3
Conceptualizing, Measuring,
and Studying Reproducibility
The second session of the workshop provided an overview of how to conceptualize, measure, and study reproducibility. Steven Goodman (Stanford University School of Medicine) and Yoav Benjamini (Tel Aviv University) discussed definitions and measures of reproducibility. Dennis Boos (North Carolina State University), Andrea Buja (Wharton School of the University of Pennsylvania), and Valen Johnson (Texas A&M University) discussed reproducibility and statistical significance. Marc Suchard (University of California, Los Angeles), Courtney Soderberg (Center for Open Science), and John Ioannidis (Stanford University) discussed assessment of factors affecting reproducibility. Mark Liberman (University of Pennsylvania) and Micah Altman (Massachusetts Institute of Technology) discussed reproducibility from the informatics perspective.
In addition to the references cited in this chapter, the planning committee would like to highlight the following background references: Altman et al. (2004); Begley (2013); Bernau et al. (2014); Berry (2012); Clayton and Collins (2014); Colquhoun (2014); Cossins (2014); Donoho (2010); Goodman et al. (1998); Jager and Leek (2014); Li et al. (2011); Liberati et al. (2009); Nosek et al. (2012); Peers et al. (2012); Rekdal (2014); Schooler (2014); Spence (2014); and Stodden (2013).
DEFINITIONS AND MEASURES OF REPRODUCIBILITY
Steven Goodman, Stanford University School of Medicine
Steven Goodman began his presentation by explaining that defining and measuring reproducibility is difficult; thus, his goal was not to list every measure but rather to provide an ontology of ways to think about reproducibility and to identify some challenges going forward. When trying to define reproducibility, Goodman noted, one needs to distinguish among reproducibility, replicability, repeatability, reliability, robustness, and generalizability because these terms are often used in widely different ways. Similarly, related issues such as open science, transparency, and truth are often ill defined. He noted that although the word “reproducibility” is often used in place of “truth,” this association is often inaccurate.
Goodman mentioned a 2015 Academy Award acceptance speech in which Julianne Moore said, “I read an article that said that winning an Oscar could lead to living five years longer. If that’s true, I’d really like to thank the Academy because my husband is younger than me.” According to Goodman, the article that Moore referenced (Redelmeier and Singh, 2001) had analysis and data access issues. He noted that an examination of some of these issues was later added to the discussion section of the paper and when the study was repeated (Sylvestre et al., 2006), no statistically significant added life expectancy was found for Oscar winners. In spite of the flawed original analysis, it is repeatedly referenced as fact in news stories. Goodman said this is a testament to how difficult it is to rectify the impact of nonreproducible findings.
Many philosophers and researchers have thought about the essence of truth over the past 100 years. William Whewell, Goodman noted, was one of the most important philosophers of science of the 19th century. He coined many common words such as scientist, physicist, ion, anode, cathode, and dielectric. He was an influential thinker who inspired Darwin, Faraday, Babbage, and John Stuart Mill. Goodman explained that Whewell also came up with the notion of consilience: that when a phenomenon is observed through multiple independent means, its validity may be greater than could be ascribed to any one of the individual inferences. This notion is used in the Bradford Hill criteria for causation1 (Hill, 1965) and in a range of scientific teaching. Edwin Wilson reenergized the notion of consilience in the late 1990s (Wilson, 1998). Goodman explained that the consilience discussions are in many ways a precursor to the current discussions of what kind of reproducibility can be designed and what actually gets science closer to the truth.
___________________
1 A group of minimal conditions is necessary to provide adequate evidence of a causal relationship between an incidence and a possible consequence.
Ronald Aylmer Fisher, Goodman explained, is one of the most important statisticians of the 20th century. Fisher wrote, “Personally, the writer prefers to set a low standard of significance at the 5 percent point. . . . A scientific fact should be regarded as experimentally established only if a properly designed experiment rarely fails to give this level of significance” (1926). Goodman stated that if this were how science was practiced by even by a small percentage of researchers, there would be significant progress in the realm of reproducibility.
Goodman noted that it is challenging to discuss reproducibility across the disciplines because there are both definitional and procedural differences among the various branches of science. He suggested that disciplines may cluster somewhat into groups with similar cultures as follows:
- Clinical and population-based sciences (e.g., epidemiology, clinical research, social science),
- Laboratory science (e.g., chemistry, physics, biology),
- Natural world-based science (e.g., astronomy, ecology),
- Computational sciences (e.g., statistics, applied mathematics, computer science, informatics), and
- Psychology.
Statistics, however, works in all sciences, and most statisticians cross multiple domain boundaries. This makes them ideal ambassadors of methods and carriers of ideas across disciplines, giving them the scope to see commonalities where other people might not.
Goodman noted that while some mergers are occurring—such as among genomics, proteomics, and economics—a few key differences exist between these communities of disciplines:
- Signal-measurement error ratio. When dealing with human beings, it is difficult to increase or decrease the error simply by increasing the sample size. In physics, however, engineering a device better can often reduce error. The calibration is different across fields and it results in a big difference in terms of what replication or reproducibility means.
- Statistical methods and standards for claims.
- Complexity of designs/measurement tools.
- Closeness of fit between a hypothesis and experimental design/data. In psychology, if a researcher is studying the influence of patriotic symbols on election choices, there are many different kinds of experiments that would fit that broad hypothesis. In contrast, a biomedical researcher studying the acceptable dose of aspirin to prevent heart attacks is faced with a much narrower range of experimental design. The closeness of fit has tremendous
-
bearing on what is considered a meaningful replication. While in many disciplines a language of direct replication, close replication, and conceptual replication exists, that language does not exist in biomedicine.
- Culture of replication, transparency, and cumulative knowledge. In clinical research and trials, it is routine to do studies and add their results. If one study is significant but the next one is not, neither is disconfirmed; rather, all the findings are added and analyzed together. However, in other fields it would be seen as a crisis if one study shows no effect while another study shows a significant effect.
- Purpose to which findings will be put (consequences of false claims). This is something that lurks in the background, especially with high-stakes research where lives are at risk. The bar has to be very high, as it is in clinical trials (Lash, 2015).
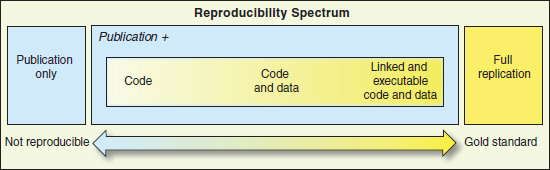
Goodman highlighted some of the literature that describes how reproducible research and replications are defined. Peng et al. (2006) defined criteria for reproducible epidemiology research (see Table 3.1), which state that replication of results requires that the analytical data set, the methods of the computer code, and all of the necessary metadata in the documentation necessary to run that code be available and that standard methods for distribution be used. Goodman explained that, in this case, reproducible research was research where you could see the data, run the code, and go from there to see if you could replicate the results or make adjustments. These criteria were applied to computational science a few years later (Peng, 2011), as shown in Figure 3.1. In this figure, the reproducibility spectrum progresses from code only, to code and data, to link and executable code and data. Goodman noted that this spectrum stops short of “full replication,” and Peng stated that “the fact that an analysis is reproducible does not guarantee the quality, correctness, or validity of the published results” (2011).
Goodman gave several examples to illustrate difficulties in reproducibility, including missing or misrepresented data in an analytical data set and limited access to important information not typically published or shared (such as case study reports in the case of Doshi et al. [2012]).
The National Science Foundation’s Subcommittee on Robust Research defined the following terms in their 2015 report on reproducibility, replicability, and generalization in the social, behavioral, and economic sciences:
- Reproducibility. “The ability of a researcher to duplicate the results of a prior study using the same materials . . . as were used by the original investigator. . . . A second researcher might use the same raw data to build the same analysis files and implement the same statistical analysis . . . [in an attempt to] yield the same results. . . . If the same results were not obtained,
TABLE 3.1 Criteria for Reproducible Epidemiologic Research
| Research Component | Requirement |
|---|---|
| Data | Analytical data set is available. |
| Methods | Computer code underlying figures, tables, and other principal results is made available in a human-readable form. In addition, the software environment necessary to execute that code is available. |
| Documentation | Adequate documentation of the computer code, software environment, and analytical data set is available to enable others to repeat the analyses and to conduct other similar ones. |
| Distribution | Standard methods of distribution are used for others to access the software, data, and documentation. |
SOURCE: Peng et al. (2006).

