardization workflow described in Mansouri et al. (2016a,b) and developed in the Konstanz Information Miner (KNIME), an open-source data analytics, reporting, and integration platform (Berthold et al. 2009). Figure B-1 shows the QSAR-ready standardization workflow process.
Two entries (CAS numbers: 52907-07-0 and 13654-09-6) were identified as mixtures and were not included in the present analyses. An additional 11 entries (CAS numbers: 32534-81-9, 59447-55-1, 855992-98-2, 3194-55-6/CMS201444, 3194-55-6/CMS201445, 3194-55-6/CMS2 02190, 3194-55-6/CMS220023, 3194-55-6/CMS220024, 3194-55-6/CMS220025, 3194-55-6/CMS220026, and 31 94-55-6/CMS220027) were considered duplicate structures.
That process resulted in 148 unique chemical structures in the OFR seed set (see list at OFR_QSARready_120318.sdf). The chemical space of the curated set of 148 seed chemicals was characterized by generating QSAR predictions with an open-source application called OPEn structure–activity/property Relationship App (OPERA) (v2.0) (Mansouri et al. 2016b, 2018), which is freely available from the National Institute of Environ-
TABLE B-1 Sources Used to Identify Chemicals in the OFR Inventorya
| Data Sources | Description | No. Chemicals | Year |
|---|---|---|---|
| Eastmond | Flame retardants | 90 | 2015 |
| Danish EPA | Brominated flame retardants | 66 | 2016 |
| UK ENV | Flame retardants | 68 | 2003 |
| IPCS | Flame retardants | 61 | 1997 |
| EFSA | Brominated flame retardants in food | 79 | 2010-2012 |
| CPSC | OFR exposure assessment | 5 | 2016 |
| EPA | Flame retardants | 9 | 2015 |
| OFR Inventory | 161a | — |
aSome chemicals appear in more than one inventory. See OFR_Categories_OFR-inventory_02-15-2019.xlsx. File available at www.nap.edu/25412.
Abbreviations: CPSC, Consumer Product Safety Commission; Danish EPA, Environmental Protection Agency of Denmark; EFSA, European Food Safety Authority; EPA, US Environmental Protection Agency; IPCS, International Programme on Chemical Safety; UK ENV, UK Environment Agency.
mental Health Sciences Github1 and EPA’s CompTox Dashboard.2 The predictions considered physicochemical properties, environmental fate, and toxicity end points, including estrogen and androgen receptor activities (Mansouri et al. 2016a, 2017), and acute oral toxicity (Kleinstreuer et al. 2018).3 In addition to predictions, OPERA provides applicability domains and accuracy estimates for each prediction. More information about OPERA outputs can be found on the EPA CompTox Dashboard and OPERA’s QSAR model reporting format (QMRF) reports that are registered and validated by the European Commission’s Joint Research Center to be OECD-compliant.4
STEP 2. GENERATE AN “EXPANDED” SET OF CHEMICAL ANALOGUES ON THE BASIS OF COMBINED FUNCTIONAL, STRUCTURAL, AND PREDICTED BIOACTIVITY INFORMATION
To identify the analogues that are structurally similar to the OFR seed set, the committee developed an automated KNIME workflow to identify all organohalogens (about 200,000 structures) from the EPA Distributed Structure-Searchable Toxicity (DSSTox) database (Richard 2004; Richard et al. 2006) (Figure B-2).
The chemicals were then compared with the 148 OFR seed structures to determine the list of most similar OFR analogues. The chemistry-development kit (CDK) fingerprints were used with a Tanimoto similarity-index threshold of 80% (Steinbeck et al. 2003, 2006). That step resulted in an output of 1,073 analogues, which were then analyzed by using the QSAR-ready standardization workflow shown in Figure B-25 (Mansouri et al. 2016a,b).
The final set of analogue compounds used for comparative analysis (the expanded set) consisted of 1,073 unique structures.6 Procedures described earlier were then repeated on the expanded set of OFR analogues, including collection of data by running OPERA predictions7 and retrieval of data on these chemicals from the EPA Dashboard.8
STEP 3: EVALUATE THE CHEMINFORMATIC SIMILARITY OF THE SEED SET TO THE ANALOGUES TO EVALUATE WHETHER THE OFRs ARE DISTINGUISHABLE AS A SINGLE CLASS
Comparing OFR Seed Set to Analogues: Unsupervised Analyses
Two unsupervised methods were used to compare features of the OFR seed set with those of the expanded
___________________
1 See https://github.com/NIEHS/OPERA.
2 See https://comptox.epa.gov/dashboard.
3 See: prediction file: pred_OPERA_OFR.csv and list of OPERA models: OPERA2.0_models.xlsx and OFRs.xlsx. Files available at www.nap.edu/25412.
