In this session, panelists considered the financial aspects of genomics-based programs, including demonstrating value and the return on investment of screening programs. Bradford Powell, an assistant professor in the Department of Genetics at the University of North Carolina at Chapel Hill, provided an overview of the economic issues related to the implementation of genomics-based screening programs. Josh Peterson, an associate professor of biomedical informatics and medicine at Vanderbilt University Medical Center, described two ongoing programs to illustrate the drivers of value for pharmacogenomics panel testing. Dean Regier, an assistant professor at the University of British Columbia, discussed the concept of personal utility from his perspective as a health economist.
CLINICAL COSTS AND EFFECTS OF GENOMIC SCREENING
The difference between diagnostic testing and screening becomes important when considering the potential value of a genomic test. Diagnostic testing is performed for an individual who either has or is suspected of having a particular disorder because of clinical symptoms, Powell said. The prior probability that the person has the condition is relatively high, and testing can inform treatment or expectations of prognosis. In contrast, Powell said, screening is a population-based method for identifying persons with a condition or predisposition to a condition when the prior probability of having that condition is low. Screening may “inflict” health care on apparently healthy individuals who might never become sick with whatever condition is being screened for, he said.
Opportunistic screening occurs when a patient comes to a clinic for specific testing (e.g., pharmacogenomic variant testing for warfarin) and is tested at the same time for several high-yield variants related to common complex diseases (e.g., familial hypercholesterolemia, hereditary breast and ovarian cancer, and Lynch syndrome). Although opportunistic screening itself has a relatively low marginal cost, Powell said, there are challenges with generalizing it to population-level screening. The American College of Medical Genetics and Genomics (ACMG) has suggested that providers who offer genomic screening should provide pretest counseling, and it has developed a list of highly actionable gene variants for which it recommends reporting of incidental findings (Green et al., 2013). However, Powell said, it is not possible to provide the same degree of pretest counseling on a
population level with the currently available workforce. There are also questions about how the estimated penetrance of these mutations will hold up against the ascertainment bias under which they were initially described.
When considering screening for additional conditions with new technologies, it is worth revisiting the screening criteria developed by Wilson and Jungner (1968). Powell presented those criteria reorganized by the characteristics of the condition, the characteristics of the case finding, and the characteristics of the health care system (see Box 3-1). By weighing the criteria listed above, it is possible to determine if it makes sense to implement screening for a given condition.
When Should Genomic Screening Be Performed?
Screening is most effective when performed prior to the age at which a condition’s symptoms are likely to appear or the age at which the earliest stage of treatment can begin, Powell said. Prenatal screening—or even preconception screening—might be maximally effective with regard to actionability, he said, as it creates the opportunity to inform reproductive decision making. Because of the ethical quandaries that come with
newborn (or earlier) genomic screening, it has been suggested that certain conditions should not be screened for until adulthood. Powell explained that according to the current ethical framework within clinical genetics, in the case of conditions with adult onset or for which an intervention is not started until adulthood, screening should be deferred and the child’s future autonomy maintained (Ross et al., 2013). However, screening for different conditions may be indicated at different times throughout a person’s life, and the financial feasibility of screening for genomic information may be different at different points in time. One potential solution, Powell said, is to “sequence first and ask questions later,” obtaining the genetic sequence that can then be queried over time. Taking this approach raises questions of how the costs would be handled (initially, and over time). For example, relatively low-risk individuals may not benefit from genomic screening for a long time, and, with the current health care system, payers do not have a long enough time horizon with a given patient to benefit financially from the screening.
When Should Results Be Returned?1
When considering genomic screening in children, the issue of genetic exceptionalism2 becomes particularly relevant, Powell said. Genomic screening in children involves proxy decision making by the parents on behalf of the child. As mentioned above, there are concerns about preserving the child’s future autonomy. Deferring screening for adult onset conditions is one approach, but some patients and stakeholders have pushed back on this, Powell said. Concerns expressed include: What if this is the only genomic screening the child gets? What if testing indicates the parent might be at risk for a treatable, adult onset condition (e.g., hereditary breast and ovarian cancer)?
The question of what results should be returned when healthy infants undergo genomic screening has been incorporated as part of the North Carolina Newborn Exome Sequencing for Universal Screening (NC Nexus) Study.3 This is being done in a controlled environment because of the potential risks, including financial risks, that might be carried by the children
___________________
1 The National Academies of Sciences, Engineering, and Medicine has convened a committee to examine the return of individual-specific research results generated in research laboratories. For more information about the committee and consensus study, see http://nationalacademies.org/hmd/Activities/Research/ResearchResultsGeneratedinResearchLaboratories.aspx (accessed January 18, 2018).