- Replicability. “The ability of a researcher to duplicate the results of a prior study if the same procedures are followed but new data are collected . . . a failure to replicate a scientific finding is commonly thought to occur when one study documents [statistically significant] relations between two or more variables and a subsequent attempt to implement the same operations fails to yield the same [statistically significant] relations” (NSF, 2015). Failure to replicate can occur when methods from either study are flawed
the discrepancy could be due to differences in processing of the data, differences in the application of statistical tools, differences in the operations performed by the statistical tools, accidental errors by an investigator, and other factors. . . . Reproducibility is a minimum necessary condition for a finding to be believable and informative” (NSF, 2015).
or sufficiently different or when results are in fact statistically compatible in spite of differences in significance. Goodman questioned if this is an appropriate and sufficient definition.
Goodman referenced several recent and current reproducibility efforts, including the Many Labs Replication Project,2 aimed at replicating important findings in social psychology. He noted that replication in this context means repeating the experiment to see if the findings are the same. He also discussed the work of researchers from Bayer who are trying to replicate important findings in basic science for oncology; Prinz et al. stated the following:
To substantiate our incidental observations that published reports are frequently not reproducible with quantitative data, we performed an analysis of our early (target identification and validation) in-house projects in our strategic research fields of oncology, women’s health and cardiovascular diseases that were performed over the past 4 years. . . . In almost two-thirds of the projects, there were inconsistencies between published data and in-house data that either considerably prolonged the duration of the target validation process or, in most cases resulted in termination of the projects because the evidence that was generated for the therapeutic hypothesis was insufficient to justify further investment in these projects. (2011)
Goodman pointed out that “reproducible” in this context denotes repeating the experiment again. C. Glenn Begley and Lee Ellis found that only 6 of 53 landmark experiments in preclinical research were reproducible: “When findings could not be reproduced, an attempt was made to contact the original authors, discuss the discrepant findings, exchange reagents and repeat experiments under the author’s direction, occasionally even in the laboratory of the original investigator” (2012). Sharon Begley wrote about C. Glenn Begley’s experiences in a later Reuters article: [C. Glenn] Begley met for breakfast at a cancer conference with the lead scientist of one of the problematic studies. “We went through the paper line by line, figure by figure,” said [C. Glenn] Begley. “I explained that we re-did their experiment 50 times and never got their results.” He said they’d done it six times and got this result once, but put it in the paper because it made the best story” (2012).
Goodman noted that the approach taken by Begley and Ellis is a valuable attempt to uncover the truth. He stressed that they did not just do a one-off attempt
___________________
2 The Many Labs Replication Project was a 36-site, 12-country, 6,344-subject effort to try to replicate a variety of important findings in social psychology (Klein et al., 2014; Nosek, 2014; Nauts et al., 2014; Wesselmann et al., 2014; Sinclair et al., 2014; Vermeulen et al., 2014; Müller and Rothermund, 2014; Gibson et al., 2014; Moon and Roeder, 2014; IJzerman et al., 2014; Johnson et al., 2014; Lynott et al., 2014; Žeželj and Jokić, 2014; Blanken et al., 2014; Calin-Jageman and Caldwell, 2014; Brandt et al., 2014). More information is available at Open Science Framework, “Investigating Variation in Replicability: A “Many Labs” Replication Project,” last updated September 24, 2015, https://osf.io/wx7ck/.
to replicate and compare the results; they tried every way possible to try to demonstrate the underlying phenomenon, even if that took attempting an experiment 50 times. This goes beyond replication and aims to reveal the true phenomenon; however, he noted that the same language is being used.
There is also no clear consensus about what constitutes reproducibility, according to Goodman. He quoted from a paper by C. Glenn Begley and John Ioannidis:
This has been highlighted empirically in preclinical research by the inability to replicate the majority of findings presented in high-profile journals. The estimates for irreproducibility based on these empirical observations range from 75% to 90%. . . . There is no clear consensus at to what constitutes a reproducible study. The inherent variability in biological systems means there is no expectation that results will necessarily be precisely replicated. So it is not reasonable to expect that each component of a research report will be replicated in perfect detail. However, it seems completely reasonable that the one or two big ideas or major conclusions that emerge from a scientific report should be validated and withstand close interrogation. (2015)
Goodman observed that in this text the terms “inability to replicate” and “irreproducibility” are used synonymously, and the words “reproducible,” “replicable,” and “validated” all relate to the process of getting at the truth of the claims.
Goodman also mentioned an article from Francis Collins and Larry Tabak (2014) on the importance of reproducibility:
A complex array of other factors seems to have contributed to the lack of reproducibility. Factors include poor training of researchers in experimental design; increased emphasis on making provocative statements rather than presenting technical details; and publications that do not report basic elements of experimental design. . . . Some irreproducible reports are probably the result of coincidental findings that happened to reach statistical significance, coupled with publication bias. Another pitfall is the overinterpretation of creative “hypothesis-generating” experiments, which are designed to uncover new avenues of inquiry rather than to provide definitive proof for any single question. Still, there remains a troubling frequency of published reports that claim a significant result, but fail to be reproducible.
Goodman summarized the three meanings of “reproducibility” as follows:
- Reproducing the processes of investigations: looking at what a study did and determining if it is clear enough to know how to interpret results, which is particularly relevant in computational domains. These processes include factors such as transparency and reporting, and key design elements (e.g., blinding and control groups).
- Reproducing the results of investigations: finding the same evidence or data, with the same strength.
- Reproducing the interpretation of results: reaching the same conclusions or inferences based on the results. Goodman noted that there are many cases in epidemiology and clinical research in which an investigation leads to a significant finding, but no associated claims or interpretations are stated, other than to assert that the finding is interesting and should be examined further. He does not think this is necessarily a false finding, and it may be a proper conclusion.
Goodman offered a related parsing of the goals of replication from Bayarri and Mayoral (2002):
- Goal 1: Reduction of random error. Conclusions are based on a combined analysis of (original and replicated) experiments.
- Goal 2: Validation (confirmation) of conclusions. Original conclusions are validated through reproduction of similar conclusions by independent replications.
- Goal 3: Extension of conclusions. The extent to which the conclusions are still valid is investigated when slight (or moderate) changes in the co-variables, experimental conditions, and so on, are introduced.
- Goal 4: Detection of bias. Bias is suspected of having been introduced in the original experiment. This is an interesting (although not always clearly stated) goal in some replications.
Goodman offered two additional goals that may be achieved through replication:
- Learning about the robustness of results: resistance to (minor or moderate) changes in the experimental or analytic procedures and assumptions, and
- Learning about the generalizability (also known as transportability) of the results: true findings outside the experimental frame or in a not-yet-tested situation. Goodman added that the border between robustness and generalizability is indistinct because all scientific findings must have some degree of generalizability; otherwise, the findings are not actually science.
Measuring reproducibility is challenging, in Goodman’s view, because different conceptions of reproducibility have different measures. However, many measures fall generally into the following four categories: (1) process (including key design elements), (2) replicability, (3) evidence, and (4) truth.
When discussing process, the key question is whether the research adhered to reproducible research-sharing standards. More specifically, did it provide code, data, and metadata (including research plan), and was it registered (for many clinical research designs)? Did it adhere to reporting (transparency) standards (e.g.,
CONSORT,3 STROBE,4 QUADAS,5 QUORUM,6 CAMARADES,7 REMARK8)? Did it adhere to various design and conduct standards, such as validation, selection thresholds, blinding, and controls? Can the work be evaluated using quality scores (e.g., GRADE9)? Did the researchers assess meta-biases that are hard to detect except through meta-research? Goodman noted that in terms of assessing meta-biases, one of the most important factors is selective reporting in publication. This is captured by multiple terms and often varies by discipline:
- Multiple comparisons (1950s, most statisticians),
- File drawer problem (Rosenthal, 1979),
- Significance questing (Rothman and Boice, 1979),
- Data mining, dredging, torturing (Mills, 1993),
- Data snooping (White, 2000),
- Selective outcome reporting (Chan et al., 2004),
- Bias (Ioannidis, 2005),
- Hidden multiplicity (Berry, 2007),
- Specification searching (Leamer, 1978), and
- p-hacking (Simmons et al., 2011).
Goodman observed that all of these terms refer to essentially the same phenomenon in different fields, which underscores the importance of clarifying the language.