4 See https://qsardb.jrc.ec.europa.eu/qmrf/.
5 See OFR.knwf. File available at www.nap.edu/25412.
6 See OFR_Analogs_QSAR-ready_120318.sdf. File available at www.nap.edu/25412.
7 See pred_OPERA_OFR_Analogs_120318.csv. File available at www.nap.edu/25412.
8 See OFR_Analogs_ChemistryDashboard-Batch-Search_201812-03_17_05_44.xls. File available at www.nap.edu/25412.
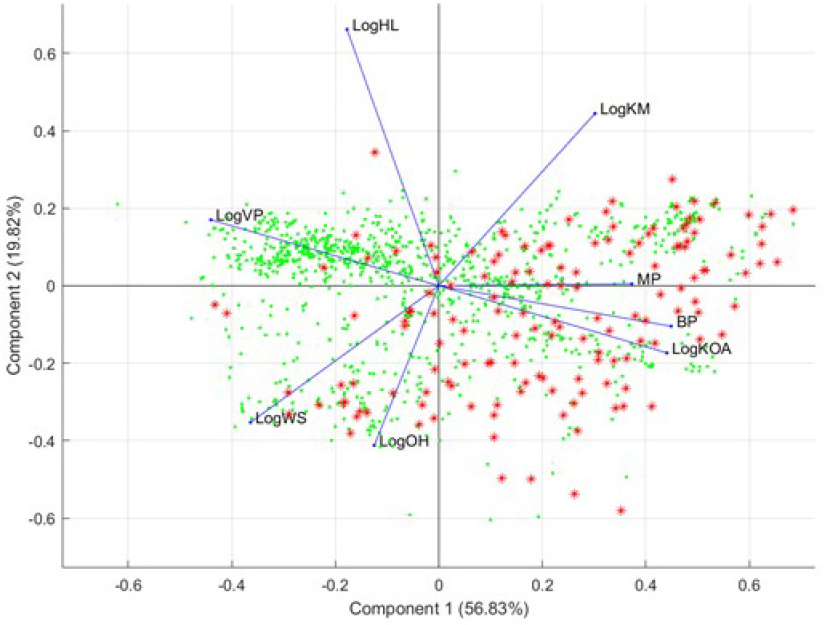
set (analogues): principal component analysis (PCA) of OPERA physicochemical properties and ToxPrint Chemotype Enrichment. PCA is a statistical procedure that uses a number of orthogonal transformations to convert a set of observations of possibly correlated variables into a set of new uncorrelated variables called principal components. The transformations are defined in such a way that the first principal components encode the largest possible variance (condensing the information on the original variables). The results of a PCA are usually discussed in terms of component loadings and scores; the loadings represent the weights of the original variables required to get the scores of the components (Pearson 1901). ToxPrint is a publicly available database of over 700 chemotypes that were developed from a chemical space on which some toxicity data were available (Yang et al. 2015). Chemotypes represent structural fragments expressed with imbedded atomic, electronic, and physicochemical properties. A chemotype can be used to represent a generic query feature (for example, chain:alkene) or a feature informed by biologic activity or as a component of an SAR model as described in this section. Most of the chemotypes in ToxPrint are not structural alerts; rather, more than 90% of them represent generic query features that can help to search and cluster substructures.
PCA of OPERA physicochemical properties. Figure B-3 shows the samples (OFRs) plotted on the basis of their coordinates (scores) for the first two components with the highest accumulated variance among all components encoding the information present in the 45 original OPERA variables. The lengths of the blue lines (loadings) show the importance of each of the 45 OPERA variables. The directions in which the chemicals (OFRs shown as red stars and analogues shown as green dots) are plotted show the similarity according to each specific variable.9 However, because PCA is a multivariate analysis, the PCA plot shows the interactions and the similarities between the samples (chemicals) in a multivariate fashion. Apart from the cluster of mostly volatile analogue structures that have different vapor-pressure properties, the OFR seed set and the expanded set have similar OPERA-predicted physicochemical, toxicologic, and environmental-fate properties. Also, the first two components encoded less than 50% of the variance, and this indicates the high variability within the properties of the analyzed structures.
