Subreport
Methods and Mechanisms of Genetic Manipulation and Cloning of Animals
As part of its charge, the committee was asked to prepare a subreport evaluating methods for detecting potential unintended compositional changes across the spectrum of messenger ribonucleic acid (mRNA), proteins, metabolites and nutrients that may occur in food derived from cloned animals that have not been genetically modified via genetic engineering methods. Detailed descriptions of methods used in animal cloning and biotechnology are provided in the report Animal Biotechnology: Science-Based Concerns (NRC, 2002). In addition, the committee was charged with evaluating methods to detect potential, unintended, adverse health effects of foods derived from cloned animals.
INTRODUCTION
Since the onset of modern biotechnology, scientists have made discoveries leading to the development of new techniques for animal agriculture. Applications of biotechnology to animal agriculture include improving milk production and composition; increasing growth rate of meat animals; improving productive efficiency, or gain-to-feed ratios, and carcass composition; increasing disease resistance; enhancing reproductive performance; increasing prolificacy; and altering cell and tissue characteristics for biomedical research and manufacturing. Continued development of new biotechnologies also will allow farm animals to serve as sources of both biopharmaceuticals for human medicine and organs for transplantation.
ANIMAL BIOTECHNOLOGY
A number of animal biotechnologies have already been developed and are in commercial use. One such example is recombinant-derived bovine somatotropin (bST). Recombinant bST has been approved by the Food and Drug Administration for use in the U.S. dairy industry and is also approved for use by 18 other countries (CAST, 2003; Etherton and Bauman, 1998). Commercial use in the United States began in early 1994 and has increased to the point that about one-half of all U.S. dairy herds, comprising more than 3 million cows, are receiving bST (Bauman, 1999). Milk yield increases in response to bST typically range from 10 to 15 percent (about 4–6 kg/d), although larger increases may occur when the management and care of the animals are excellent (Bauman, 1992; Chillard, 1989; NRC, 1994). It is, however, important to distinguish the use of bST from other biotechnologies, such as transgenic or cloned animals. Application of recombinant bST is a biotechnology in which a recombinant-derived protein is administered by injection to the recipient animal without changing the animal’s genetic composition or genome.
The application of genomics—the study of how the genes in deoxyribonucleic acid (DNA) are organized and expressed—and bioinformatics in animal agriculture will provide new genetic markers for improved selection for desired traits in all livestock species. Transgenic biology provides a means of altering animal genomes to achieve desired production and health outcomes of commercial value and societal importance. For example, genetic modification of animals may lead to technologies that reduce the major losses that occur during the first months of embryogenesis. Biotechnology also offers potential to animal agriculture as a means to reduce nutrients and odors from manure and volume of manure produced, resulting in animals that are more environmentally friendly (CAST, 2003).
The advent of techniques to propagate animals by nuclear transfer, also known as cloning, potentially offers many important applications to animal agriculture, including reproducing highly desired elite sires and dams. Animals selected for cloning will be of great value because of their increased genetic merit for increased food production, disease resistance, and reproductive efficiency, or will be valued because they have been genetically modified to produce organs for transplantation or products with biomedical application.
Before entering the marketplace, new agricultural biotechnologies are evaluated rigorously by the appropriate federal regulatory agencies to ensure efficacy, consumer safety, and animal health and well-being. The development of technologies to clone animals used for food production has raised the question of whether there are unintended compositional changes in food derived from these animals that may, in turn, result in unintended health effects.
CLONING
Cloning, a term originally used primarily in horticulture to describe asexually produced progeny, means to make a copy of an individual organism, or in cellular and molecular biology, groups of identical cells and replicas of DNA and other molecules. For example, monozygotic twins are clones. Although clone is descriptive for multiple approaches for cloning animals, in this report clone is used as a descriptor for somatic cell nuclear transfer.
Animal cloning during the late 1980s resulted from the transfer of nuclei from blastomeres of early cleavage-stage embryos into enucleated oocytes, and cloning of livestock and laboratory animals has resulted from transferring a nucleus from a somatic cell into an oocyte from which the nucleus has been removed (Westhusin et al., 2001; Wilmut et al., 1997).
Somatic cell nuclear transfer can also be used to produce undifferentiated embryonic stem cells, which are matched to the recipient for research and therapy that is independent of the reproductive cloning of animals. The progeny from cloning using nuclei from either blastomeres or somatic cells are not exact replicas of an individual animal due to cytoplasmic inheritance of mitochondrial DNA from the donor egg, other cytoplasmic factors that may influence “reprogramming” of the genome of the transferred nucleus, and subsequent development of the cloned organism (Cummins, 2001; Jaenisch and Wilmut, 2001).