In terms of results, Goodman explained, the methods used to assess replication are not clear. He noted some methods that are agreed upon within different disciplinary cultures include contrasting statistical significance, assessing statistical
___________________
3 The Consolidated Standards of Reporting Trials website is http://www.consort-statement.org, accessed January 6, 2016.
4 The Strengthening the Reporting of Observational Studies in Epidemiology website is http://www.strobe-statement.org, accessed January 12, 2016.
5 The Quality Assessment of Diagnostic Accuracy Studies website is http://www.bris.ac.uk/quadas/, accessed January 12, 2016.
6 See, for example, the Quality of Reporting of Meta-Analyses checklist for K.B. Filion, F.E. Khoury, M. Bielinski, I. Schiller, N. Dendukuri, and J.M. Brophy, “Omega-3 fatty acids in high-risk cardiovascular patients: A meta-analysis of randomized controlled trials,” BMC Cardiovascular Disorders 10:24, 2010, electronic supplementary material, Additional file 1: QUORUM Checklist, http://www.biomedcentral.com/content/supplementary/1471-2261-10-24-s1.pdf.
7 The Collaborative Approach to Meta-Analysis and Review of Animal Data from Experimental Studies website is http://www.dcn.ed.ac.uk/camarades/, accessed January 12, 2016.
8 The REporting recommendations for tumour MARKer prognostic studies website is http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2361579/, accessed January 12, 2016.
9 The Grading of Recommendations, Assessment, Development and Evaluations website is http://www.gradeworkinggroup.org/, accessed January 12, 2016.
compatibility, improving estimation and precision, and assessing contributors to bias/variability.
Assessing replicability from individual studies can be approached using probability of replication, weight of evidence (e.g., p-values, traditional meta-analysis and estimation, Bayes factors, likelihood curves, and the probability of replication as a substitute evidential measure), probability of truth (e.g., Bayesian posteriors, false discovery rate), and inference/knowledge claims. Goodman noted that there are many ways to determine the probability of replication, many of which include accounting for predictive power and resampling statistics internally.
Goodman discussed some simple ways to connect evidence to truth and to replicability, as shown in Table 3.1. However, he noted that very few scientists are even aware that these measures can show how replication is directly related to strength of evidence and prior probability. A participant questioned shifting the conversation from reproducibility to truth and how to quantify uncertainty in truth. Specifically, since so much of the reproducibility discussion is fundamentally about error, is truth a specific target number in the analysis or is it a construct for which there is a range that would be acceptable? Goodman explained that replicability and reproducibility is a critical operational step on the road to increasing certainty about anything. Ultimately the goal is to accumulate enough certainty to declare something as established. He explained that the reproducibility in and of itself is not the goal; rather, it is the calculus that leads a community to agree after a certain number of experiments or different confirmations of the calculations that the result is true. This is why, according to Goodman, a hierarchy of evidence exists, and the results from an observational study are not treated in quite the same way as a clinical trial even though the numbers might look exactly the same in the standard errors. Some baseline level of uncertainty might be associated with the phenomenon under study, and additional uncertainties arise from measurement issues and covariance. To evaluate the strength of evidence, quantitative and qualitative assessments need to be combined. Goodman noted that some measures directly put a probability statement on truth as opposed to using operational surrogates. Only through that focus can we understand exactly how reproducibility, additional experiments, or increased precision raises the probability that we have a true result. However, he commented that there are no formal equations that incorporate all of the qualitative dimensions of assessing experimental quality and covariant inclusion or exclusion. To some extent, this can be captured through sensitivity analyses, which can be incorporated into that calculus. While the uncertainty can be captured in a Bayesian posterior through Bayesian model averaging, this is not always done and it is unclear if this is even the best way to proceed.
Goodman mentioned a highly discussed editorial (Trafimow and Marks, 2015) in Basic and Applied Social Psychology whose authors banned the use of indices related to null hypothesis significance testing procedures in that journal (including
p-values, t-values, F-values, statements about significant difference, and confidence intervals). Goodman concluded that reproducibility does not have a single agreed-upon definition, but there is a widely accepted belief that being able to demonstrate a physical phenomenon in repeated independent contexts is an indispensable and essential component of scientific truth. This requires, at minimum, a complete knowledge of both experimental and analytic procedures (both process-related reproducibility and the strength of evidence). He speculated that reproducibility is the focus instead of truth, per se, because frequentist logic, language, and measures do not provide proper evidential or probabilistic measures of truth. The statistical measures related to reproducibility depend on the criteria by which truth or true claims are established in a given discipline. Goodman stated that in some ways, addressing the problems would be a discipline-by-discipline reformation effort where statisticians have a special role.
Goodman reiterated that statisticians are in a position to see the commonalities of problems and solutions across disciplines and to help harmonize and improve the methods that we use to enhance reproducibility on the journey to truth. Statisticians communicate through papers, teaching, and development of curricula, but the curricula to cross many disciplines do not currently exist. He noted that these curricula have to cover the foundation of inference, threats to validity, and the calculus of uncertainty. Statisticians can also develop software that incorporates measures and methods that contribute to the understanding and maximization of reproducibility.
Goodman concluded his presentation by noting that enhancing reproducibility is a problem of collective action that requires journals, funders, and academic communities to develop and enforce the standards. No one entity can do it alone. In response to a participant question about why a journal could not force authors to release their data and methods without the collaboration of other journals, Goodman explained that many journals are in direct competition and no one journal wants to create more impediments for the contributors than necessary. Data sharing is still uncommon in many fields, and a journal may hesitate to require something of authors that other competitive journals do not. However, Goodman noted that many journals are starting to move in this direction. The National Institutes of Health and other funders are beginning to mandate data sharing. The analogous movement toward clinical trial registration worked only when the top journals declared they would not publish clinical trials that were not preregistered. While any one journal could stand up and be the first to similarly require data sharing, Goodman stressed that this is unlikely given the highly competitive publishing environment.
Another participant stated that even if all data were available, the basic questions about what constitutes reproducible research remain because there is not a clear conceptual framework. Goodman observed that there may be more opportunity for agreement about the underlying constructs than about language
because language seems to differ by field. Mapping out the underlying conceptual framework, however, and pointing out that while these words are applied here or there in different ways and cultures, everyone is clear about the underlying issues (e.g., the repeatability of the experiment versus the calculations versus the adjustments). If the broader community can agree on that conceptual understanding, and specific disciplines can map onto that conceptualization, Goodman is optimistic.
A participant questioned the data-sharing obligation in the case of a large data set that a researcher plans to use to produce many papers. Goodman suggested the community envision a new world where researchers mutually benefit from people making their data more available. Many data-sharing disciplines are already realizing this benefit.
Yoav Benjamini, Tel Aviv University
Yoav Benjamini began his presentation by discussing the underlying effort of reproducibility: to discover the truth. From a historical perspective, the main principle protecting scientific discoveries is a constant subjection to scrutiny by other scientists. Replicability became the gold standard of science during the 17th century, as illustrated by the debate associated with the air pump vacuum. That air pump was complicated and expensive to build, with only two existing in England. When the Dutch mathematician and scientist Christiaan Huygens observed unique properties within a vacuum, he traveled to England to demonstrate the phenomenon to scientists such as Robert Boyle (whom Benjamini cited as the first to introduce the methods section into scientific papers) and Thomas Hobbes. Huygens did not believe the phenomenon would be believed unless he could demonstrate it on a vacuum in England.
According to Fisher (1935), Benjamini explained, “We may say that a phenomenon is experimentally demonstrable when we know how to conduct an experiment which will rarely fail to give us statistically significant results.” Benjamini noted that this is the p-value, but since the p-value cannot be replicated a threshold is needed. However, there has been little in the literature about quantifying what it means to be replicable (Wolfinger, 2013) until recently with the introduction of the r-value.
Benjamini offered the following definitions to differentiate between reproducibility and replicability:
- Reproducibility of the study. Subject the study’s raw data to the same analysis used in the study, and arrive at the same outputs and conclusions.
- Replicability of results. Replicate the entire study, from enlisting subjects through collecting data, analyze the results in a similar but not necessarily identical way, and obtain results that are essentially the same as those reported in the original study (Peng, 2009; Nature Neuroscience, 2013; NSF, 2014).
He noted that this is not merely terminology because reproducibility is a property of a study, and replicability is a property of a result that can be proved only by inspecting other results of similar experiments. Therefore, the reproducibility of a result from a single study can be assured, and improving the statistical analysis can enhance its replicability.