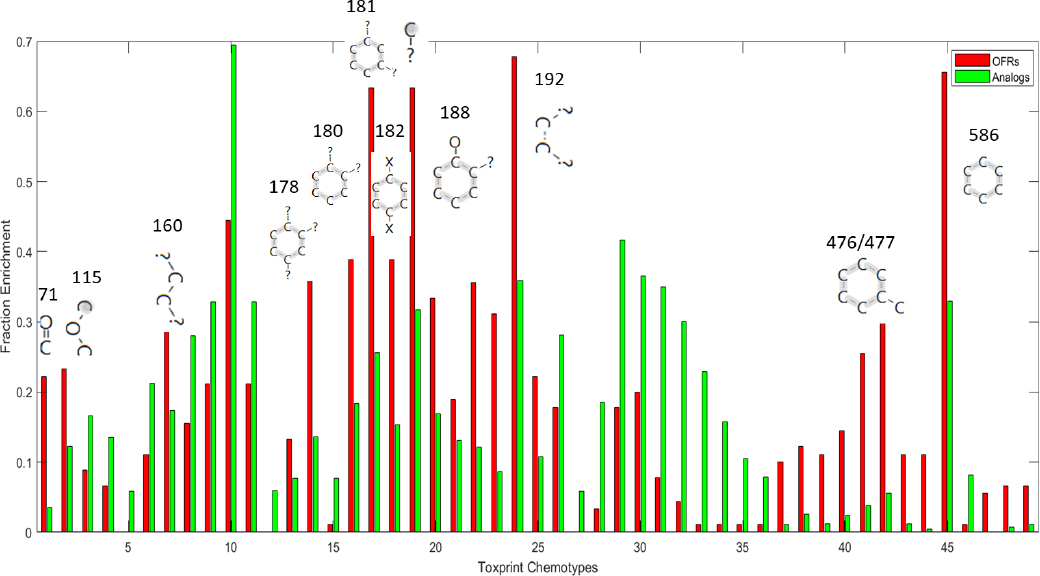
ToxPrint Chemotype Enrichment. A second unsupervised analysis was performed by using Toxprint Chemotype Enrichment (Richard et al. 2016) to identify the most enriched Toxprint Chemotypes in the OFR seed and expanded sets. The OFR seed set was found to consist mostly of aromatic chemicals. The analogues contained various types of chemical structures, both aromatic and aliphatic enrichment sites (Figure B-4). Furthermore, sev-
___________________
9 List of abbreviations provided in the supplemental material OPERA2.0_models.xlsx. File available at www.nap.edu/25412.
eral ToxPrint chemotypes are highly enriched in both the seed list and the analogues.
Comparing OFR Seed Set with Analogues: Supervised Analyses
The goal of supervised analyses was to determine whether a selected set of properties can differentiate the OFR seed set from the analogues accurately. Two approaches, Machine Learning Classification and Supervised PCA, were used.
Machine Learning Classification. The well-known, reliable k-nearest-neighbors (kNN) coupled to genetic algorithms (GAs) were used to find the optimal subset of molecular descriptors that differentiate the OFR seed set and analogues (Todeschini 1989). GA begins with an initial random population of chromosomes, which are binary vectors that represent the presence or absence of molecular descriptors. An evolutionary process is then simulated to optimize a defined fitness function, and new chromosomes are obtained by coupling the chromosomes of the initial population with genetic operations, such as crossover and mutation (Ballabio et al. 2011; Leardi and González 1998). Classification-balanced accuracy (BA) was used as the fitness function and calculated in a fivefold cross-validation procedure. To adapt that method for the present purpose, the committee used OPERA properties and CDK descriptors in the model. Results of the analyses show that the variable selection techniques were able to classify the OFR set and the analogue chemicals with up to 80% balanced accuracy when the highest selected descriptors were used.
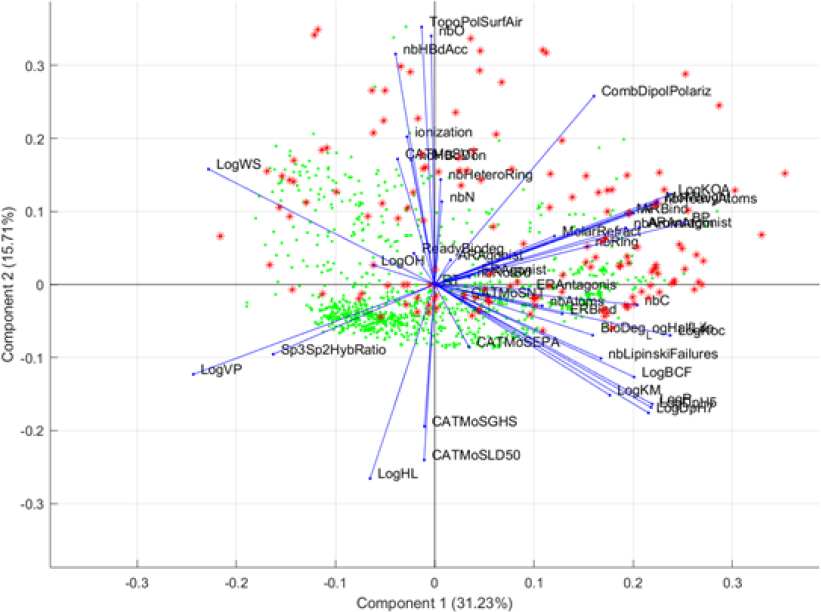