Cloning by nuclear transfer from embryonic blastomeres (Willadsen, 1989; Willadsen and Polge, 1981) or from a differentiated cell of an adult (Kuhholzer and Prather, 2000; Polejaeva et al., 2000; Wilmut et al., 1997) requires that the introduced nucleus be reprogrammed by the cytoplasm of the egg and direct development of a new embryo, which is then transferred to a recipient mother for development to term. The offspring will be identical to their siblings and to the original donor animal in terms of their nuclear DNA, but will differ in their mitochondrial genes; variances in the manner nuclear genes are expressed are also possible.
Epigenetic Change in the Genome
Epigenetics is the study of factors that influence behavior of a cell without directly affecting its DNA or other genetic components. The epigenetic view of differentiation is that cells undergo differentiation events that depend on correct temporal and spatial repression, derepression, or activation of genes affecting the fate of cells, tissues, organs, and ultimately, organisms. Thus epigenetic changes in an organism are normal and result in alterations in gene expression. For example, epigenetic transformation of a normal cell to a tumor cell can occur without mutation of any gene.
Effects of Cloning
Santos and colleagues (2003) have reviewed current thinking on epigenetic marking that correlates with developmental potential of cloned bovine preimplantation embryos. They indicate that reprogramming of DNA methylation affects the entire genome of mammalian embryos both during phase I germline development when DNA methylation imprints are eliminated and during phase II preimplantation development of mammalian embryos.
Phase II DNA reprogramming is initiated upon fertilization of the oocyte for remodeling of chromatin in the male pronucleus and selective demethylation of its DNA, while subsequent DNA demethylation in early cleavage stage embryos is passive. At the blastocyst stage, de novo methylation is lineage-specific as the inner cell mass, or embryonic disc, becomes highly methylated and trophectoderm becomes hypomethylated. These epigenetic reprogramming events appear to be deficient in cloned embryos that have abnormal patterns of DNA methylation and gene expression.
Cloning by nuclear transfer using present methods is very inefficient. This is likely due to the limited ability of oocyte cytoplasm to reprogram somatic cell nuclear DNA, whereas it readily reprograms sperm cell nuclear DNA (Wilmut et al., 2002). Consequently, there are very high rates of embryonic, fetal, perinatal, and neonatal deaths, as well as birth of offspring with various abnormalities. These losses are assumed to result from inappropriate expression of genes during various stages of development, with less than 4 percent of embryos reconstructed using adult or fetal somatic cell nuclei being born as live young.
A very high percentage of fertilized eggs that develop and survive beyond the first 30 to 60 days of gestation in sheep and cattle have abnormal placental morphology, such as reduced vascularity and abnormal or few cotyledons; abnormal placental functions, such as hydroallantois and hydroamnios; and abnormal fetal development, such as enlarged liver, hydrops fetalis, dermal hemorrhaging, and swollen brain. Neonates also often experience respiratory distress and cardiovascular abnormalities (De Sousa et al., 2001; Hill et al., 1999, 2000). In addition, there is evidence for abnormalities of the immune system, brain, and digestive system of cloned animals. Mice cloned from cumulus cells become obese, but this trait is not heritable after sexual reproduction, which is direct evidence that obesity in the clones result from epigenetic events (Tamashiro et al., 2002; Wilmut et al., 2002).
Wilmut and colleagues (2002) have reported that a number of cellular factors influence the outcome of cloned offspring. First, normal development depends on the embryo having normal ploidy, which is achieved by coordinating stage of cell cycle of the recipient oocyte and donor nucleus. For donor cells, the G2/M/G1/G0 phases of the cell cycle are associated with effective nuclear reprogramming; however, there are also conflicting results as to whether there is an advantage for donor cell nuclei to be in G1 versus G0.
Donor cells at metaphase seem to be most compatible with metaphase II oocytes with respect to development of embryos to the blastocyst stage, but high rates of mortality occur later. This mortality likely reflects inappropriate expression of genes at different stages of development, which is lethal. For example, perturbation of expression of H10 and/or insulin-like growth factor II (IGF-II), as well as reduced expression of IGF-II receptor in liver, kidney, heart, and muscle is associated with large lamb syndrome following embryo culture.
Changes in expression of the IGF-II receptor gene have been associated with loss of methylation at a differentially methylated region involved with genetic imprinting. There are also reports of improper expression of both imprinted and nonimprinted genes, inappropriate reactivation of the inactive X chromosome (Xue et al., 2002), and reprogramming of telomerase activity (Rideout et al., 2001) to restore telomere length in cloned animals.
Inappropriate gene expression can result from defects in the organization of nuclear material, chromatin structure, and/or activity of regulatory molecules. Chromatin proteins regulate access of transcription factors to chromosomal DNA and expression patterns for individual genes. Linker histone H1 and somatic cell histone H1 are important regulators of gene expression. Their expression changes from absent to very low during early embryonic development to low to moderate at the time of activation of the embryonic genome, and somatic cell histone H1 is lost from most mouse nuclei soon after transfer depending on cell cycle stage for donor and recipient cells. Enzymatic modifications of histones include phosphorylation, methylation, acetylation, and ubiquitination or removal of these modifications, which lead to epigenetic regulation of gene expression.