Replicability assessment, Benjamini asserted, requires multiple studies, so it can usually be done within a meta-analysis. However, meta-analysis and replicability assessment are not the same. Meta-analysis answers the following question: Is there evidence for effects in at least one study? In contrast, a quantitative replicability assessment should answer this question: Is there evidence for effect in at least two (or more) studies? This is not in order to buy power but in order to buy a stronger scientific statement.
An assessment establishes replicability in a precise statistical sense, according to Benjamini. If two studies identify the same finding as statistically significant (with p-values P1 and P2), replicability is established if the union hypothesis H01 ∪ H02 is rejected in favor of the conjunction of alternatives
![]()
The r-value is therefore the level of the test according to which the alternate is true in at least two studies. Similar findings in at least two studies are a minimal requirement, but the strength of the replicability argument increases as the number of studies with findings in agreement increases. When screening many potential findings, the r-value is the smallest false discovery rate at which the finding will be among those where the alternate is true in at least two studies.
Benjamini concluded his presentation with several comments:
- Different designs need different methods. For example, an empirical Bayes approach can be used for genome-wide association studies (Heller and Yekutieli, 2014).
- The mixed-model analysis can be used as evidence for replicability.10 It has the advantage of being useful for enhancing replicability in single-laboratory experiments. The relation between the strength of evidence and power needs to be explored, perhaps through the use of r-values.
- Selective inference and uncertainty need to be introduced into basic statistical education (e.g., the Bonferroni correction method, false discovery rate, selective confidence intervals, post-selection inference, and data splitting).
___________________
10 See Chapter 2 for Benjamini’s previous discussion of this topic.
REPRODUCIBILITY AND STATISTICAL SIGNIFICANCE
Dennis Boos, North Carolina State University
Dennis Boos began his workshop presentation by explaining that variability refers to how results differ when an experiment is repeated, which naturally relates to reproducibility of results. The subject of his presentation was the variability of p-values (Boos and Stefanski, 2011); he asserted that p-values are valuable indicators, but their use needs to be recalibrated to account for replication in future experiments.
To Boos, reproducibility can be thought of in terms of two identical experiments producing similar conclusions. Assume summary measures T1 and T2 (such as p-values) are continuous and exchangeable.
![]()
then
![]()
Thus, before the two experiments are performed, there is a 50 percent chance that T2 > T1. Boos noted that for p-values, it is not unusual for weak effects that produce a p1 near, but less than, 0.05 to be followed by p2 larger than 0.05, which indicates that the first experiment is not reproducible. Boos commented that this is not limited to p-values; regardless of what measure is used, if the results are near the threshold of what is declared significant, there is a good chance that that standard will not be met in the subsequent experiment.
Boos paused to provide some background on standard errors. If a sample mean ![]() is being reported, the standard deviation s divided by the square root of the sample n is usually reported as the standard error,
is being reported, the standard deviation s divided by the square root of the sample n is usually reported as the standard error, ![]() , although this is not always meaningful (Miller, 1986). If regression coefficients
, although this is not always meaningful (Miller, 1986). If regression coefficients ![]() are reported, the standard error is easy to produce unless model selection is used. However, standard errors are not typically included when using Monte Carlo estimates, p-values, or R2, all of which can have high variability (Lambert and Hall, 1982; Boos and Stefanski, 2011). He noted that the log scale of p-values is more reliable.
are reported, the standard error is easy to produce unless model selection is used. However, standard errors are not typically included when using Monte Carlo estimates, p-values, or R2, all of which can have high variability (Lambert and Hall, 1982; Boos and Stefanski, 2011). He noted that the log scale of p-values is more reliable.
Boos argued that only the orders of magnitude of p-values, as opposed to their exact values, are accurate enough to be reported, with three rough ranges being of interest:
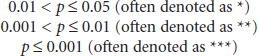
He suggested that some alternate methodologies could be useful in reducing variability:
- Bootstrap and jackknife standard errors for log10 (p-value), as suggested by Shao and Tu (1996);
- Bootstrap prediction intervals and bounds for pnew of a replicated experiment, as suggested by Mojirsheibani and Tibshirani (1996); and
- Estimate of the reproducibility probability P (pnew ≤ 0.05), which assesses the probability that a repeated experiment will produce a statistically significant result (Goodman, 1992; Shao and Chow, 2002).
Using the reproducibility probability, Boos argued that p-values of p ≤ 0.01 are necessary if the reported results are to an acceptable degree of reproducibility. However, the reproducibility probability has a large variance itself, so the standard error is large. Boos suggested using the reproducibility probability as a calibration but remaining aware of its limitations.
Boos also discussed Bayes calculations in the one-sided normal test (Casella and Berger, 1987). He gave an example of looking at the posterior probability over a class of priors. Using the Bayes factor, which is the posterior probability of the alternative over the null hypothesis, he illustrated that Bayes factors show similar repeatability problems as p-values when the values are near the threshold.
To conclude, Boos summarized that under the null hypothesis, p-values have large variability. The implications for reproducibility (in terms of the reproducibility probabilities ![]() ) are as follows:
) are as follows:
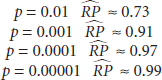
Reporting p-values in terms of ranges (specifically *, **, or ***) may be a reasonable approach for the broader community, he suggested.
Andreas Buja, Wharton School of the University of Pennsylvania
Andreas Buja stated that the research from Boos and Stefanski (2011) made many significant contributions to the scientific community’s awareness of sampling variability in p-values and showed that this variability can be quantified. That awareness led to a fundamental question: When seeing a p-value, is it believable that something similar would appear again under replication? Because a positive
answer is not a given, Buja argues that more stringent cutoffs than p = 0.05 are important to achieve replicability.
There are two basic pedagogical problems, according to Buja: (1) p-values are random variables; while they appear to be probabilities, they are transformed and inverted test statistics. (2) The p-value random variables exhibit sampling variability, but the depth of this understanding is limited and should be more broadly stated as data set–to–data set variability. Moving from p-value variability to bias, Buja suggested that multiple studies might be more advantageous than a single study with a larger sample because interstudy correlation can be significant.
Buja argued that statistics and economics need to work together to tackle reproducibility. Statistical thinking, he explained, views statistics as a quantitative epistemology and the science that creates protocols for the acquisition of qualified knowledge. The absence of protocols is damaging, and important distinctions are made between replicability and reproducibility in empirical, computational, and statistical respects. Economic thinking situates research within the economic system where incentives must be set right to solve the reproducibility problem. Buja argued that economic incentives (e.g., journals and their policies) should to be approached in conjunction with the statistical protocols.
Journals need to fight the file drawer problem, Buja said, by stopping the chase of “breakthrough science” and publishing, soliciting, and favorably treating replicated results and negative outcomes. He argued that institutions and researchers are lacking suitable incentives. Researchers will self-censor if journals treat replicated results and negative outcomes even slightly less favorably. Also, researchers will lose interest as soon as negative outcomes are apparent. In Buja’s view, negative results are important but often are not viewed as such by researchers because publishing them is challenging. Following the example of Young and Karr (2011), Buja argued that journals should accept or reject a paper based on the merit and interest of the research problem, the study design, and the quality of researcher, without knowing the outcomes of the study.
Statistical methods should take into account all data analytic activity, according to Buja. This includes the following:
- Revealing all exploratory data analysis, in particular visualizations;
- Revealing all model searching (e.g., lasso, forward/backward/all-subsets, Bayesian, cross validated, Akaike information criterion, Bayesian information criterion, residual information criterion);
- Revealing all model diagnostics and actions resulting from them; and
- Inferencing steps that attempt to account for all of the above.
In principle, Buja stated that any data-analytic action that could result in a different outcome in another data set should be documented, and the complete set of data-analysis inference should be undertaken in an integrated fashion.
While these objectives are not yet attained, Buja identified some work that is contributing to these goals. Post-selection inference, for example, is a method for statistical inference that is valid after model selection in linear models (Berk et al., 2013). Buja explained that this helps secure against attempts at model selection, including p-hacking. Inference for data visualization is also progressing, according to Buja. In principle, synthetic data can be plotted with and compared to actual data. Sources of synthetic data include permutations for independence tests, parametric bootstrap for model diagnostics, and sampling conditional data distributions given sufficient statistics. If the actual data plot can be differentiated from the synthetic plots, significance can be demonstrated (Buja et al., 2009).