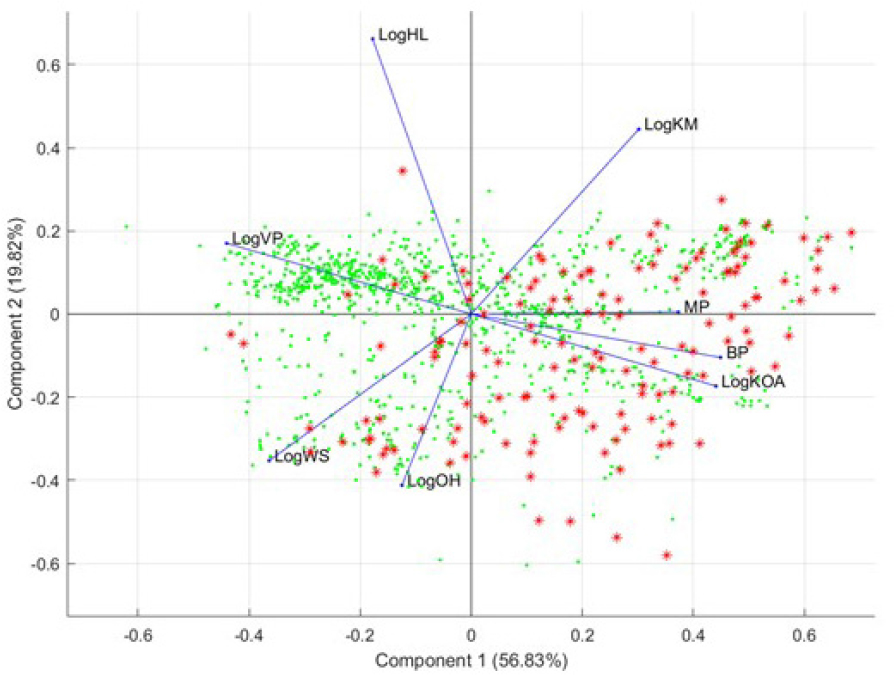
Supervised PCA. Following the supervised GA learning techniques, the committee used results on the descriptors and conducted an additional supervised PCA analysis. Specifically, the highest selected descriptors were used in this supervised PCA analysis to judge visually whether these descriptors were able to differentiate the two groups of chemicals. Figure B-5 shows the output from this supervised PCA analysis.
Results of this PCA show that the highest selected descriptors separate analogue chemicals (expanded set) from the OFR seed set; this explains the high balanced accuracy that was reached in the GA-kNN procedure. However, most of the analogue compounds are separated by vapor pressure (LogVP), soil adsorption (LogKM), and water solubility (LogWS). That finding demonstrates that analogues that easily separate from the OFR seed set do not share physicochemical and environmental properties with the chemicals in the seed set. However, the fact that many analogues are not easily separable from the seed set chemicals suggests that the analogues share properties with the OFR seed set.
Conclusion
The analyses show that many existing organohalogen chemicals, although never used as flame retardants, share properties with the known OFRs and might have the potential of being used for the same purpose. Thus, the seed list of OFR chemicals can be separated only on the basis of use category and cannot be considered to be a scientifically defined class by itself on the basis of the structural features or on chemical properties. Consequently, to study the toxicologic and biologic properties, one can categorize the OFRs into subclasses that share functional groups.
STEP 4: DEFINE SUBCLASSES FOR HAZARD EVALUATION
The chemical space of the OFR inventory was profiled in more detail to identify major chemical groups. ToxPrint Chemotypes (Yang et al. 2015; Richard et al. 2016), available in ChemoTyper,10 were used initially to characterize the 148 structures in the seed set. The full inventory of 161 chemicals was covered by 143 (of 729 possible) chemotypes that are available in ToxPrint.