2Genetic exceptionalism is the concept that genetic information should be treated differently than other medical information and deserves special privacy protections (Rothstein, 2005).
3 For more information about the NC Nexus Study, see https://www.med.unc.edu/genetics/berglab/Research/nc-nexus-project (accessed January 18, 2018).
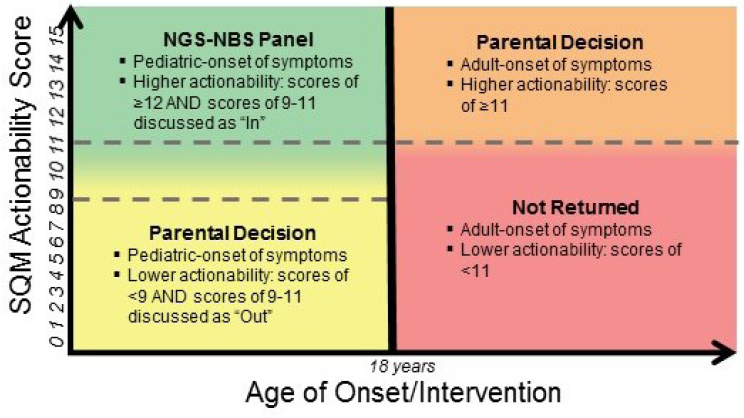
NOTE: NGS-NBS = next-generation sequencing–newborn screening; SQM = semi-quantitative metric.
SOURCE: Bradford Powell, National Academies of Sciences, Engineering, and Medicine workshop presentation, November 1, 2017.
over their lifetimes, Powell said. For a given childhood condition or gene variant, the actionability score (a semi-quantitative metric that weighs the severity of the disease, the likelihood of the outcome, the efficacy of a given intervention, the acceptability of the intervention, and the knowledge base supporting the disease and intervention [Powell, 2016]) is considered relative to the age of onset or intervention of the condition (see Figure 3-1). If a result is considered to be actionable within childhood, that condition/gene variant could potentially be included in a next-generation genomic sequencing newborn screening panel. Another part of the study randomizes parents into two groups which are then asked to decide whether they want to receive genomic screening information: those parents whose child screens positive for a pediatric-onset condition with a low actionability score, and those whose child screens positive for an adult-onset condition with a higher actionability score.4
___________________
4 For more information about parental decision making about newborn genomic screening, see http://pediatrics.aappublications.org/content/137/Supplement_1/S16 (accessed January 18, 2018).
Genomic screening in adults involves different issues, in part because adults make their own decisions. The GeneScreen project5 at the University of North Carolina at Chapel Hill is looking at the best ways to provide genomic screening to the general adult population. The study is designed to screen for a smaller subset of genes that are expected to provide the majority of health benefits across the population (including 17 genes for 11 conditions), Powell said, noting that when analyzing the results of the study, it will be important to take into account the fact that the prevalence of these individual conditions is low and there are concerns about the false discovery rate. The most common conditions among the 59 medically actionable genes listed by the ACMG have incomplete penetrance, Powell added. When prevalence is low in the general population, the number of false positive results can be greater than the number of true positive results. For example, if a condition is present in 1 in 10,000 individuals, and the screening test is 99 percent sensitive and 99.94 percent specific, it would be expected to find 699 positive screens in 1 million people. One hundred people will actually have the condition, and the sensitivity of the assay means that of those people, 99 will have a positive result, and 1 person will have a false negative result. Even though this is a very specific screen, and the vast majority of the 999,900 individuals who are negative will appropriately screen negative, 600 negative individuals will screen falsely positive (i.e., for each true positive result, there will be 6 false positives). Those individuals will receive medical care that they do not need.
For metabolic conditions in the newborn screen (e.g., phenylketonuria), secondary testing is performed to confirm positive results before any intervention. However, such secondary testing is not possible for many of the conditions that might be considered for a genomic screening panel, Powell said. Importantly, when “likely pathogenic” screening results have a 90 to 95 percent probability of being truly pathogenic, the screen is identifying people who would not have needed treatment (Plon et al., 2008; Richards et al., 2015).
Who Pays and Who Benefits?
Because health care in the United States uses a third-party payer system, there are different potential incentives to participate in genomic screening programs. Ideally, genomic screening programs should result in better patient care. There are also potential conflicts of interest to keep in mind, Powell said. For example, leaders at a health care system might be interested in identifying people at risk for colon cancer because the health care
___________________
5 For more information about the GeneScreen project, see http://genomics.unc.edu/genomicsandsociety/GeneScreen.html (accessed January 24, 2018).
system may benefit from performing additional colonoscopies. A pharmaceutical company might benefit from individuals being identified as being at higher risk for a certain side effect because those individuals might be prescribed a more costly agent. Whether these actions have a net benefit to patients or society depends on the prevalence of the condition and on how severe or frequent the adverse reaction would be, Powell said.