Valen Johnson, Texas A&M University
Valen Johnson began by noting that the nature of p-values does not answer the question of whether a test statistic is bigger than it would be if the experiments were repeated; it does prove whether the null hypothesis is true.
Johnson offered the following review of the Bayesian approach to hypothesis testing: The Bayes theorem provides the posterior odds between two hypotheses after seeing the data. This demonstrates that the posterior probability of each hypothesis is equal to the Bayes factor times the prior odds between the two hypotheses.
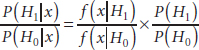
or
![]()
where the Bayes factor is the integral of the sampling density with respect to the prior distribution under that hypothesis:
![]()
and when doing null hypothesis significant testing, the prior distribution for the parameter under the null hypothesis is just a point mass π0(θ,H0). Bayesian methods are not used frequently because they make it difficult to specify what the prior distribution of the parameter is under an alternative hypothesis.
Recently there has been some methodology developed called uniformly most powerful Bayesian tests (UMPBTs) that provides a default specification of the prior
π1(θ,H1) under the alternative hypothesis (Johnson, 2013a). Johnson explained that the rejection region for UMPBT can often be matched to the rejection regions of classical uniformly most powerful tests. Interestingly, the UMPBT alternative places its prior mass at the boundary of the rejection region, which can be interpreted as the implicit alternative hypothesis that is tested by researchers in the classical paradigm. This methodology provides a direct correspondence among p-values, Bayes factors, and the posterior probability. Johnson noted that the prior distribution under the alternative is selected such that it will maximize the probability that the Bayes factor exceeds the threshold against all other possible prior distributions. He commented that it is surprising that such a prior distribution exists because it has to sustain for every data-generating value of the parameter.
For single-parameter exponential family models, Johnson explained that specifying the significance level of the classical uniformly most powerful test is equivalent to specifying the threshold that a Bayes factor must exceed to reject the null hypothesis in the Bayesian tests. So, the UMPBT has the same rejection region as the classical test. If one assumes that equal prior probability is assigned to the null hypothesis and the alternative hypothesis, then there is equivalence between the p-value and the posterior value and the null hypothesis is true.
Under this assumption and using UMPBT to calculate posterior probabilities and prior odds, a p-value of 0.05 leads to the posterior probability of the null hypothesis of 20 percent. Johnson stressed that the UMPBT was selected to give a high probability to rejecting the null at the threshold so the posterior probability would be greater than 20 percent if another prior were used. Johnson speculated that the widespread acceptance of a p-value of 0.05 reduces reproducibility in scientific studies. To get to a posterior probability that the null hypothesis is true with 0.05, he asserted that one really needs a p-value of about 0.005.
The next question Johnson posed was whether the assumed prior probability of the null hypothesis of only 0.5 is correct. He noted that when researchers do experiments, they do multiple experiments with the same data in multiple testing, conduct subgroup analysis, and have file drawer biases, among other issues. Given all these potential concerns, he speculated that the prior probability that should be assigned to the null should be much greater than 0.5. Assuming a higher prior probability of the null hypothesis would result in an even higher posterior probability. That is, a p-value of 0.05 would be associated with a posterior probability of the null hypothesis that was much greater than 20 percent.
In conclusion, Johnson recommends that minimum p-values of 0.005 and 0.001 should be designated as standards for significant and highly significant results, respectively (Johnson, 2013b).
Panel Discussion
After their presentations, Dennis Boos, Andreas Buja, and Valen Johnson participated in a panel discussion. During this session, two key themes emerged: the value of raising the bar for demonstrating statistical significance (be it through p-values, Bayes factors, or another measure) and the importance of protocols that contribute to the replicability of research.
Statistical Significance
A participant asked how individuals and communities could convince clinical researchers of the benefits of smaller p-values. Johnson agreed that there has been pushback from scientists about the proposal to raise the bar for statistical significance. But he noted that the evidence against a false null hypothesis grows exponentially fast, so even reducing the p-value threshold by a factor of 10 may require only a 60 percent increase in an experiment’s sample size. This is in contrast to the argument that massively larger sample sizes would be needed to get smaller p-values. He also noted that modern statistical methodology provides many ways to perform sequential hypothesis tests, and it is not necessary to conduct every test with the full sample size. Instead, Johnson said that a preliminary analysis could be done to see if there appears to be an effect and, if the results look promising, sample size could be increased. Buja commented that, from an economic perspective, fixed thresholds do not work. He argued that there should be a gradual kind of decision making to avoid the intrinsic problems of thresholds.
A participant wondered if the most effective approach might be to try to move people away from p-values to Bayes factors because the Bayes factor of 0.05 is more statistically significant than a p-value of 0.05. Since most researchers are only familiar with the operational meaning of the p-value, the participant argued that keeping the 0.05 value but changing the calculation leading to it may be easier. Johnson noted the challenge of bringing a Bayesian perspective into introductory courses, so educating researchers on Bayes factors is going to be difficult. Because of this challenge, he asserted that changing the p-value threshold to 0.005 might be easier.
A participant proposed an alternative way to measure reproducibility: tracking the ratio of the number of papers that attempted to verify the main conclusion of the report to the number that succeeded (Nicholson and Lazebnik, 2014). Another workshop participant questioned how well such a counting measure would work because many replication studies are underpowered and not truly conclusive on their own. Johnson explained that Bayes factors between experiments multiply together naturally so they serve as an easy way to combine information across multiple experiments.
In response to a participant’s question about how much reproducibility might improve if Bayes factors were used rather than p-values, Buja commented that many factors other than statistical methodology could play bigger roles. Boos added that both frequentist and Bayesian analyses will generally come to the same conclusions if done correctly. Johnson agreed that there are many sources of irreproducibility in scientific studies and statistics, and the use of elevated significance thresholds is just one of many factors.
A participant summarized that the level of evidence needs to be much higher than the current standard, be it through Bayes factors or p-values. The participant wondered how to get a higher standard codified in a situation where a variety of models and approaches are being used to analyze a set of data and come up with a conclusion. While a more stringent p-value or higher Bayes factor will help, it may be that neither will provide as much value as single hypothesis tests. But the prior probabilities of various kinds of hypotheses may matter, and that is more difficult to model. Johnson agreed that it is more difficult when working with more complicated models that have high-dimensional parameters. He agreed that the Bayesian principles are in place, but that implementing and specifying the priors is a much more difficult task.
A participant noted that whenever a threshold is used, regardless of what method, approach, process, or philosophy is in place, a result near the threshold would have more uncertainty than a finding within the threshold. Boos agreed that the threshold is just a placeholder and it is up to the community to realize that anything close to it may not reproduce. Johnson disagreed, stating that there needs to be a point at which a journal editor has to decide whether to publish a result or whether a physicist can claim a new discovery. The important takeaway, in Johnson’s view, is that this bar needs to be higher.
Protocols
A participant observed that although there are many suggested alternatives to p-values, none of them have been widely accepted by researchers. Johnson responded that UMPBT cannot be computed on all models and that p-values may be a reasonable methodology to use as long as the standards are tightened. Buja reinforced the notion that protocols are important even if they are suboptimal; reporting a suboptimal p-value, in other words, is an important standard.
A participant noted that Buja called for “accounting for all data analytical actions,” including all kinds of model selection and tuning; however, the participant would instead encourage scientists to work with an initial data set and figure out how to replicate the study in an interesting way. Buja agreed that replication is the goal, but protocols are essential to make a study fully reproducible by someone else. With good reporting, it may be possible to follow the research path taken by a re-
searcher, but it is difficult to know why particular paths were not taken when those results are not reported. He stressed that there is no substitute for real replication.
ASSESSMENT OF FACTORS AFFECTING REPRODUCIBILITY
Marc Suchard, University of California, Los Angeles
Marc Suchard began his presentation by discussing some recent cases of conflicting observational studies using nationwide electronic health records. The first example given was the case of assessing the exposure to oral bisphosphonates and the risk of esophageal cancer. In this case, Cardwell et al. (2010) and Green et al. (2010) had different analyses of the same data set to show “no significant association” and “a significantly increased risk,” respectively. Another example evaluated oral fluoroquinolones and the risk of retinal detachment; Etminan et al. (2012) and Pasternak et al. (2013) used different data sources and analytical methods to infer “a higher risk” and “not associated with increased risk,” respectively. Suchard’s final example was the case of pioglitazone and bladder cancer, as examined by Wei et al. (2013) and Azoulay et al. (2012), which “[do] not appear to be significantly associated” and “[have] an increased risk,” respectively, using the same population but different analytical methods.