Expression of genes in higher organisms is regulated by histones H2A, H2B, H3, and H4, as well as histone H1 and other nonhistone proteins, which result in the condensed state DNA that is inaccessible to transcription factors. Transcription of genes requires “unraveling” of DNA and a higher order chromatin structure. This is achieved by a family of adenosine triphosphate-hydrolyzing enzymes that remodel chromatin by shuffling nucleosomes and a family of enzymes that modify histones covalently by acetylation, phosphorylation, methylation, or ubiquination of histone tails. If this does not occur, the genes are silent. For details on histone methylation and gene transcription, refer to Kouzarides (2002).
Widespread disruptions in DNA methylation occur in cloned embryos of mice and cattle, with abnormally high rates of methylation being retained through several cell divisions. In pigs, changes in methylation are initially similar to that for embryos resulting from in vitro fertilization, but by the four- to eight-cell stage, embryonic DNA is more extensively methylated than for embryos resulting from oocytes fertilized in vivo.
Santos and colleagues (2003) note that epigenetic reprogramming is severely deficient in cloned bovine embryos and involves histone H3 lysine 9 (H3-K9). In control bovine embryos, DNA methylation is reduced between the two- and fourcell stages followed by de novo methylation after the eight-cell stage. There is
close correlation between H3-K9 methylation and overall DNA methylation. DNA methylation is up to tenfold greater at all stages of development in cloned bovine embryos, indicating that more genes are silent. Further, H3-K9 can be acetylated, indicating active gene expression, and cloned embryos are also hyperacetylated. These results suggest failure to both activate and to silence genes appropriately in cloned bovine embryos.
In normal blastocyts, there is lineage-specific hypermethylation of the inner cell mass compared with trophectoderm, but this asymmetry is not found in cloned embryos. Differences in acetylation of H3-K9 were also found, but differences were not as dramatic as those for methylation. Failure of trophectoderm to be hypomethylated may explain the aberrant placental development that is characteristic of cloned animals.
Nuclei from bovine granulosa cells yield a higher percentage of cloned blastocysts than nuclei from fetal fibroblasts and, as predicted, granulosa cell-derived clones had a higher proportion of normal DNA and histone methylation. One example of inappropriate expression of a gene is major histocompatibility (MHC) class I antigens that are not expressed by normal bovine trophectoderm/chorion. However, major histocompatibility class I genes are expressed by trophectoderm/ chorion of cloned conceptuses (Hill et al., 2002), which may put the cloned conceptus at risk for rejection by the maternal immune system.
Epigenetic programming is a normal part of development, including differentiation of primordial germ cells, sperm, oocytes, and conceptus tissues. This programming allows for both temporal and spatial repression and derepression of genes and development of healthy offspring. There is no evidence that the basic genome of cloned animals differs from that of the donor of cells used for somatic cell nuclear cloning.
While most cloned animals do not survive, there is considerable evidence that a very low percentage of cloned embryos do survive because of inappropriate silencing or over-expression of genes due to abnormalities such as genome methylation, histone assembly into nucleosomes, or chromatin remodeling by linker histones, polycomb group proteins, nuclear scaffold proteins, and transcription factors. Those clones that do survive beyond the neonatal period have an apparently normal phenotype. In either case, the basic genome of the cloned embryo is not modified. However, temporal and spatial aspects of gene expression during the course of development are apparently flawed in cloned embryos and the severity of these epigenetic events likely dictate the fate of clone.
Little evidence is available in the scientific literature to assess whether the progeny of cloned animals are at increased risk for inherited or developmental defects. The recently released 2003 Food and Drug Administration (FDA) draft report Animal Cloning: A Risk Assessment makes the following statement about the progeny of clones: “The underlying biological assumption for progeny animals is that generation of the cells that ultimately become ova and sperm naturally resets the epigenetic signals for gene expression. This process is thought to effectively “clear”
the genome of incomplete or inappropriate signals. The data to confirm this underlying assumption are limited but consistent across species” (FDA, 2003).
EVALUATING METHODS TO DETECT POTENTIAL UNINTENDED COMPOSITIONAL CHANGES AND ADVERSE HEALTH EFFECTS OF FOODS DERIVED FROM CLONED ANIMALS
Background
Historically, equivalence of tissue or food composition has been an important component of the regulatory process to evaluate food safety (CAST, 2001; Falk et al., 2002; Juskevich and Guyer, 1990). For genetically modified (GM) plants and the animal biotechnologies reviewed by FDA, the evaluation has included comprehensive compositional analyses of plants, tissues, and milk when appropriate. The committee found that a comparable approach for products from cloned animals—primarily meat and milk—would be an appropriate, scientifically based method for assessing compositional equivalence. Implicit to such assessments is that no increased health risk would be expected if the compositional analyses of animal products from cloned and noncloned animals were substantially equivalent.