Drivers of Cost
There are three elements that drive the overall cost of genomic testing, Powell said: direct costs, downstream costs, and ancillary costs. At this stage, there are many hypotheses about the utility and applicability of genomic testing, but not enough data. Of the three types of costs, the direct costs (which include the cost of assays, analysis, and return of results) are the best characterized to date, although the cost of returning results is likely to change as current genomic screening efforts expand to population-level screening. Downstream costs include provider education, confirmatory testing, interventions and surveillance, and complications of interventions or surveillance. Ancillary costs include false reassurance (misunderstanding of information which may lead to people not seeking indicated care), interventions or surveillance in response to clinical false positive screens, patient anxiety or discomfort, and effects on insurance or employment. Powell noted that these ancillary costs can be quantified as quality-adjusted life years (QALYs) for comparison across different potential interventions.
Unknowns
There are still unknowns to be clarified before the balance of costs and benefits can be fully understood, Powell said. Unknowns include the prevalence of the conditions (the proportion of a population that has—or had—a specific condition in a given time period) (NIMH, 2017); the penetrance of the conditions (the percentage of individuals with a given genotype who exhibit the phenotype associated with that genotype) (Griffiths et al., 2000); and the efficacy of pre-symptomatic intervention, that is, how much of a difference identifying people at risk really makes. Until these variables are better characterized for each condition, there is a risk of treating people that do not need treatment, Powell said. In closing, he stressed the importance of being transparent and making sure that the patients and populations served by health care systems fully understand when screening is used to further research and when it is intended for clinical testing.
THE VALUE OF PHARMACOGENOMIC PANEL TESTING
To illustrate the drivers of value for pharmacogenomics panel testing, Peterson described two Vanderbilt University programs, a pharmocogenomic screening program integrated with the electronic health record (EHR) system called the Pharmacogenomic Resource for Enhanced Decisions in Care Treatment (PREDICT)6 and an accompanying cost-effectiveness study designed to determine the long-term value of pharmocogenomic panel testing, Rational Integration of Genomic Healthcare Technology (RIGHT).7
Multiplex Pharmacogenomic Screening Panel
The typical testing approach at Vanderbilt University Medical Center has been evolving from serial, single gene testing to multiplexed panel testing. Multiplexed panels offer economies of scale, Peterson said, and the cost of assaying additional genetic variants approaches zero. Panels also broaden the opportunities to perform testing: preemptive screening (before clinically indicated) becomes feasible as well as reactive testing. Importantly, with preemptive testing clinicians do not need to remember to order each individual genetic test since the results of the panel are embedded in the EHR. This was one of the drivers of institutional investment in this area, Peterson said. On the other hand, he continued, clinicians may not want the responsibility of dealing with the additional patient data from pharmacogenomics testing. There is some additional cost associated with panel testing, both for the assay itself and for data management downstream, and any benefits are accrued in the future (when the test will presumably be cheaper and better). Peterson also noted that the genetic data might never be used if patients are not prescribed a relevant medication. There is also concern, especially by payers, about unintended or unwarranted costs related to cascade testing.
Common clinical scenarios motivated Vanderbilt’s program for multiplexed pharmacogenomics screening, Peterson said; that program is integrated with the EHR as a way to provide opportunities to use a patient’s genomic data over time. For example, a patient in the health system who has risk factors for coronary disease can be preemptively screened with the panel test and the results saved in his or her EHR. Later, if the patient develops coronary artery disease, he or she might receive percutaneous coronary intervention (a stent) and be prescribed antiplatelet therapy and statin therapy, which can be tailored according to the pharmacogenomic information in the EHR. Perhaps years later, the patient might develop
___________________
6 For more information about PREDICT, see https://www.mydruggenome.org (accessed January 17, 2018).
7 For more information about RIGHT, see http://rightsim.org or http://www.nber.org/papers/w24134 (accessed January 22, 2018).
atrial fibrillation and need anticoagulants, which can again be informed by the pharmacogenomic data.
Patients can enter the Vanderbilt PREDICT program through targeted preemptive screening or as a result of reactive testing. Preemptive screening is not reimbursed by payers, Peterson said, and the institution must cover the costs. In the case of reactive testing, there is a clinical indication and an ICD-9 code that can be used for billing purposes, and payers will cover the costs of at least one of the components on the panel. Once a patient is genotyped, the results are entered into the EHR, which includes clinical decision support for identified genetic risk variants. An example of using this information is the best practice alert, which interrupts the e-prescribing process and alerts the prescriber that the patient has a gene variant associated with an adverse reaction or variability of response for the drug being prescribed. The alert also includes recommendations for treatment modifications.