Suchard explained that he is interested in what he terms subjective reproducibility to gain as much useful information from a reproducibility study as possible by utilizing other available data and methods. In particular, he is interested in quantitatively investigating how research choices in data and analysis might generate an answer that is going to be more reproducible, or more likely to get the same answer, if the same type of experiment were to be repeated.
For example, when examining associations between adverse drug events and different pharmaceutical agents using large-scale observational studies, several studies can be implemented using different methods and design choices to help identify the true operating characteristics of these methods and design choices. This could help guide the choice of what type of design to use in terms of reliability or reproducibility and better establish a ground truth of the positive and negative associations. Another advantage of this approach, Suchard explained, is to better identify the type I error rate and establish the 95 percent confidence interval. He stressed that reproducibility of large-scale observational studies needs to be improved.
As an empirical approach toward improving reproducibility, Suchard and others set up a 3-year study called the Observational Medical Outcomes Partnership (OMOP)11 to develop an analysis platform for conducting reproducible obser-
___________________
11 The Observational Medical Outcomes Partnership website is http://omop.org, accessed January 12, 2016.
vational studies. The first difficulty with looking at observational data across the world, he explained, is that observational data from electronic health records are stored in many different formats. Addressing this requires a universal set of tools and a common data model that can store all this information, so that everyone is extracting it in the same way when specifying particular inclusion and exclusion criteria. To do this, the data from multiple sources need to be translated into a common data model without losing any information. Then one of many study methods is specified and the associated design decisions are made. Currently, Suchard noted that very few of these choices are documented in the manuscripts that report the work or in the protocols that were set up to initially approve the studies, even though the information might exist somewhere in a database that is specialized for the data researchers were using. If everything is on a common platform, the choices can be streamlined and reported.
Through the OMOP experiment, Suchard and others developed a cross-platform analysis base to run thousands of studies using open-source methods. The results of these studies are fully integrated with vignettes, from the first step of data extraction all the way to publication. He noted that doing this makes everything reproducible both objectively (can the study be reproduced exactly?) and subjectively (can a researcher make minor changes to the study?).
Subject-matter experts were then asked to provide their best understanding of known associations and nonassociations between drugs and adverse events. The experiment now has about 500 such statements about “ground truth,” which provide some information on the null distribution of any statistic under any method. Suchard noted that this is important to counter some of the confounders that cannot be controlled in observational studies.
Database heterogeneity can be a problem for reproducible results even when everything else can be held constant, according to Suchard. He explained that looking at different data sets could result in very different answers using the same methodology. Of the first 53 outcomes examined in the OMOP experiment, about half of them had so much heterogeneity that it would not be advisable to combine them, and 20 percent of them had at least one data source with a significant positive effect and one with a significant negative effect.
Suchard then discussed what happens to study results when the data and design are held constant but design choices are varied. He explained that just a small change in a design choice could have a large impact on results.
Another issue is that most observational methods do not have nominal statistical operating characteristics for estimates, according to Suchard. Using the example of point estimates of the relative risk for a large number of negative controls (Ryan et al., 2013), he explained that if the process is modeled correctly and the confidence intervals are constructed using asymptotic normality assumptions, the estimates should have 95 percent coverage over the true value. However, this
proved not to be the case and the estimated coverage was only about 50 percent with both negatively and positively biased estimates. He explained that there needs to be a way to control for the many factors affecting bias so that the point estimates at the end are believable.
When assessing which types of study designs tend to perform best in the real world, Suchard suggested one way to look at the overall performance of the method is to think of it as a binary classifier (Ryan et al., 2013). If a given data source and an open black box analyzer produced either a positive or negative association, the result could be compared with the current set of ground truths and, if these estimates appear to be true, the area under the operator-receiver curve for that binary classification gives the probability of being able to identify a positive result as having a higher value than a negative result. Suchard noted that a group of self-controlled methods that compare the rate of adverse events when an individual is and is not taking a drug tend to perform better in these data with this ground truth than the cohort methods.
Suchard also noted that empirical calibration could help restore interpretation of study findings, specifically by constructing a Bayesian model to estimate what the distribution of the normal values might be using a normal mixture model (Schuemie et al., 2013). The p-value can then be computed as the error under the predicted distribution under the null hypothesis for the observed relative risk. The relative risk versus standard error can then be computed.
To address the findings that observational data are heterogeneous and methods are not reliable (Madigan et al., 2014), the public-private partnership Observational Health Data Sciences and Informatics12 group was established to construct reproducible tools for large-scale observational studies in health care. This group consists of more than 80 collaborators across 10 countries and has a shared, open data model (tracking more than 600 million people) with open analysis tools.
A participant wondered if medical research could be partitioned into categories, such as randomized clinical trials and observational studies, to assess which is making the most advances in medicine. Suchard pointed out that both randomized control trials and observational studies have limitations; the former is expensive and can put patients at risk, but the latter can provide misinformation. Balancing the two will be important in the future.
Courtney Soderberg, Center for Open Science
Courtney Soderberg began her presentation by giving an overview of the Center for Open Science’s recent work to assess the level of reproducibility in
___________________
12 The Observational Health Data Sciences and Informatics website is http://www.ohdsi.org/, accessed January 12, 2016.
experimental research, predict reproducibility rates, and change behaviors to improve reproducibility. The Many Labs project brought together 36 laboratories to perform direct replication experiments of 13 studies. While 10 out of 13 studies were ultimately replicable, Soderberg noted a high level of heterogeneity among the replication attempts. There are different ways to measure reproducibility, she explained, such as through p-value or effect sizes, but if there is a high level of heterogeneity in the chosen measure, it may indicate a problem in the analysis such as a missed variable or an overgeneralization of the theory. This knowledge of the heterogeneity can in itself move science forward and help identify new hypotheses to test. However, Soderberg noted that irreproducibility caused by false positives does not advance science, and the rate of these must be reduced. In addition, research needs to be more transparent so exploratory research is not disguised as confirmatory research. As noted by Simmons et al. (2011), the likelihood of obtaining a false-positive result when a researcher has multiple degrees of freedom can be as high as 60 percent. A large population of researchers admits to engaging in questionable research behavior, but many define it as research degrees of freedom and not as problematic (John et al., 2012).
Soderberg explained that a good way to assess reproducibility rates across disciplines is through a large-scale reproducibility project, such as the Center for Open Science’s two reproducibility projects in psychology13 and cancer biology.14 The idea behind both reproducibility projects is to better understand reproducibility rates in these two disciplines. In the case of Reproducibility Project: Psychology, 100 direct replications are conducted to determine the reproducibility rate of papers from 2008 and analyze findings to understand what factors can predict whether a study is likely to replicate (Open Science Collaboration, 2015).15 Moving forward, the project aims to look at initiatives, mandates, and tools to change behaviors. For example, Soderberg explained that open data sharing and study preregistration are not traditional practices in psychology, but the proportion of studies engaging in both has been increasing in recent years. Conducting future reproducibility projects can determine whether behavior change has resulted in higher levels of reproducibility.
John Ioannidis, Stanford University
In his presentation, John Ioannidis noted that a lack of reproducibility could often be attributed to the typical flaws of research:
___________________
13 The Center for Open Science’s reproducibility project in psychology website is https://osf.io/ezcuj/wiki/home/, accessed January 12, 2016.
14 The Center for Open Science’s reproducibility project in cancer biology website is https://osf.io/e81xl/wiki/home/, accessed January 12, 2016.
15 Results reported subsequent to the time of the February 2015 workshop.
- Solo, siloed investigator;
- Suboptimal power due either to using small sample sizes or to looking at small effect sizes;
- Extreme multiplicity, with many factors that can be analyzed but often not taken fully into account;
- Cherry-picking, choosing the best hypothesis, and using post-hoc selections to present a result that is exciting and appears to be significant;
- p-values of p < 0.05 being accepted in many fields, even though that threshold is often insufficient;
- Lack of registration;
- Lack of replication; and
- Lack of data sharing.
He explained that empirical studies in fields where replication practices are common suggest that most effects that are initially claimed to be statistically significant turn out to be false positives or substantially exaggerated (Ioannidis et al., 2011; Fanelli and Ioannidis, 2013). Another factor that complicates reproducibility is that large effects, while desired, are not common in repeated experiments (Pereira et al., 2012).