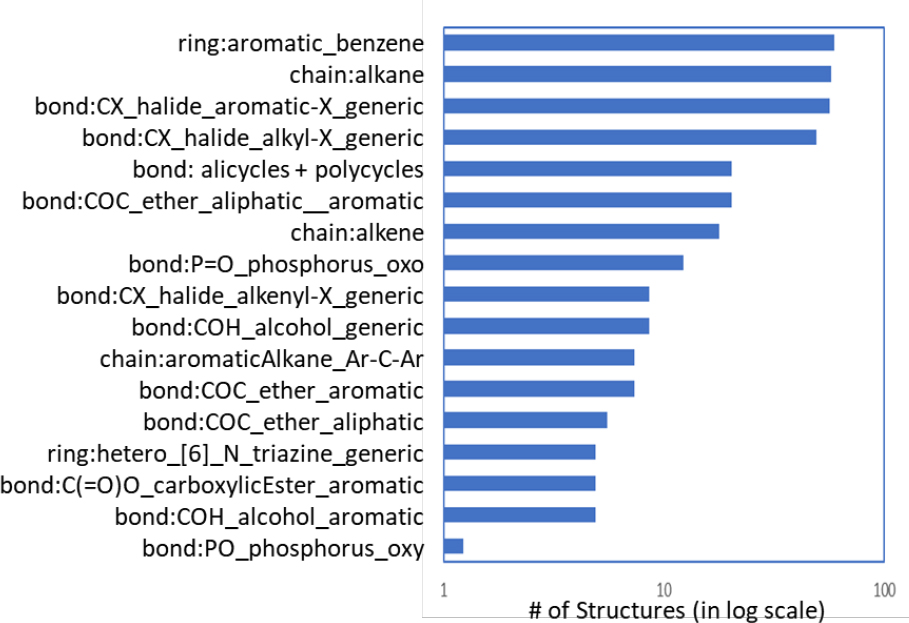
Some of the major ToxPrint chemotypes encountered in the OFR inventory in this study are listed in Figure B-6. These chemotypes roughly identify the top-level, large categories within the OFR chemical space. They include polyhalogenated analogues of benzenes, aliphatic chains, alicycles and carbocycles (polycycles), organophosphorus chemicals, aromatic and aliphatic ethers, phenols and their derivatives, and aromatic carboxylic esters (phthalates). These observations suggest a subclass approach based on the generic ToxPrint features as chemical classes.
The committee’s approach was to develop large commonly occurring core features of the OFR chemicals to represent each group that can also be associated with a predicted biologic activity whenever possible. Using expert judgment, the committee grouped the OFR inventory into eight structural classes on the basis of predicted biologic activity (such as GABA receptors, aromatase activity, and ER/AR modulators) (see Table 3-1). Merging the biology-informed groups with the chemotypes identified in Figure B-6 led to the formulation of 14 OFR categories for the inventory of 161 OR chemicals (Table B-2). Seven OFR chemicals were placed into two subclasses on the basis of their predicted biologic activity and chemical structures. The OFR subclasses were used to support the case studies described in Chapter 3.
___________________
10 See http://chemotyper.org.
| OFR Subclass | No. Chemicals | CAS No. of Chemicals |
|---|---|---|
| Polyhalogenated alicycles | 17 | 25495-98-1; 25637-99-4; 3194-55-6; 3194-57-8; 134237-50-6; 134237-51-7; 134237-52-8; 678970-17-7; 678970-16-6; 678970-15-5; 169102-57-2; 138257-19-9; 138257-18-8; 3322-93-8; 77-47-4; 87-84-3; 1837-91-8 |
| Polyhalogenated aliphatic carboxylate | 4 | 3066-70-4; 5445-17-0; 5445-19-2; 19660-16-3 |
| Polyhalogenated aliphatic chains | 12 | 52434-59-0; 1522-92-5; 3296-90-0; 3234-02-4; 96-13-9; 109678-33-3; 85535-84-8; 71011-12-6; 85535-85-9; 63449-39-8; 75-95-6; 79-27-6 |
| Polyhalogenated benzene alicycles | 4 | 1084889-51-9; 893843-07-7; 1025956-65-3; 155613-93-7 |
| Polyhalogenated benzene aliphatics and functionalized | 19 | 168434-45-5; 23488-38-2; 39569-21-6; 87-83-2; 85-22-3; 38521-51-6; 58495-09-3; 31780-26-4; 84852-53-9; 497107-13-8; 59447-55-1; 34571-16-9*; 855993-01-0*; 855992-98-2*; 147-82-0; 57011-47-9; 61368-34-1; 93-52-7; 39568-99-5 |
| Polyhalogenated benzenes | 19 | 608-90-2; 87-82-1; 84303-46-8; 60044-26-0; 67733-52-2; 67889-00-3; 69278-62-2; 59080-40-9; 13654-09-6; 36355-01-8; 92-66-0; 92-86-4; 115245-07-3; 60044-24-8; 59080-37-4; 77102-82-0; 16400-50-3; 67888-96-4; 59080-39-6 |
| Polyhalogenated bisphenol aliphatics and functionalized | 11 | 66710-97-2; 55205-38-4; 37853-61-5; 37419-42-4; 3072-84-2; 33798-02-6; 79-94-7; 25327-89-3; 21850-44-2; 4162-45-2; 79-95-8 |