PREDICT relies on frontline clinicians to explain pharmacogenomic test results to their patients; however, there are not enough genetic counselors to be able to intervene with everyone during the prescribing process, Peterson said. To help with this, patients are also informed of their pharmacogenomic results through the patient portal, which provides high-level information about their pharmacogenetic test results and about how their genes affect their medications.
Determining the Value of Pharmacogenomic Testing
There are several lessons that have informed the economic modeling of pharmacogenomic screening, Peterson said.
- Cost is a concern. Cost does matter to clinicians, especially the expectation of reimbursement and the out-of-pocket costs for the patients they care for. There is also concern about the overall cost to the health system for something that is new and relatively unfamiliar.
- Strength of evidence and guidelines matter. Providers are particularly interested in guidelines from their clinical specialty. Guidelines from genetic societies have not yet been incorporated into guidelines from other specialties.
- Clinical behavior is diverse. Pharmacogenomic screening data are not deterministic. There are many reasons why providers might not follow the advice in the EHR alert. However, retrospective analysis shows that pharmacogenomic data do change prescribing substantially, with between 30 and 60 percent of prescriptions modified based on variants reported (Peterson et al., 2016).
Discrete event simulation is used to model indication (how often pharmacogenomic data would be used) and outcome (what the benefit of using those data would be), comparing genotyped and non-genotyped populations.8 Peterson’s group at Vanderbilt has created a model that incorporates all 46 drug–gene interactions on the level A list compiled by the Clinical Pharmacogenetics Implementation Consortium (CPIC).9 To fully model cost effectiveness, Peterson said, it would be necessary to create a model for each individual interaction—all 46 of them. To simplify the simulation, the interactions were grouped into 7 categories according to frequency of prescribing, frequency of the adverse event, and severity of the adverse event, and 7 different models were created instead of 46.
In a simple genotype-tailored therapy model, Peterson explained, simulated patients receive the primary treatment or an alternate treatment, based on genotyping results. As in a real-life scenario, a certain number of adverse events are expected with the primary treatment. The simulated patients are followed through their lifetimes to death. The model relies on several base assumptions: that pharmacogenomics-guided therapy costs threefold more; pharmacogenomics guidance conveys 0.70 relative risk of adverse events; if not preemptively screened, a genetic test is ordered 50 percent of the time; and any genetic information obtained upstream is used 75 percent of the time. These are optimistic assumptions, Peterson said, and the simulation is run millions of times to determine what is driving the economic results.
Different strategies (no genotyping, reactive serial single-gene sequencing, preemptive panel, and reactive panel) have different likelihoods of being cost effective, given a certain willingness to pay for an extra QALY, Peterson said. Figure 3-2 depicts the projected likelihood for each scenario. An example of a procedure with a very high incremental cost-effectiveness ratio (ICER) is a left ventricular assist device, where the ICER exceeds $500,000; an example of a low incremental cost-effectiveness ratio is a colonoscopy, where patients gain an extra QALY for every $25,000 spent on the intervention. In this analysis, preemptive genotyping has a 50 percent chance of being cost effective relative to a willingness to pay of $150,000, although many groups set willingness to pay thresholds at $50,000 or $100,000, Peterson shared. Reactive serial single-gene testing
___________________
8 For more detailed information, see “The Value of Genomic Pharmacogenomic Information,” a working paper to be included in a forthcoming conference proceedings of the National Bureau of Economic Research. Available at http://www.nber.org/chapters/c13989 (accessed January 3, 2018).
9 CPIC Level A indicates that genetic information should be used to change prescribing of an affected drug. To be defined as CPIC Level A, the preponderance of evidence must be high or moderate in favor of changing prescribing. For more information about the level definitions for CPIC gene–drug pairs see https://cpicpgx.org/prioritization/#flowchart (accessed February 8, 2018).
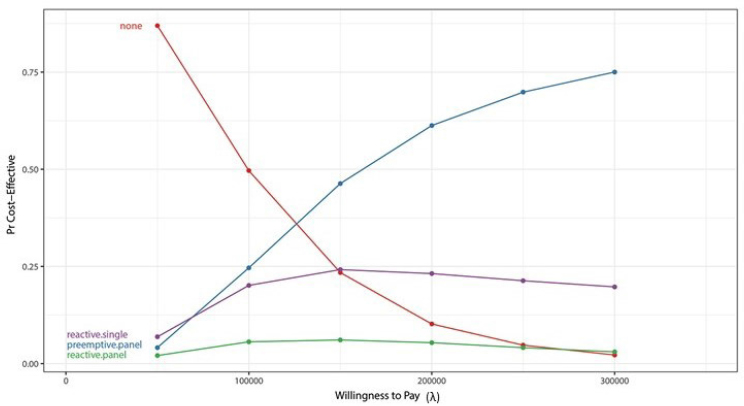
NOTES: Pr = plausible range. Willingness to pay (x axis) is from the decision makers’ perspective.