Ioannidis explained that a problem in making discoveries is searching for causality among correlation; the existence of large data sets is making this even more difficult (Patel and Ioannidis, 2014). He proposed that instead of focusing on a measure of statistical inference, such as a p-value, adjusted p-value, normalized p-value, or a base factor equivalent, it may be useful to see where an effect size falls in the distribution of the effect size seen across a field in typical situations.
Another possibility with large databases is to think about how all available data can be analyzed using large consortia and conglomerates of data sets. This would allow for an analysis of effects and correlations to see if a specific association or correlation is more or less than average. However, Ioannidis cautioned that even if a particular correlation is shown, it is difficult to tell if the effect is real. Another approach to the problem is trying to understand the variability of the results researchers want to get when using different analytical approaches (Patel et al., 2015). Ioannidis suggested that instead of focusing on a single result, communities should reach a consensus on which essential choices enter into one analysis and then identify the distribution of results that one could obtain once these analytical choices are taken into account.
He concluded by discussing the potential for changing the paradigms that contribute to irreproducibility problems. He asserted that funding agencies, journals, reviewers, researchers, and others in the community need to identify what goals should be given highest priority (e.g., productivity, quality, reporting quality, reproducibility, sharing data and resources, translational impact of applied research)
and reward those goals appropriately. However, Ioannidis admitted that it is an open question how such goals can be operationalized.
Q&A
A participant congratulated Suchard on his important work in quantifying uncertainty associated with different designs in pharmacoepidemology applications but wondered to what extent the methodology is extendable to other applications, including to studies that are not dependent on databases. Suchard explained that it is difficult to know how to transfer the methodology to studies that are not easy to replicate, but there may be some similarities in applying it to observational studies with respect to comparative effectiveness research. He noted that the framework was not specifically designed for pharmacoepidemology; it was just the first problem to which it was applied.
A participant asked Ioannidis whether—if one accepts that the false discovery rate is 26 percent using a p-value of 0.05—a result with a p-value of 0.05 should be described in papers as “worth another look” instead of “significant”? Ioannidis responded that it would depend on the field and what the performance characteristics of that field would look like, although most fields should be skeptical of a p-value close to 0.05. Recently, he and his colleagues started analyzing data from a national database in Sweden that includes information on 9 million people. Using this database, they are able to get associations with p-values of 10−100; however, he speculates that most are false associations, with such small p-values being possible due to the size of the data set. Similarly for pharmacoepidemology, he stated that almost all drugs could be associated with almost any disease if a large enough data set is used.
A participant asked if there are data about the increase in the number of problematic research papers. Soderberg observed that Simmons et al. (2011) ran simulations looking at how researcher degrees of freedom and flexibility in the analytic choice can cause false positives to go as high as 61 percent. She explained that it is difficult to tell if the real literature has such a high false-positive rate because it is unclear how many degrees of freedom, and how much analytic flexibility, are faced by researchers. However, surveys suggest that it is quite common to make such choices while executing research, and she believes the false-positive rates are higher than the 5 percent that a p-value of 0.05 would suggest. She noted that it is difficult to tell how behavior has evolved over time, so it is hard to measure how widespread reproducibility issues have been in the past.
A participant noted that much of Ioannidis’s work is centered on studies with a relatively small sample size, while Suchard’s work deals with tens of millions or hundreds of millions of patients, and wondered if there needs to be a fundamental rethinking on how to interpret statistics in the context of data sets where sampling variability is effectively converging to epsilon but bias is still persistent in the
analyses. Ioannidis agreed that with huge sample sizes, power is no longer an issue but bias is a major determinant. He suggested that many potential confounders be examined.
REPRODUCIBILITY FROM THE INFORMATICS PERSPECTIVE
Mark Liberman, University of Pennsylvania
Mark Liberman began his presentation by giving an overview of a replicability experiment using the “common task method,” which is a research paradigm from experimental computational science that involves shared training and testing data; a well-defined evaluation metric typically implemented by a computer program managed by some responsible third party; and a variety of techniques to avoid overfitting, including holding test data until after the bulk of the research is done. He explained that the setting for this experiment is an algorithmic analysis of the natural world, which often falls within the discipline of engineering rather than science, although many areas of science have a similar structure.
There are dozens of current examples of the common task method applied in the context of natural language research. Some involve shared task workshops (such as the Conference on Natural Language Learning,16 Open Keyword Search Evaluation,17 Open Machine Translation Evaluation,18 Reconnaissance de Personnes dans les Émissions Audiovisuelles,19 Speaker Recognition Evaluation,20 Text REtrieval Conference,21 TREC Video Retrieval Evaluation,22 and Text Analysis Conference23) where all participants utilize a given data set or evaluation metrics to produce a result and report on it. There are also available data sets, such as the Street View House Numbers,24 for developing machine-learning and object-recognition algorithms
___________________
16 The Conference on Natural Language Learning website is http://ifarm.nl/signll/conll/, accessed January 12, 2016.
17 The Open Keyword Search Evaluation website is http://www.nist.gov/itl/iad/mig/openkws.cfm, accessed January 12, 2016.
18 The Open Machine Translation Evaluation website is http://www.nist.gov/itl/iad/mig/openmt.cfm, accessed January 12, 2016.
19 The Reconnaissance de Personnes dans les Émissions Audiovisuelles website is https://tel.archives-ouvertes.fr/tel-01114399v1, accessed January 12, 2016.
20 The Speaker Recognition Evaluation website is http://www.itl.nist.gov/iad/mig/tests/spk/, accessed January 12, 2016.
21 The Text REtrieval Conference website is http://trec.nist.gov, accessed January 12, 2016.
22 The TREC Video Retrieval Evaluation website is http://trecvid.nist.gov, accessed January 12, 2016.
23 The Text Analysis Conference website is http://www.nist.gov/tac/, accessed January 12, 2016.
24 The Street View House Numbers website is http://ufldl.stanford.edu/housenumbers/, accessed January 12, 2016.
with minimal requirement on data preprocessing and formatting. In the Street View House Numbers case, there has been significant improvement in the performance, especially in terms of reducing the error rate using different methods (Netzer et al., 2011; Goodfellow et al., 2013; Lee et al., 2015).
Liberman pointed out that while the common task method is part of the culture today, it was not so 30 years ago. In the 1960s, he explained, there was strong resistance to such group approaches to natural language research, the implication being that basic scientific work was needed first. For example, Liberman cited Languages and Machines: Computers in Translation and Linguistics, which recommended that machine translational funding “should be spent hardheadedly toward important, realistic, and relatively short-range goals” (NRC, 1966). Following the release of that report, U.S. funding for machine translational research decreased essentially to zero for more than 20 years. The committee felt that science should precede engineering in such cases.
Bell Laboratories’ John Pierce, the chair of that National Research Council study, added his personal opinions a few years later in a letter to the Journal of the Acoustical Society of America:
A general phonetic typewriter is simply impossible unless the typewriter has an intelligence and a knowledge of language comparable to those of the native speaker of English. . . . Most recognizers behave, not like scientists, but like mad inventors or untrustworthy engineers. The typical recognizer gets into his head that he can solve “the problem.” The basis for this is either individual inspiration (the mad “inventor” source of knowledge) or acceptance of untested rules, schemes, or information (the untrustworthy engineer approach) . . . The typical recognizer . . . builds or programs an elaborate system that either does very little or flops in an obscure way. A lot of money and time are spent. No simple, clear, sure knowledge is gained. The work has been an experience, not an experiment. (Pierce, 1969)
While Pierce was not referring to p-values or replicability, Liberman pointed out that he was talking about whether scientific progress was being made toward a reasonable goal.
The Defense Advanced Research Projects Agency (DARPA) conducted its Speech Understanding Research Project from 1972 through 1975, which used classical artificial intelligence to improve speech understanding. According to Liberman, this project was viewed as a failure, and funding ceased after 3 years. After this, there was no U.S. research funding for machine translation or automatic speech recognition until 1986.
Liberman noted that Pierce was not the only person to be skeptical of research and development investment in the area of human language technology. By the 1980s, many informed American research managers were equally skeptical about the prospects, and relatively few companies worked in this area. However,
there were people who thought various sorts of human language technology were needed and that the principle ought to be feasible. In the mid-1980s, according to Liberman, the Department of Defense (DoD) began discussing whether it should restart research in this area.