Substantial Equivalence
Establishing equivalence of composition is evidence that substantive compositional changes did not occur in the animal as the result of the genetic modification event. Based on studies with GM plants, substantial equivalence is analogous to “as safe as its conventional counterpart” (CAST, 2001). The approach of substantial equivalence, however, is not absolute. Numerous biotechnology approaches developed in animal agriculture have been to intentionally design genetic modification to effect changes in the composition of target nutrients or other molecules (for review, see Karatzas, 2003; Niemann and Kues, 2000). Thus the process of assessing compositional equivalence needs to be undertaken and accommodated for with this in mind.
For example, cloned transgenic cows have been developed that produce milk with a marked increase in β-casein and k-casein (Brophy et al., 2003). Transgenic pigs have been produced that overexpress the bGH gene, which is associated with a dramatic reduction in carcass fat (85 percent reduction) and constituent fatty acid classes (Solomon et al., 1994). Genetic modification to change levels of selected nutrients in plants and animals has been, and is, an important objective of genetic engineering strategies to create designer foods (CAST, 2003; Falk et al., 2002). From the perspective of modifying the nutrient profile of foods, this has been done to increase beneficial nutrients or to decrease nutrients associated with adverse health effects, such as saturated fatty acids.
Safety Assessment
There is a long history of assessing the safety of foods introduced into the marketplace, involving an integrated multidisciplinary approach that incorporates molecular biology, protein chemistry and biochemistry, food chemistry, nutritional sciences, and toxicology. It is important to appreciate that absolute safety is not the objective with respect to any methodology or combination of methodologies used to evaluate complex substances such as food. The standard that customarily has been applied is that the food under evaluation should be as safe as an appropriate counterpart that has a long history of safe use. This comparative evaluation process is the foundation of establishing substantial equivalence of the food being evaluated. It also is important to emphasize that the food product itself, rather than the biotechnology process used to generate GM animals and cloned animals, should be the focal point of the evaluation. The primary objective of the safety review is to assess food safety; embedded in this is whether the process might affect the food. In addition, it is important to recognize that a statistically significant difference in one or more compounds in the food evaluated and the appropriate comparator does not necessarily imply an outcome with respect to human health. This must be evaluated on a case-by-case basis as part of the regulatory framework.
The current regulatory view of FDA is that gene-based modification of animals for food production falls under the Center for Veterinary Medicine regulations as new “animal drugs.” Since epigenetic changes in the genome may lead to changes in expression of one or more genes in a manner that may be analogous to gene expression changes observed in transgenic animals, the committee determined that cloned animals should initially be evaluated in a manner that is comparable to that for animals in which genetic engineering has been used to make specific genetic modifications.
The committee also determined that cloned animals developed from transgenic parent stock for the purpose of producing pharmaceutical compounds, biomaterials, and other products not related to food production, not be allowed to enter the food chain. The committee believes this approach reduces the regulatory burden of evaluating novel biotechnology products designed for purposes other than food production.
The committee further determined that epigenetic events are normal with respect to germ cells and conceptus development, but as the result of cloning there may be changes in the extent of epigenetic events that influence expression of one or more genes. Consequently, an integral component of our evaluation was to assess the existing and evolving methods available to assess changes, both targeted and globally, in gene expression.
If a change in gene expression or alteration in mRNA abundance does occur as the result of cloning, this would be expected to result in “downstream” changes in the level of a protein encoded for by the particular mRNA species. Depending
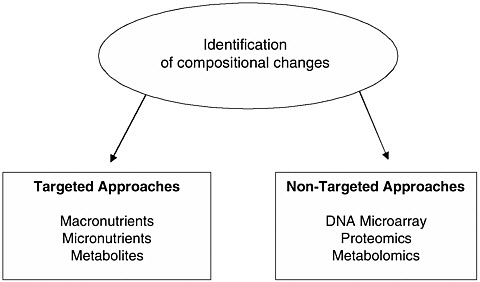
FIGURE 1 Assessment of compositional changes of food. Adapted from Kuiper et al. (2003).
upon the biological role of the protein, this could result in a change in the abundance of a cell constituent, such as a metabolite, protein, lipid, or carbohydrate. Thus quantifying whether any of these constituents are changed in a food is important to determine whether epigenetic modification is associated with any unintended compositional change. Consequently, methodologies that are currently used or are under development to detect changes in mRNA, protein, metabolites, and nutrients will be evaluated.
