Session 2: What Do We Need to Know?
During the second workshop session, speakers and attendees considered the types of information capable of moving research forward in this area. For example, given the expected research on living organisms and disease organisms in permafrost, participants shared ideas on the specific areas of research and other activities (e.g., oil extraction) that may expose scientists or local communities to microbial risks. Speakers were asked to share their thoughts on the kinds of pathogens that might be uncovered in the permafrost, which of these might be still viable, what scientific approaches could be used, and the kinds of facilities or institutions that may have the capacity to do the research. Perhaps most importantly, attendees shared views on the critical gaps in scientific understanding as well as gaps in surveillance and observational capabilities.
Wildlife Health Surveillance
The purpose of surveillance, as outlined by Dr. Craig Stephen, Canadian Wildlife Health Cooperative, is to provide timely intelligence on the health of the population and to inform effective responses to emerging and ongoing issues. On the other hand, monitoring (one part of surveillance) is the ongoing, continuous, routine observation to assess the status of a population. Most wildlife surveillance takes place in the form of a survey, typically because research is time-limited and not continuous or routine. Surveillance can also come in the form of monitoring, though this is not linked to action. Dr. Stephen shared his thoughts on challenges associated with wildlife surveillance including biased population sampling (e.g., bias towards species that can be detected and those that are considered “charismatic”) and hard-to-determine trends (e.g., non-representative samples and lack of denominator data to measure prevalence). In designing effective wildlife surveillance, it is important to consider whether the goal is really tied to monitoring (e.g., characterizing a status of a population, disease, or hazard) or surveillance (e.g., a signal of change to provoke investigation or action). Both of these have distinct tradeoffs and needs to ensure a successful effort. Dr. Stephen suggested that elements for successful monitoring likely include the ability to track populations and hazards, while surveillance calls for options, thresholds, and resources to take action.
The methods through which surveillance is conducted depends on the type: health, risk, or disease surveillance. Dr. Stephen noted that currently, most wildlife surveillance takes the form of hazard surveillance (mostly infectious and parasitic surveillance, though contaminants research is emerging). Risk surveillance is very limited because researchers are rarely characterizing exposure potential. According to Dr. Stephen, there is hardly ever health surveillance of wildlife (where “health” means the capacity to cope). More often, the well-being of animals and their susceptibility to an emerging issue is tracked in population monitoring. Other crucial considerations
include (1) the difference between surveillance “systems” versus time-limited projects, with implications for the identification of trends; (2) examination of specific populations versus scanning for a suite of signals, which affects ability to reveal the unexpected; and (3) looking for health outcomes versus signals of threats and hazards and who can use the data that is gathered.
A wildlife surveillance system likely aims to be an interconnected and interdependent intergovernmental enterprise done in the public good. Acknowledging the difficulty of accomplishing such a robust goal, Dr. Stephen raised a number of related questions about who may have legal authority to do so. Who has stakeholders, rights-holders, and community trust? Who has the expertise and resources to operate and sustain activities? Who has the contextual understanding to assess and communicate the findings? He also posed the important question of how one might determine that these efforts are working. A recent literature review found no systematic evaluations of wildlife surveillance systems (Stephen et al., 2019). To achieve a successful surveillance assessment, one must consider utility, acceptability, feasibility, and accuracy of the systems as well as the system’s ability to meet standards. Dr. Stephen pointed out that many of these factors remain unknown (excluding perhaps utility and acceptability), impeding evaluations and quality improvement.
Dr. Stephen highlighted several key elements for surveillance success, namely a commitment among partners to work together, respect for jurisdictional roles, and emphasis on the linkages among organizations. The design of a successful system is purposeful, supportive of data sharing, accessible to a variety of stakeholders, strategic in linking people with processes and technologies, and includes a combination of epidemiological and laboratory sciences. In particular, he emphasized the importance of having clarity of goals, roles, and activities as well as equal attention towards finding and testing together with sharing data and information. In summary, development of a robust logic framework can highlight the need to support wildlife surveillance systems, especially in terms of requesting needed support and resources.
At the conclusion of his remarks, Dr. Stephen shared special considerations for threat detection surveillance. He noted that a reliable surveillance system could help provide strategic early warning in the context of scenario planning (Fink et al., 2020). Multiple sources of information are needed to move from a surveillance mindset towards one of “epidemic intelligence;” however, the holders of this information are not in the traditional wildlife health field, rather they are community members, landscape managers, meteorologists, and resource managers. It is key to engage these stakeholders in the development and use of these “intelligence-approach” systems, Dr. Stephen added.
Human Health Surveillance
Ms. Luise Müller, Statens Serum Institut, opened her remarks by defining surveillance as “systematic ongoing collection, collation and analysis of data and the timely dissemination of information to those who need to know so that action can be
taken.”1 She emphasized that a key component of surveillance is data that are intended for action. Objectives of a human health surveillance system may include estimates of the magnitude of the health problem in at-risk populations, determination of disease trends, detection of outbreaks, control strategy evaluation, development of hypotheses on etiology and risk factors, detection of changes in isolation and health practices, information on microbiological evolution, and obligations to international entities (e.g., European Commission and the World Health Organization).