Charles Wayne, a DoD program manager at the time, proposed to design a speech recognition research program with a well-defined objective evaluation metric that would be applied by a neutral agent, the National Institute of Standards and Technology (NIST), on shared data sets. Liberman explained that this approach would ensure that “simple, clear, sure knowledge” was gained because it would require the participants to reveal their methods at the time the evaluation was revealed. The program was set up accordingly in 1985 (Pallett, 1985).
From this experiment the common task structure was born, in which a detailed evaluation plan is developed in consultation with researchers and published as a first step in the project. Automatic evaluation software (originally written and maintained by NIST but developed more widely now) was published at the start of the project to share data and withhold evaluation test data. Liberman noted that not everyone liked this idea, and many people were skeptical that the research would lead anywhere. Others were disgruntled because the task and evaluation metrics were too explicit, leading some to declare this type of research undignified and untrustworthy.
However, in spite of this resistance and skepticism, Liberman reported that the plan worked. And because funders could measure progress over time, this model for research funding facilitated the resumption of funding for the field. The mechanism also allowed the community to methodically surmount research challenges, in Liberman’s view, because the evaluation metrics were automatic and the evaluation code was public. The same researchers who had objected to being tested twice a year began testing themselves hour by hour to try algorithms and see what worked best. He also credited the common task method with creating a new culture where researchers exchanged methods and results on shared data with a common metric. This participation in the culture became so valuable that many research groups joined without funding.
As an example of a project that used the common task method, Liberman described a text retrieval program in 1990 in which four contractors were given money for working on a project. Because of the community engagement across the field, 40 groups from all around the world participated in the evaluation workshop because it offered them an opportunity to access research results for free.
An additional benefit of the common task method is that it creates a positive feedback loop. Liberman explained that when every program has to interpret the same ambiguous evidence, ambiguity resolution becomes a gambling game that rewards the use of statistical methods. That, in turn, led to what has become known as machine learning. Liberman noted that, given the nature of speech and language,
statistical methods need large training sets, which reinforces the value of shared data. He also asserted that researchers appreciated iterative training cycles because they seem to create simple, clear, and sure knowledge, which motivated participation in the common-task culture.
Over the past 30 years, variants of this method have been applied to many other problems, according to Liberman, including machine translation, speaker identification, language identification, parsing, sense disambiguation, information retrieval, information extraction, summarization, question answering, optical character recognition, sentiment analysis, image analysis, and video analysis. Liberman said that the general experience is that error rates decline by a fixed percentage each year to an asymptote, depending on task and data quality. He noted that progress usually comes from many small improvements and that shared data play a crucial role because they can be reused in unexpected ways.
Liberman concluded by stating that while science and engineering cultures vary, sharing data and problems lowers the cost of the data entry, creates intellectual communities, and speeds up replication and extension.
A participant commented that there is a debate about whether subject matter experts can cooperate with computational scientists and statisticians, particularly at this point in time when some statistical methods are outperforming expert rule-based systems. Liberman observed that DARPA essentially forced these communities to work together and managed to change the culture of both fields.
Micah Altman, Massachusetts Institute of Technology
To begin his presentation, Micah Altman surmised how informatics could improve reproducibility. He partitioned these thoughts into two categories: formal properties and systems properties. Formal properties, Altman explained, are properties of information flows and their management that tend to support reproducibility-related inferences. These include transparency, auditability, provenance, fixity, identification, durability, integrity, repeatability, self-documentation, and nonrepudiation. These properties can be applied to different stages and entities, as well as to components of the information system. Systems properties describe how the system interacts with users and what incentives and culture they engender. They include factors such as barriers to entry, ease of use, support for intellectual communities, speed and performance, security, access control, personalization, credit and attribution, and incentives for well-founded trust among actors. There is also the larger question of how the system integrates into the research ecosystem, which includes issues such as sustainability, cost, and the capability for inducing well-founded trust in the system and its outputs.
Altman noted that reproducibility claims are not formulated as direct claims about the world. Instead, they are formulated as questions such as the following:
- What claims about information are implied by reproducibility claims or issues?
- What properties of information and information flow are related to those claims?
- What would possible changes to information processing and flow yield? How much would they cost?
He noted that many elements of the system might be targeted to improve today’s situation. People are at the root of the ecosystem, where they affect decisions of theory; select and apply methods; observe and edit data; select, design, and perform analyses; and create and apply documents. The methods also interact, intervene, and simulate in the real world. The researcher’s information flow over time ranges among creation/collection, storage/ingest, processing, internal sharing, analysis, external dissemination/publication, reuse (scientific, educational, scientometric, and institutional), and long-term access. Altman reiterated the point made by many of the workshop’s speakers that there are many operational reproducibility claims and interventions, such as those outlined in Table 3.2, that are often referred to by the same or inverted names in different communities.
There is an overlap between social science, empirical methods, and statistical computation, according to Altman. In all cases there is a theory guiding the choice of how to conceptualize the world or conceptualize the approach, which leads to algorithms and protocols for solving a particular inference problem. Those protocols or algorithms are implemented in particular coding rules (e.g., instrumentation design and software) and are executed to produce details and context. Informatics tools can be used in a number of environments to deal with information flow, depending on what properties and reproducibility claims are being examined.
TABLE 3.2 Some Types of Reproducibility Issues and Use Cases
| Common Labels | Reproducibility Related Issue | Example Interventions |
|---|---|---|
| Misconduct, bit rot, author responsibility | Data were fabricated, corrupted, or radically misinterpreted prior to analysis |
Discipline/community data archives. NIH genomic data sharing policy
RetractionWatch; collaborative data collection projects |
| Misconduct, negligence, confusion, typo, proofreader error, dynamic data problem, versioning problem | Data (referenced by identifier, provided as an instance, described by method) have nontrivial set of semantic differences from that used as input to the publication | Dat, DataHub, DataVerse (versioning) |
| Common Labels | Reproducibility Related Issue | Example Interventions |
|---|---|---|
| Misconduct, negligence, harmless error | Published analysis algorithm does not correspond to implemented analysis | S/Weave; Compendia; Vistrails |
| Reproducibility [NSF, 2014; Buckheit and Donoho, 1995] | Variance of estimates given data instance and analysis implementation | Journal replication data and code archives. |
| Replicability [King, 1995] | Virtual machine archiving | |
|
Replication [NSF, 2014]
Reproducibility [King, 1995] independent replication |
Variance of estimates given method algorithm and analysis algorithm | Protocol archive, Journal of Visual Experiments |
| Result validation, fact checking | Variance of estimates given data identifier and analysis algorithm | Data citation standards |
| Calibration, extension, reuse | Produce new analysis given data identifier | Data archives |
| File drawer problem | Publisher bias toward significant (or expected) results | APS preregistration badge, Journal of Null Results |
| Underreporting (adverse events); data dredging (multiple comparisons) | Author bias toward publishing favored outcomes | Clinical trial preregistration |
| Data dredging: multiple comparisons; p-hacking | Author bias to creating significant results resulting in difference between stated method/analysis and actual (complete) method/analysis | Holdout data escrow |
| Sensitivity, robustness | Variance of support for claims across specification change | Sensitivity analysis |
| Reliability | Variance of support for claims across repeated measures, samples | Meta-analysis; Cochrane review |
| Data integration | ||
| Generalizability | Variance of support for claims across different frames | Cochrane review |
| Laws, truth | Variance of support for claims to other populations | Grand challenge? |
SOURCE: Courtesy of Micah Altman, Massachusetts Institute of Technology, presentation to the workshop.
To conclude, Altman posed the following questions about how to better support reproducibility with information infrastructure:
- How can we better identify the inferential claims implied by a specific set of (non)reproducibility claims and issues?
- Which information flows and systems are most closely associated with these inferential claims?
- Which properties of information systems support generating these inferential claims?
Altman also thanked his collaborators, namely Kobbi Nissim, Michael Bar-Sinai, Salil Vadhan, Jeff Gill, and Michael P. McDonald, and provided the following references to related work: Allen et al. (2014); Altman and Crosas (2013); Garnett et al. (2013); Altman et al. (2001, 2011); Altman (2002, 2008); Altman and King (2007); Altman et al. (2004); and Altman and McDonald (2001, 2003).

