SOURCES: Joshua Peterson, National Academies of Sciences, Engineering, and Medicine workshop presentation, November 1, 2017. Originally from Graves et al., 2017.
is somewhat cost effective at the low end of willingness to pay, but cost effectiveness levels off and then decreases as willingness to pay increases. Reactive panel testing has a low probability of cost effectiveness across the spectrum of willingness to pay. A sensitivity analysis that plots the probability that pharmacogenomic information is used versus risk reduction from use of the pharmacogenomic-guided therapy is another way to analyze cost effectiveness of multiplexed testing strategies, Peterson said. If there is no reduction of the risk of an adverse event, then not testing is clearly preferred. If there is significant risk reduction, preemptive panel testing is the preferred option only if the probability that the clinician is going to use the data is high. If that probability is low, reactive single-gene testing is the better approach. Efforts to create implementation programs that are effective matter, Peterson said. As implementation approaches improve to a point where clinicians use pharmacogenomic data 100 percent of the time, preemptive panel testing becomes the optimal strategy.
Summary of Lessons Learned from the PREDICT Project
The key goals of the PREDICT project are to create a methodology with which to evaluate the use of pharmacogenomic information over a patient’s lifetime and assess the value of panel testing. There are a number of assumptions under which multiplexed pharmacogenomic screening is cost effective, Peterson said, but the time frame for accrual of benefits to achieve cost effectiveness may be many years, especially in relatively low-risk individuals. In the current health care system, payers do not work under time horizons that are that long. The analyses conducted thus far are a traditional approach to cost effectiveness; there may be other kinds of value that are not being captured, such as the value of patients knowing their genomic test results and being confident in the safety and efficacy of the prescriptions, which can affect adherence and engagement in care.
COLORECTAL CANCER SCREENING AND RETURN OF SECONDARY FINDINGS: A VALUE FRAMEWORK ANALYSIS
The value-for-money approach considers costs relative to health outcomes, Regier said, describing the reference cases for estimating value for money used in Canada, the United Kingdom, and the United States. A reference case is used to lay out the principles and methods that are appropriate for particular institutional objectives. In Canada and the United Kingdom, the objective of the analysis is to maximize health status gains subject to limited budgets. To do this, an ICER is calculated by dividing the incremental cost by the QALY and then compared to how much the decision makers are willing to pay for each QALY gained. The reference case in Canada and the United Kingdom is the health system perspective—that is, the perspective of those who manage the budgets and need to meet the objective of maximum health gains across the population. The recently published guidance from the Second Panel on Cost Effectiveness in Health and Medicine for the United States takes this analysis a step further, calling for the inclusion of the societal perspective as well as the health care system perspective (Neumann et al., 2017). To achieve a societal perspective, researchers may want to go beyond standard analyses to consider broader costs and broader concepts of what patients value in their health care.
Regier explained that health economists typically estimate QALYs using the EQ5D questionnaire, a standardized tool created by the EuroQol group for measuring health status. The EQ5D consists of a set of questions that asks patients to rate their quality of life in five domains: mobility, self-care, usual activities, pain/discomfort, and anxiety/depression. In the EQ-5D-3L, each dimension has three possible responses that indicate whether the patient has no problems, some/moderate difficulties, or extreme difficulties/-
incapacity in that dimension. These results are then calculated to inform the QALY, whose value lies between one, which is perfect health, and zero, which is death.10
Traditional measures of value and value for money in health economics are somewhat limited in their application to precision medicine and genomic technologies, Regier said. The value of precision medicine is dependent on the information that patients receive and the benefits that patients and providers ascribe to that information (Marshall et al., 2017). The Second Panel on Cost Effectiveness in Health and Medicine said that decision makers need a “quantification and valuation of all health and non-health effects of interventions, and to summarize those effects in a single quantitative measure” (Neumann et al., 2017, p. 370). The EQ5D estimate of QALY does not incorporate those non-health effects, Regier noted.