Evaluation of Unintended Composition Changes
Various analytical methods can be utilized to identify compositional changes in food, irrespective of whether these are intended or unintended (Kuiper et al., 2003; also discussed in detail in Chapter 4 of the main report). These strategies can be broadly categorized as either targeted or nontargeted (see Figure 1).
Targeted Approach
Historically, the targeted analytical approach has been an important component of the FDA review process. In this approach, known individual compounds, such as nutrients, are quantified (see Box 1a and 1b for representative listing of important macronutrients and micronutrients).
|
Box 1a
|
While analysis of specific, known compounds is important, the targeted approach may miss some unexpected changes because the list of compounds selected for assay is not all-inclusive. For GM crops, the Organization for Economic Cooperation and Development has published a list of analytes to quantify in assessing compositional changes (OECD, 2001). For GM plants, the analyses have provided information on macronutrients, micronutrients, antinutritive factors, and naturally occurring toxins. For cloned animals, it will be important to establish a comparable list. Box 1a and 1b present proposed lists of nutrients that could be quantified as part of the compositional equivalence determination for cloned animals.
|
Box 1b
|
Nontargeted Approach
Because of the vast number of individual constituents found in animal products, quantifying all of the macronutrients and micronutrients found in various tissues and in milk as a means to determine compositional equivalence is neither feasible nor practical. To enhance the likelihood of detecting unintended changes, nontargeted analytical approaches, or profiling methods such as DNA array, proteomics, and metabolomics, have been proposed as an approach for characterizing changes in GM plants (Kuiper et al., 2003; Schilter and Constable, 2002; also see Chapter 5 of the main report). The use of these profiling methodologies has the potential to provide a more extensive or global quantification of mRNA, protein, and metabolites to determine whether changes in one or more have occurred as the result of the cloning procedure.
Integration of Targeted and Nontargeted Approaches
Integration of both targeted and nontargeted approaches is a promising means to assess whether cloning has induced unintended changes in composition of meat or milk. Figure 2 presents a flow chart illustrating sequential and parallel assay steps that could be part of the profiling approach for detecting unintended effects. It must be emphasized, however, that both this report and information in the committee’s main report underscore the need to establish and validate the requisite methodological specificity and sensitivity of the various nontargeted profiling methods before they can be used in practice to reliably determine whether

FIGURE 2 Flow chart for conducting profiling analytical techniques.
unintended changes have occurred as the result of cloning (for review, see Kuiper et al., 2003).
Regardless of the analytical approaches taken to establish compositional equivalence, the following two issues must be addressed.
-
Defining threshold criteria for concluding that a significant change has occurred in a constituent of an animal product and whether this change is associated with an unintended (and adverse) health effect in humans; and
-
Standardization of sampling procedures for profiling techniques and
-
interlaboratory testing and assay validation (including assay sensitivity) among laboratories.
Comparator Databases
In addition to validating profiling methodologies as reliable analytical approaches to evaluate compositional equivalence, it is important to establish, validate, and utilize databases containing detailed information about normal variations in mRNA, protein, and metabolite profiles within a species. These comparator databases should account for the normal variation within an animal species that occurs in composition during growth and development, different physiological states—such as lactating versus nonlactating—and different animal management and rearing practices.
Consequently, the committee found that appropriate comparator databases could be derived from compositional data for the tissue or product produced by the species used for cloning during the time a particular species is used for productive purposes. In this example, the comparator database is not derived from animals that have been genetically modified by the application of transgenic biology.
Assessment of Methods to Detect Unintended Health Effects
Because each cloned animal is expected to be unique from an epigenetic basis, the profile of nutrients and metabolites to be assayed may need to be established on a case-by-case basis. The committee found that cloning creates a potential for temporal and spatial changes in gene expression due to epigenetic events, but without changing the basic genome of any species. There is no scientific evidence that cloning is associated with any unintended compositional change that results in an unintended health consequence in humans.
Since FDA treats genetic modification of animals as a new animal drug, it is beneficial to use the review process for recombinant bST as a case study to comment on methods of approach for evaluating health effects of foods derived from cloned animals. Bovine somatotropin was the first product of biotechnology approved for commercial use in animal agriculture. The approach for supplementing cows with bST is to provide the recombinant protein by injection.
Administration of bST increases blood levels of the protein leading to the consequent biological effects. The biological cascade invoked by administration of a protein, such as bST, is comparable to the situation where overexpression of a gene occurs, resulting in an increase in the level of a gene product due to epigenetic events. In turn, the increased level of the gene product increases the target protein in the blood and tissues with the attendant biological effect.
An integral component of the FDA review process is to establish food safety of the compound undergoing investigation. In the case of cloned animals, there is no compound per se to review since no single drug or compound is being ad-
ministered to the target animal. Rather, any change that might occur as the result of cloning would be alterations in expression of one or more genes in its basic genome.