| Polyhalogenated carbocycles | 15 | 13560-89-9; 51936-55-1; 13560-92-4; 34571-16-9*; 855993-01-0*; 855992-98-2*; 2385-85-5; 18300-04-4; 115-28-6; 1773-89-3; 1770-80-5; 115-27-5; 31107-44-5; 40703-79-5; 52907-07-0 |
| Polyhalogenated diphenyl ethers | 12 | 1163-19-5; 32534-81-9; 60348-60-9; 32536-52-0; 58965-66-5; 5436-43-1; 207122-16-5; 189084-67-1; 41318-75-6; 189084-64-8; 68631-49-2; 207122-15-4 |
| Polyhalogenated organophosphates | 22 | 114955-21-4*; 1373346-90-7*; 126-72-7; 19186-97-1; 115-96-8; 13674-84-5; 13674-87-8; 38051-10-4; 66108-37-0; 78-43-3; 6145-73-9; 33125-86-9; 49690-63-3; 7046-64-2; 5412-25-9; 53461-82-8; 61090-89-9; 140-08-9; 6749-73-1; 4351-70-6; 6294-34-4; 115-98-0 |
| Polyhalogenated phenol derivatives | 7 | 118-79-6; 608-71-9; 615-58-7; 42757-55-1; 39635-79-5; 70156-79-5; 25713-60-4* |
| Polyhalogenated phenol-aliphatic ethers | 9 | 3278-89-5; 31977-87-4; 35109-60-5; 37853-59-1; 61262-53-1; 3555-11-1; 607-99-8; 26762-91-4; 20217-01-0 |
| Polyhalogenated phthalates/benzoates/imides | 11 | 32588-76-4; 183658-27-7; 90075-91-5; 82001-21-6; 20566-35-2; 26040-51-7; 7415-86-3; 55481-60-2; 632-79-1; 117-08-8; 57011-47-9 |
| Polyhalogenated triazines | 6 | 52434-90-9; 57829-89-7; 75795-16-3; 25713-60-4*; 114955-21-4*; 1373346-90-7* |
*An asterisk indicates that the chemical occurs in more than one category.
The categories were confirmed by comparison with unsupervised clustering techniques. The committee used 148 unique structures from the OFR seed set and applied an agglomerative nesting (clustering) procedure that used ToxPrint Chemotypes as the independent variables. ToxPrint Chemotypes that were found in at least two structures were then used to generate a distance matrix by calculating Jaccard distances (distances between clusters were calculated by using the average linkage method). The cluster-analysis results are in good agreement with the OFR chemotype categories that the committee generated.
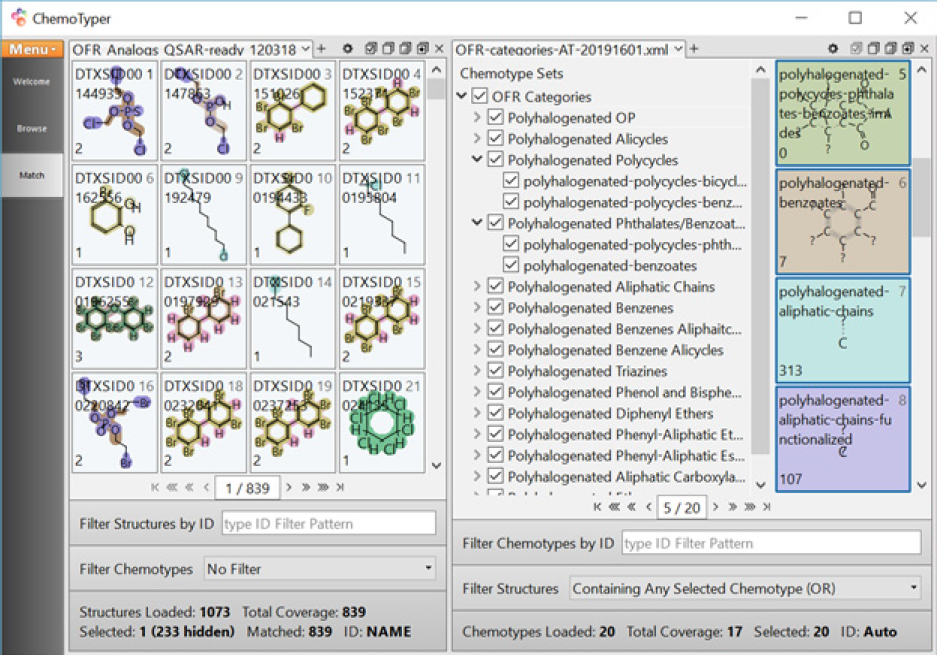
To validate whether the OFR categories designed with the committee’s approach can group chemicals of interest adequately, the 14 OFR categories were coded in CSRML11 and implemented with the public ChemoTyper application. Figure B-7 shows the current OFR-categories-20192501.xml12 in ChemoTyper to search against the “expanded set (1,073)” with highlighting to show structural features matched with the OFR categories.13
The organohalogen structures that are not covered by the OFR categories include organohalides with only one halogen, cyclic rings smaller than five, spiro polycyclic rings, halogens directly attached to phosphorus, or trivalent phosphorus. In accord with the supervised analyses in the preceding section, the chemical space covered by the expanded set is similar to that in the seed set in that the OFR categories cover nearly 80% of the set of 1,073.