The Value of Knowing: Incorporating Preference-Based Utility
Regier said that from his perspective as an economist, value equals preference-based utility. In other words, the preferences of the individuals making decisions among alternative health care goods can inform the value of those goods. For precision medicine and genomics, Grosse and colleagues have said that personal utility is the utility that individuals and families ascribe to genomic information apart from health outcomes (Grosse et al., 2010). Personal utility enhances one’s sense of control, informs self-identity, and can resolve uncertainties surrounding an individual’s diagnosis and prognosis (Foster et al., 2009; Regier et al., 2009). The valuation of personal utility, Regier said, can be done using the stated preference discrete choice experiment method, which is based on an attribute-based measure of value that is based on random utility theory (McFadden, 1974). A discrete choice experiment is a well-established way of eliciting preferences for non-market health care products and programs and determining how individuals value various health states (Lancsar and Louviere, 2008).
Next-generation sequencing and secondary findings provide information on diseases that are not related to a patient’s current diagnosis, Regier said. For example, a patient might be tested for Lynch syndrome, with the secondary findings indicating a risk for long QT syndrome (which is treatable) and Alzheimer’s disease (for which there is no effective treatment). As discussed earlier in the day, the ACMG recommends the return of highly penetrant, clinically actionable results, without patient preference being fully taken into account; however, Regier said, the ACMG did not ask individuals what kind of information they wanted. In contrast, the Cana-
___________________
10 For more information about the EQ5D and the EuroQol Group, see https://euroqol.org/eq-5d-instruments (accessed January 18, 2018).
dian College of Medical Genetics does not endorse returning actionable secondary findings for monogenic conditions outside of the research context because of the high cost, both psychological and monetary.
In a discrete choice experiment, Regier and colleagues asked a representative population of 1,200 Canadians what kind of information they wanted from secondary findings, if such information could be provided (Regier et al., 2015). Participants were presented with a series of exercises, and each time they were asked to choose between two scenarios for receiving information that differed with regard to disease risk, treatability, severity, carrier status, and cost to the patient. An econometric behavioral model was used to predict preference-based utility, which informed the uptake of different policy scenarios for the return of results and individuals’ willingness to pay for the return of incidental findings. Willingness to pay is a measure of value, Regier clarified, and it is estimated as the monetary equivalent of preference-based utility. The results of the experiment showed that the Canadian population that had been surveyed valued secondary genomic information regardless of whether the condition was treatable. Participants decidedly wanted to have actionable findings returned, Regier said, but they also wanted to receive incidental findings regarding non-treatable conditions.
The cost effectiveness of returning secondary genomic findings was studied by Bennette and colleagues, who calculated the ICER (i.e., the incremental cost per QALY gain) (Bennette et al., 2015). Analyzing hypothetical cohorts of cardiomyopathy and colorectal cancer patients as well as the general population, they found that the ICER for return of secondary findings varied depending on the specific patient population. A key aspect of this analysis, Regier said, is decision uncertainty, which is the level of confidence that the ICER will fall below a certain willingness to pay for a QALY gain. There are some limits to this quantitative model, Regier said, in that there is no allowance for personal utility because it is not incorporated into the QALY metric. In addition, the analysis begins at the stage of return (or not) of secondary findings, and upstream costs and consequences are not examined.
Regier explained how he adapted Bennette and colleagues’ model (Bennette et al., 2015) to answer questions about the cost effectiveness of genomic screening for colorectal cancer and polyposis (CRCP) syndromes and the return of secondary findings. By incorporating an allowance for personal utility (i.e., the value of knowing), Regier adapted the ICER equation to calculate net monetary benefit and include personal utility using willingness to pay. His model projected life expectancy and costs for two trajectories: one for an individual who received standard care for CRCP, and another for an individual who received a genomic screening panel and was given the secondary findings. For the latter trajectory, life expectancy
and costs were also projected for relatives who were offered—and accepted or declined—genetic counseling and genetic testing. These models are data intensive, Regier said. Overall, the results suggest that the possible return of secondary findings was likely to be cost effective; however, he noted that there is a lot of decision uncertainty. When using the reference case for the United Kingdom and Canada (which does not factor in personal utility), Regier finds that the probability that CRCP screening and the return of secondary findings is cost effective to be 72 percent, which may be too much uncertainty for the typical decision maker in Canada or the United Kingdom, he said. When the concepts of personal utility and net benefit were incorporated, decision uncertainty was reduced. The probability that CRCP screening and the return of secondary findings was cost effective was 82 percent. Sequencing in Canada is currently fairly expensive, Regier said. The monetary cost of the hypothetical panel was calculated to be $4,600. Further analysis suggested that the probability of cost effectiveness (incorporating personal utility) could be increased to 95 percent if the hypothetical cost of the genomic screening panel and analysis was $3,200.