The uncertainty of what gene or genes might be affected at the expression level by cloning greatly hinders the use of any targeted approach for selecting a specific gene or few genes a priori as a means to identify an unintended effect in gene expression. Consequently, nontargeted profiling methods that provide a more global assessment of compositional changes are needed. As previously discussed, however, the nontargeted profiling techniques are not yet appropriate to use as a reliable and reproducible means to assess unintended changes in mRNA, protein, or metabolite levels. The committee concluded it is important that research be supported to evolve and refine the nontargeted profiling methodologies that, as mentioned previously, include DNA microarray, proteomics, and metabolomics.
In the future, using profiling methods to establish which genes are expressed differently will be important to not only to establish identity of the gene, but also to help establish the knowledge base about the biology of the gene product. As discussed above for gene products for which there is an understanding of the biology, safety evaluations could be made of a known gene product in a manner consistent with present FDA guidelines. However, for many gene products there will be little or no information about the biology. This is problematic because of the time and cost associated with establishing a sufficient scientific understanding of the biology.
In the case of a known gene product, FDA’s review of bST provides a case study of what methods can be used to evaluate food safety. For a protein like bST, an understanding of the chemical nature, biological activity, and potential for harmful residues is important (Juskevich and Guyer, 1990). A determination of oral activity of the protein “drug” is initially required. The design of the oral toxicity studies is based on the known biological activities of the protein.
For bST, FDA required that rats be treated orally with “up to 100 times or more the dose of bST administered daily to cattle (on the basis of mg/kg of body weight) for at least 14 days” (Juskevich and Guyer, 1990). In this case, a known gene product, recombinant bST, was available for oral feeding studies.
The approach of feeding a food product derived from cloned animals to meet the 100-fold food safety margin is not feasible. Feeding this large quantity of a single food, which at some point on the dose-response curve the test food would be the only dietary source, to the test animal, may result in associated problems caused by nutrient deficiencies that may occur as the result of one food being predominant in the diet. This could be interpreted as an unintended effect when, in fact, the effect observed reflects inadequate intake of an essential nutrient.
It is important to identify each gene product affected by cloning to establish biological function of the gene product. The gene product, as a defined chemical, could be evaluated for hazard potential using conventional toxicology approaches
in a way approved for bST (see Juskevich and Guyer, 1990). If expression of numerous genes is affected by the cloning event, this evaluation will be challenging, costly, and burdensome, both for the petitioner and the regulatory agency. Since there is no evidence that food from cloned animals poses any increased health risk to the consumer, it could be concluded that food from cloned animals should be approved for consumption. However, the paucity of evidence in the literature on this topic makes it impossible to provide scientific evidence to support this position.
In view of the challenges discussed about the application of profiling methods to evaluate unintended changes in compositional profile, coupled with the lack of data in the scientific literature, other methods to evaluate such changes are those wherein the analysis is targeted for a specific macronutrient, micronutrient, or metabolite. The methods for the nutrients listed in Box 1a and 1b have been developed, validated, and are sufficient for the compositional assessment. Clearly, this approach is dependent upon an approach wherein a number of targeted compounds are assayed. Determining the list of compounds to assay is important.
Since many constituents of food are biologically degraded by heat processing or cooking and the digestive process, it is expected that this would render them biologically inactive. There are notable exceptions, specifically those proteins that induce an allergic response. For the vast majority of other molecules, biological inactivation via cooking or digestion provide another means to assure that any unintended change in a food component are not associated with increased health risks to the consumer.
On October 31, 2003, FDA released a preliminary report in which it was concluded that “Edible products from normal, healthy clones or their progeny do not appear to pose increased food consumption risks relative to conventional animals” (FDA, 2003). At present, there is no supportive evidence for increased risk to consumers of animal products from cloned animals, with the exception that it would not be prudent to allow animals that are genetically engineered to produce pharmaceuticals or other biologics, such as silk, to enter the food chain.
Other Considerations
The committee was asked to present a position about whether transgenic animals classified as “no-takes” should be permitted to enter the food chain. No-takes are animals in which the transgene is not incorporated into the genome. Implicit to this is the use of molecular techniques, including southern blotting, noncoding gene integration tags to detect integration of a gene, and real-time polymerase chain reaction to detect the presence of the transgene and the number of copies integrated into the genome. If the transgene DNA is not detected, the animal is considered a no-take. A list of tissues to sample, which will assure the transgene is not expressed in one tissue, but is expressed in another, must be
established. If animals are determined to be no-takes, there is no scientifically based rationale to exclude them from entering the food chain.
Animal Identification
A challenge to regulatory oversight of cloned and transgenic animals is development and implementation of effective programs to monitor the presence of these animals. The challenges this poses for transgenic animals have been reviewed by Howard and colleagues (2001), and an important issue to be resolved is whether cloned animals should be differentiated from noncloned animals at the point of entry of animals into the food system. Currently, there is no analytical method available to differentiate cloned from noncloned animals. However, should subsequent scientific review establish an increased risk to human health associated with the consumption of food products from cloned animals, it will be necessary to distinguish cloned animals from noncloned animals prior to entry into the food chain.