REFERENCES
Ballabio, D., M. Vasighi, V. Consonni, and M. Kompany-Zareh. 2011. Genetic algorithms for architecture optimisation of counter-propagation artificial neural networks. Chemometr. Intell. Lab. Syst. 105(1):56-64.
Berthold, M.R., N. Cebron, F. Dill, T.R. Gabriel, T. Kötter, T. Meinl, P. Ohl, K. Thiel, B. Wiswedel. 2009. KNIME – The Konstanz Information Miner, Version 2.0 and beyond. ACM SIGKDD Explorations Newsletter. 11(1):26-31.
Danish EPA (Environmental Protection Agency, Denmark). 2016. Category approach for selected brominated flame retardants: Preliminary structural grouping of brominated flame retardants. Environmental project No. 1872. Copenhagen, Denmark: Danish Environmental Protection Agency [online]. Available: https://www2.mst.dk/Udgiv/publications/2016/07/978-87-93435-90-2.pdf [accessed September 25, 2018].
Eastmond, D.A. 2015. Letter to Consumer Product Safety Commission (CPSC). Exhibit C in CPSC staff briefing package in response to petition HP15-1, requesting rule-
___________________
11 See OFR-categories-20192501.xml. File available at www.nap.edu/25412.
12 File available at www.nap.edu/25412.
13 Downloadable from https://chemotyper.org.
making on certain products containing organohalogen flame retardants. May 24, 2017. Available: https://www.cpsc.gov/content/ballot-vote-petition-hp-15-1-requesting-rulemaking-on-certain-products-containing [accessed July 18, 2018].
EFSA (European Food Safety Authority). 2010. Scientific opinion on polybrominated biphenyls (PBBs) in food. EFSA Journal 8(10):1789 [online]. Available: www.efsa.europa.eu/efsajournal [accessed September 25, 2018].
EFSA. 2011a. Scientific opinion on tetrabromobisphenol A (TBBPA) and its derivatives in food. EFSA Journal 9(12):2477 (updated 2013) [online]. Available: www.efsa.europa.eu/efsajournal [accessed September 25, 2018].
EFSA. 2011b. Scientific opinion on polybrominated diphenyl ethers (PBDEs) in food. EFSA Journal 9(5):2156 [online]. Available: www.efsa.europa.eu/efsajournal [accessed September 25, 2018].
EFSA. 2011c. Scientific opinion on hexabromocyclododecanes (HBCDDs) in food. EFSA Journal 9(7):2296 [online]. Available: www.efsa.europa.eu/efsajournal [accessed September 25, 2018].
EFSA. 2012a. Scientific opinion on brominated flame retardants (BFRs) in food: Brominated phenols and their derivatives. EFSA Journal 10(4):2634 [online]. Available: www.efsa.europa.eu/efsajournal [accessed September 25, 2018].
EFSA. 2012b. Scientific opinion on emerging and novel brominated flame retardants (BFRs) in food. EFSA Journal 10(10):2908 [online]. Available: www.efsa.europa.eu/efsajournal [accessed September 25, 2018].
Environment Agency. 2003. Prioritisation of flame retardants for environmental risk assessment. United Kingdom: Environment Agency [online]. Available: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/291681/scho1008bote-e-e.pdf [accessed October 5, 2018].
EPA (US Environmental Protection Agency). 2015. Flame retardants used in flexible polyurethane foam: An alternatives assessment update. EPA 744-R-15-002. Design for the Environment, US EPA. Available: https://www.epa.gov/saferchoice/2015-update-report-flame-retardants-used-flexible-polyurethane-foam-publications [accessed February 21, 2019].
IPCS. 1997. Flame retardants: A general introduction. Environmental Health Criteria 192. Geneva, Switzerland: World Health Organization [online]. Available: http://www.inchem.org/documents/ehc/ehc/ehc192.htm [accessed October 5, 2018].
Kleinstreuer, N.C., A. Karmaus, K. Mansouri, D.G. Allen, J.M. Fitzpatrick, and G. Patlewicz. 2018. Predictive models for acute oral systemic toxicity: A workshop to bridge the gap from research to regulation. Comput. Toxicol. 8(11):21-24.