Considering Personal Utility and the Potential Value of Genomic Knowledge
Current guidelines for assessing the value of genomics technologies do not support the inclusion of personal utility, Regier concluded. However, when the value of precision medicine knowledge from the patient and public perspective is not incorporated, it can lead to the wrong investments by health care delivery systems (over-investment if patients have disutility for the knowledge of their own genomic variants; under-investment if patients place value on that knowledge). The proposed value framework allows for a broader analysis of value, beyond QALY.
From an applied perspective, Regier said, both upstream and downstream considerations are critical. In the absence of personal utility, decision uncertainty is quite substantial. Precision medicine and genomic technologies amplify already complex decisions regarding screening. Additional data are needed to support these complex decisions, including information from patients and the public regarding their perceptions of the value of personal genomic knowledge.
DISCUSSION
Data for Demonstrating Value
Data on penetrance is incredibly important, Powell said, describing which data he would prioritize to demonstrate the value of genomic testing.
As a clinician returning genomic results, he said, he wants to be able to give patients a sense of what they can expect based on the findings. Data sharing about drug response phenotypes across a large population is needed, Peterson added. Genotype information is often accessible, while phenotype data are more difficult to ascertain from the EHR. It will also be important, Regier said, to do a better job of engaging with patients and the public in order to develop an improved understanding of how they value different types of information and health outcomes.
While very large randomized controlled trials are ideal for demonstrating value to payers and guideline writers, some of the simulations conducted required analyzing millions of patients with certain characteristics to achieve the necessary precision, Peterson said, highlighting the need for improved methods for comparative effectiveness research with outside cohorts. There is also a need for better methods to understand cost effectiveness, Regier added. Randomized controlled trials in precision medicine will not be common or broad in scale.
Perspectives on Costs
It was observed that the costs of conducting panel-based preemptive testing are likely to vary over time, and a workshop participant asked how reductions in the cost of testing influence overall cost effectiveness and who bears these costs. Peterson replied that for a high-risk group that is likely to use the information right away, the cost of testing will be swamped by the cost of outcomes. In modeling the value of genomic information for tailoring antiplatelet therapy in coronary patients, for example, the cost of the test, which is generally modeled at $100 to $200, does not have a major influence on the overall economic output. The value proposition changes, however, when screening a very large, average-risk population. The cost of doing business in this area should be split between health care systems, which might indirectly accrue a lot of value for this kind of work, and insurers, who also have a stake, Peterson suggested. From a health care system point of view, there are costs which will not be directly covered by insurers, such as the costs of data management, data, or staff time for delivering information to clinicians or assisting with decision making. These types of costs are difficult to incorporate into the cost of the actual assay. Assay cost is a moving target, Regier agreed, noting the Personalized Oncogenomics Program at the Genome Sciences Center in British Columbia does whole-genome transcriptome analysis for patients with incurable cancers and uses forecasting methods to help understand where that moving target might go in the future.
Most large employers are self-insured and use payers as a third-party administrator, a workshop participant said. It is then the employer who is
responsible for determining health care benefits, along with other employee benefits. The participant asked if cost–benefit and return on investment from an employer’s perspective were different from an insurance company perspective and if health economists model from an employer’s perspective, noting that employee recruitment, retention, and return on investment for some industry sectors and employers can be modeled over 10 years, instead of 1 or 2 years for an insurance member relationship. There are many different people involved in making those decisions, with many different competing interests. Personal utility, for example, is not going to be important to an insurance company, Powell said. It may, however, be important to an employer, because an employer will design benefits and health care coverage to try to attract and retain the desired workforce. Economic analyses can be done from many different perspectives, Veenstra added. It was suggested that when seeking to convince payers to adopt the routine reimbursement of genomics, forward-thinking employers might be more successful targets than a large insurance company or the Centers for Medicare & Medicaid Services. Potential models should emphasize the employer’s perspective, rather than other payer perspectives.
Pharmacist Role in Pharmacogenomics
Clinical pharmacists are critical for clinical decision support, Peterson said, and most hospitals have pharmacists who are assigned to specific teams (e.g., a transplant pharmacist). Unfortunately, the prescribing alert in the EHR is not pushed out to all of the systems that the pharmacists use. There are surveillance systems that the pharmacists use which run alongside the EHR and pick up pharmacogenomic information on current patients.
Taking Clinical Practice and Uptake into Account
There are challenges associated with accounting for clinical practice in models that use retrospective data, observed a workshop participant. For example, if a retrospective economic analysis shows a certain number of actionable variants but in practice only a portion of those are actually being acted on, how can that information be accounted for in the economic analysis? How can the message be delivered to providers that improvement in clinical practice to achieve optimal use, or even appropriate use, could provide a greater cost effectiveness? The behavior of physicians and patients is an extremely important aspect, Peterson agreed, noting that one of the aims of his study is to examine how such behavior influences economic outcomes. The more likely it is that the incidental data are used, the more likely the patient is to benefit from having preemptive screening.