It is envisioned that the greatest likelihood of increased risk may arise from cloned, transgenic animals in which the genetic modification, as the result of transgenesis, has been made for the production of biomaterials or pharmaceuticals. In this case, methodological approaches are available to identify the transgene. This will be important in order to differentiate cloned, transgenic animals developed for food production purposes from cloned, transgenic animals developed for the production of biomaterials, pharmaceuticals, and other non-food purposes.
Due to the possible need to differentiate cloned, transgenic animals, a national system for animal identity and identity preservation is required. This system must be implemented at the point of slaughter or processing to rapidly and inexpensively identify the presence of cloned, transgenic animals or products derived from these animals. While the question of animal identification is beyond the scope of this report, it will likely be an important component of future oversight processes developed to monitor the entry of cloned, transgenic animals into the food system.
CONCLUSIONS
-
Profiling techniques are appropriate for establishing compositional differences between cloned and noncloned animals.
-
Profiling methods and their interpretation are not sufficiently developed to allow direct assessment of potential health effects associated with most unintended compositional changes.
-
There is no scientific basis to exclude animals deemed to be “no-takes” from entering the food chain.
-
There is a need to improve our ability to detect and assess the health
-
consequences of unintended changes in GM foods, such as better tools for toxicology assessment and a more robust knowledge base regarding which components impact health.
-
Given the possibility that foods with unintended changes could enter the marketplace, there is a need to enhance our capacity for postmarket surveillance of exposure and effects.
RECOMMENDATIONS
-
Targeted analysis of selected nutrients in animal products should be performed on a case-by-case basis.
-
Standardized sampling methodology, validation, and performance-based analyses should be performed for targeted analyses.
-
Standardized methodology, validation, and performance-based analyses should be performed for profiling analyses.
-
Publicly available compound identification databases should be developed that contain information such as mass spectra and nuclear magnetic resonance spectra where appropriate.
-
Publicly accessible databases should also be developed for compound profiles of animal-derived foods (see also Research Recommendations in Chapter 7 of the main report).
-
Animal identity and identity preservation systems should be improved for tracking animals and animal products through the food chain.
REFERENCES
Bauman DE. 1992. Bovine somatotropin: Review of an emerging animal technology. J Dairy Sci 75:3432–3451.
Bauman DE. 1999. Bovine somatotropin and lactation: From basic science to commercial application. Domest Anim Endocrinol 17:101–116.
Brophy B, Smolenski G, Wheeler T, Wells D, L’Huillier P, Laible G. 2003. Cloned transgenic cattle produce milk with higher levels of β-casein and k-casein. Nat Biotechnol 21:157–162.
CAST (Council for Agricultural Science and Technology). 2001. Evaluation of the U.S. Regulatory Process for Crops Developed Through Biotechnology. Issue 19. Washington, DC: CAST.
CAST. 2003. Biotechnology in Animal Agriculture: An Overview. Issue 23. Washington, DC: CAST.
Chilliard Y. 1989. Long-term effects of recombinant bovine somatotropin (rBST) on dairy cow performances: A review. In: Sejrsen K, Vestergaard M, Neimann-Sorensen A, eds. Use of Somatotropin in Livestock Production. New York: Elsevier Applied Science. Pp. 61–87.
Cummins JM. 2001. Cytoplasmic inheritance and its implications for animal biotechnology. Theriogenology 55:1381–1399.
De Sousa PA, King T, Harkness L, Young LE, Walker SK, Wilmut I. 2001. Evaluation of gestational deficiencies in cloned sheep fetuses and placentae. Biol Reprod 65:23–30.
Etherton TD, Bauman DE. 1998. The biology of somatotropin in growth and lactation of domestic animals. Physiol Rev 78:745–761.
Falk MC, Chassey BM, Harlander SK, Hoban TJ, McGloughlin N, Akhlaghi AR. 2002. Food biotechnology: Benefits and concerns. J Nutr 132:1384–1390.
FDA (U.S. Food and Drug Administration). 2003. Animal Cloning: A Risk Assessment, Draft Executive Summary. Washington, DC: U.S. Food and Drug Administration. Online. Available at http://www.fda.gov/cvm/index/cloning/CLRAES.doc. Accessed January 4, 2004.
Hill JR, Roussel AJ, Cibelli JB, Edwards JF, Hooper NL, Miller MW, Thompson JA, Loonery CR, Westhusin ME, Robl JM, Stice SL. 1999. Clinical and pathological features of cloned transgenic calves and fetuses (13 case studies). Theriogenology 51:1451–1465.