Leardi, R., and A. L. González. 1998. Genetic algorithms applied to feature selection in PLS regression: how and when to use them. Chemometr. Intell. Lab. Syst. 41(2):195-207
Mansouri, K., A. Abdelaziz, A. Rybacka, A. Roncaglioni, A. Tropsha, A. Varnek, A. Zakharov, A. Worth, A.M. Richard, C.M. Grulke, D. Trisciuzzi, D. Fourches, D. Horvath, E. Benfenati, E. Muratov, E.B. Wedebye, F. Grisoni, G.F. Mangiatordi, G.M. Incisivo, H. Hong, H.W. Ng, I.V. Tetko, I. Balabin, J. Kancherla, J. Shen, J. Burton, M. Nicklaus, M. Cassotti, N.G. Nikolov, O. Nicolotti, P.L. Andersson, Q. Zang, R. Politi, R.D. Beger, R. Todeschini, R. Huang, S. Farag, S.A. Rosenberg, S. Slavov, X. Hu, and R.S. Judson. 2016a. CERAPP: Collaborative Estrogen Receptor Activity Prediction Project. Environ. Health Perspect. 24(7):1023-1033.
Mansouri, K., C.M. Grulke, A.M. Richard, R.S. Judson, and A.J. Williams. 2016b. An automated curation procedure for addressing chemical errors and inconsistencies in public datasets used in QSAR modelling. SAR QSAR Environ. Res. 27(11):939-965.
Mansouri, K., N. Kleinstreuer, E. Watt, J. Harris, and R. Judson. 2017. CoMPARA: Collaborative Modeling Project for Androgen Receptor Activity. Poster presented at Society of Toxicology Annual Meeting, Baltimore, MD, March 12-16, 2017. doi:10.13140/RG.2.2.16791.78241.
Mansouri, K., C.M. Grulke, R.S. Judson, and A.J. Williams. 2018. OPERA models for predicting physicochemical properties and environmental fate endpoints. J. Cheminform. 10(1):10.
Pearson, K. 1901. On lines and planes of closest fit to systems of points in space. The London, Edinburgh, and Dublin Philosophical Magazine and Journal of Science 2(11):559-572.
Richard, A.M. 2004. DSSTox Website launch: improving public access to databases for building structure-toxicity prediction models. Preclinica 2:103-108.
Richard, A.M., L.S. Gold, and M.C. Nicklaus. 2006. Chemical structure indexing of toxicity data on the internet: moving toward a flat world. Curr. Opin. Drug Discov. Devel. 9(3):314-325.
Richard, A.M., R.S. Judson, K.A. Houck, C.M. Grulke, P. Volarath, I. Thillainadarajah, C. Yang, J. Rathman, M.T. Martin, J.F. Wambaugh, T.B. Knudsen, J. Kancherla, K. Mansouri, G. Patlewicz, A.J. Williams, S.B. Little, K.M. Crofton, and R.S. Thomas. 2016. ToxCast chemical landscape: Paving the road to 21st century toxicology. Chem. Res. Toxicol. 29(8):1225-1251.
Steinbeck, C., Y. Han, S. Kuhn, O. Horlacher, E. Luttmann, and E. Willighagen. 2003. The Chemistry Development Kit (CDK): An open-source Java library for chemo- and bioinformatics. J. Chem. Inf. Comput. Sci. 43:493-500.
Steinbeck, C., C. Hoppe, S. Kuhn, M. Floris, R. Guha, and E.L. Willighagen. 2006. Recent developments of the chemistry development kit (CDK) – an open-source java library for chemo- and bioinformatics. Curr. Pharm. Des. 12(17):2111-2120.
TERA (Toxicology Excellence for Risk Assessment). 2016. Flame Retardant Exposure Assessment Database. Conducted in conjunction with Life Line Group under a contract with the US Consumer Project Safety Commission, Task Order 15, Contract Number CPSC-D-12-0001 [online]. Available: https://www.cpsc.gov/s3fs-public/FR-exposure-assessment-contractor-report-18-09282016-with-cover.pdf?miFGGrplONrzWVCDOIQSOY1lTeGzQvWi [accessed September 25, 2018.]
Todeschini, R. 1989. K-nearest neighbour method: the influence of data transformations and metrics. Chemom Intell Lab Syst 6:213-220.
Yang, C., A. Tarkhov, J. Marusczyk, B. Bienfait, J. Gasteiger, T. Kleinoder, T. Magdziarz, O. Sacher, C.H. Schwab, J. Schwoebel, L. Terfloth, K. Arvidson, A. Richard, A. Worth, and J. Rathman. 2015. New publicly available chemical query language, CSRML, to support chemotype representations for application to data mining and modeling. J. Chem. Inf. Model 55(3):510-528.