Economic models can vary widely depending on uptake (whether the
patient wants the information, whether the clinician returns the result), Regier added. If decision models are done well (and they are often not), they will incorporate uptake into the model as a probability, he said. The difficulty comes when a novel technology is involved and the degree of uptake is not known. The challenge then is to predict uptake (i.e., patient and clinician preferences) in a way that has external validity, he said, noting that this is what the discrete choice experiment is meant to do. Incomplete uptake leads to varying budget impacts across the health care system. There are advanced techniques—referred to by the general term “value of information analysis”—that can be used to compare the value of doing more research versus the value of getting things implemented, Veenstra said. Such analyses can help to structure these issues and identify evidence gaps.
Incorporating the Heterogeneity of Personal Utility Preferences
It is important to take diversity and diverse populations into account when collecting information on how the general population values genome screening and results. There is no “average patient,” Regier said, and the challenge is how to incorporate the heterogeneity of value or preferences. There are methods being developed that are starting to address this. In the Canadian population he sampled, for example, there was a range of age representation, jurisdictional representation across the provinces, and representation across languages. One population that was missing, however, was Aboriginal First Nations representation, and there is a gap in understanding value among those in traditionally underserved communities.
Information about diverse individuals is often lacking at the biological level as well, and the understanding of founder variants and benign population variants is limited, Powell added. There is also the potential to exacerbate economic disparities with genomic screening. For example, identifying a condition that a person cannot get treatment for can cause more harm than benefit for that person.
There is variation by geography as well as by socioeconomic status, observed a workshop participant. Health literacy and economic and educational factors vary in populations and affect personal utility. Socioeconomic status, including income, geography, and age, affect preferences and personal utility, Regier agreed. Better communication and decision aids are needed to help people make very complex decisions about what information they want. There is an opportunity to engage with patients and the public and provide them with some level of genomic literacy (e.g., explaining terms such as penetrance, treatability, and clinical utility) so these individuals can have conversations with their providers and choose the options that are consistent with the underlying value of the information for them.
Opportunistic Screening
Feero raised the issue of opportunistic approaches to screening (i.e., testing for an additional set of variants when sequencing is ordered for a specific clinical question). By implementing opportunistic screening, relevant genomic data on larger segments of the population are slowly being accrued. Once there has been a commitment to conduct the original genomic test, the incremental cost for adding several additional tests would seem to be quite small, Feero said.
Decision Tools
It is challenging for providers to determine which panel will provide the most benefit for the patient from the many different panels available, a workshop participant commented. Developing decision tools for clinicians to help select the most appropriate panel from what is available may help providers identify the best panel, the participant suggested. There are ongoing efforts to develop decision aids, Peterson said, but unfortunately there is wide variability in terms of how those decision algorithms work or even what variants are on the panel. In his practice, he said, he prefers to work with companies and panels that are transparent in terms of both what is on the panel and how the company uses it to arrive at a recommendation. There are also guidelines that reflect academic agreement on how a particular variant should be used in clinical practice.
There is a risk of creating panels that are so broad that they include variants for which there is not sufficient evidence of utility, Powell added. Such testing creates the risk of doing research under the guise of clinical testing. Projects such as ClinGen will help increase understanding of the genes and the variants for which there is sufficient information to make recommendations for screening, but there is a need to be clear about what testing is research, he said. Different panels are used in different health care systems in Canada, Regier said, and some health care systems do not have access to any genomic panel testing, raising the issue of equity across the country.
Considering Ways to Move Forward
To inform the Roundtable’s development of activities, Ginsburg asked the panel what the population health community should be doing to accelerate the types of modeling discussed and to provide information to institutional decision makers who are considering implementing genomics-based programs. Costs change rapidly over time, Peterson said. While there is value to knowing the current costs, the 5- to 10-year value
of the work being done now will be reliant on the clinical outcomes. The models show that outcomes drive costs, and, from a societal health care perspective, achieving the best possible clinical outcome is the priority, he said. A standardized cost can be attached to that clinical outcome, which has some transferability across health care systems. There is a movement toward embedding health economics within many genome-scale sequencing projects, Powell said, adding that this movement should be encouraged and mentioning the National Human Genome Research Institute’s Ethical, Legal and Social Implications Research Program. It is important to engage budgetary decision makers and provide them with reliable but not overly complicated information, Regier suggested, noting that health economists often present information at a level that is too technically complex. There is also a need to bring the public and patients into the conversations around value, he concluded.




