Hill JR, Burghardt RC, Jones K, Long CR, Looney CR, Shin T, Spencer TE, Thompson JA, Winger QA, Westhusin ME. 2000. Evidence for placental abnormality as the major cause of mortality in first-trimester somatic cell cloned fetuses. Biol Reprod 63:1787–1794.
Hill JR, Schlafer DH, Fisher PJ, Davies CJ. 2002. Abnormal expression of trophoblast major histocompatibility complex class I antigens in cloned bovine pregnancies is associated with a pronounced endometrial lymphocytic response. Biol Reprod 67:55–63.
Howard TH, Homan EJ, Bremel RD. 2001. Transgenic livestock: Regulation and science in a changing environment. J Anim Sci 79:E1–E11.
Jaenisch R, Wilmut I. 2001. Developmental biology. Don’t clone humans! Science 291:2552.
Juskevich JS, Guyer CG. 1990. Bovine growth hormone: Human food safety evaluation. Science 249:875–884.
Karatzas CN. 2003. Designer milk from transgenic clones. Nat Biotechnol 21:138–139.
Kouzarides T. 2002. Histone methylation in transcriptional control. Curr Opin Genet Develop 12:198–209.
Kuhholzer B, Prather RS. 2000. Advances in livestock nuclear transfer. Proc Soc Exp Biol Med 224:240–245.
Kuiper HA, Kok EJ, Engel KH. 2003. Exploitation of molecular profiling techniques for GM food safety assessment. Curr Opin Biotechnol 14:238–243.
Niemann H, Kues WA. 2000. Transgenic livestock: Premises and promises. Anim Reprod Sci 60–61:277–293.
NRC (National Research Council). 1994. Metabolic Modifiers: Effects on the Nutrient Requirements of Food-Producing Animals. Washington, DC: National Academy Press.
NRC. 2002. Animal Biotechnology: Science-Based Concerns. Washington, DC: The National Academies Press.
OECD (Organization for Economic Cooperation and Development). 2001. Consensus Document on Compositional Considerations for New Varieties of Soybean: Key Food and Feed Nutrients and Anti-nutrients. Online. Available at htt[://www.olis.oecd.org/olis/2001doc.nsf/43bb6130e5e86e5fc12569fa005d004c/cdb400c627da47a5c1256b17002f840d/$FILE/JT00117705.PDF. Accessed January 16, 2003.
Polejaeva IA, Chen SH, Vaught TD, Page RL, Mullins J, Ball S, Dai Y, Boone J, Walker S, Ayares DL, Colman A, Campbell KH. 2000. Cloned pigs produced by nuclear transfer from adult somatic cells. Nature 407:86–90.
Rideout WM, Eggan K, Jaenisch R. 2001. Nuclear cloning and epigenetic reprogramming of the genome. Science 293:1093–1098.
Santos F, Zakhartchenko V, Stojkovic M, Peters A, Jenuwein T, Wolf E, Reik W, Dean W. 2003. Epigenetic marking correlates with developmental potential in cloned bovine preimplantation embryos. Current Biol 13:1116–1121.
Schilter B, Constable A. 2002. Regulatory control of genetically modified (GM) foods: Likely developments. Toxicol Lett 127:341–349.
Solomon MB, Pursel VG, Paroczay EW, Bolt DJ. 1994. Lipid extraction of carcass tissue from transgenic pigs expressing a bovine growth hormone gene. J Anim Sci 72:1242–1246.
Tamashiro KLK, Wakayama T, Akutsu H, Yamazaki Y, Lachey JL, Wortman MD, Seely RJ, D’Alessio DA, Woods SC, Yanagamachi R, Sakai RR. 2002. Cloned mice have an obese phenotype not transmitted to their offspring. Nat Med 8:262–267.
Westhusin ME, Long CR, Shin T, Hill JR, Looney CR, Pryor JH, Piedrahita JA. 2001. Cloning to reproduce desired genotypes. Theriogenology 55:35–49.
Willadsen SM. 1989. Cloning of sheep and cow embryos. Genome 31:956–962.
Willadsen SM, Polge C. 1981. Attempts to produce monozygotic quadruplets in cattle by blastomeres separation. Vet Rec 108:211–213.
Wilmut I, Schnieke AE, McWhir J, Kind AJ, Campbell KH. 1997. Viable offspring derived from fetal and adult mammalian cells. Nature 385:810–813.
Wilmut I, Beaujean N, DeSousa PA, Dinnyes A, King TJ, Paterson LA, Wells DN, Young LE. 2002. Somatic cell nuclear transfer. Nature 419:583–587.
Xue F, Tian C, Du F, Kubota C, Taneja M, Dinnyes A, Dai Y, Levine H, Pereira LV, Yang X. 2002. Abberant patterns of X chromosome inactivation in bovine clones. Nat Genet 31:216–220.




