Acknowledging that resources are limited, Ms. Müller identified some key selection criteria for disease surveillance: frequency, severity, communicability, associated economic costs, preventability, public and political interests, and international relevance. She highlighted some examples of surveillance systems in Denmark including the clinical mandatory notification system as well as laboratory-based surveillance systems (e.g., Danish Microbiology Database [MiBa]). Other systems include childhood vaccination coverage as well as sentinel-surveillance for influenza. Focusing on MiBa, the backbone of the laboratory- based systems, she noted that it includes all national microbiological test results, in real-time, along with information on positive and negative tests. Surveillance is facilitated by data capture with person-identifiable data, which makes it possible to link to other registers.
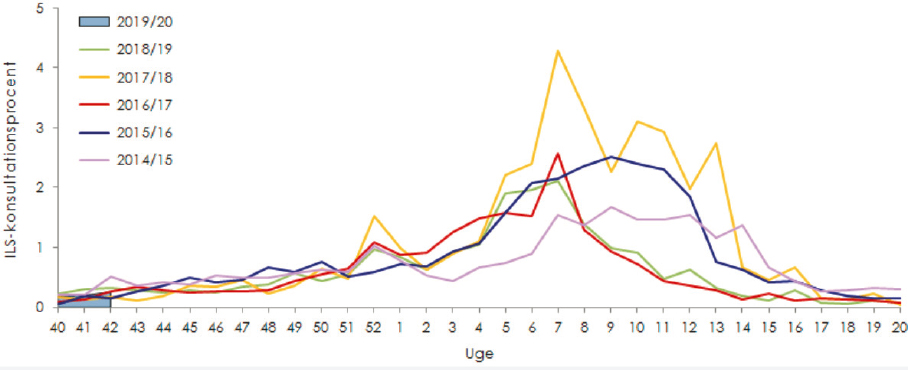
Using a variety of mechanisms including MiBa, scientists try to capture estimates for the entire “surveillance pyramid” for influenza, from symptomatic members of the public, through the general practitioners or on-call doctors, samples taken, hospital or ICU admissions, and all the way to mortalities. Ms. Müller provided an example of the sentinel system where doctors report data on the number of consultations that include a laboratory component. This allows researchers to track these cases and review trends year-to-year (Figure 16).
___________________
1 See A Dictionary of Epidemiology. 4th edition. Ed. Last J. Oxford University Press, 2001.
Ms. Müller then focused on foodborne outbreaks in Denmark, both locally and nationally. In the case of local outbreaks (i.e., point-source outbreaks), responsible institutions include regional food offices, medical officers, and local laboratories. In national outbreaks (i.e., cases in two or more regions), responsible institutions include the Statens Serum Institut reference laboratory and Department of Infectious Disease Epidemiology as well as the Veterinary and Food Administration and Food Institute. These three institutions make up the Central Outbreak Management Group to track outbreaks and coordinate investigations using a One Health approach.
As an example, Ms. Müller discussed a 2017 outbreak of Listeria monocytogenes ST8, a severe disease with relatively high mortality rates, in cold-smoked salmon. A genetic cluster of four cases was identified by whole genome sequencing, one from 2015 and three from 2017, indicating that outbreaks can potentially persist for years. The Central Outbreak Management Group met to discuss the findings, and after two additional cases were identified, comparisons were made between the food and environmental isolates from 2014 to 2017 and matched to cold-smoked salmon from Poland. Interviews of the patients revealed that they had consumed the salmon, and the product was recalled from the market. Upon starting an international inquiry, notifications were issued to the European Epidemic Intelligence Information System for Food and Waterborne Diseases, Early Warning and Response System, and the Rapid Alert System for Food and Feed. Shortly after, a case was reported in France and matched with the same salmon producer in Poland, and a month later, matching cases were reported in Germany as well. Ms. Müller emphasized the importance of whole genome sequencing as a tool to detect and compare food and human isolates, especially in cross-border outbreaks. This case illustrates the usefulness of robust communication platforms in disseminating information quickly. Ms. Müller ended her presentation by reiterating some of the ongoing challenges in human health surveillance such as ensuring that data are collected for action, prioritizing diseases to monitor, detecting unexpected diseases, comparing surveillance data in humans with food and animals using a One Health approach, and harmonizing surveillance data across national borders.
Panel on Research Needs and Gaps in Scientific Understanding and Surveillance Capabilities
During a panel discussion on critical gaps in scientific understanding and surveillance capabilities, Dr. Robyn Barbato, U.S. Army Cold Regions Research and Engineering Lab, encouraged speakers to consider potential research services and programs that may help fill these needs. Reflecting on microbial survival in ancient permafrost, Dr. Lise Øvreås, University of Bergen noted that the Arctic system, organisms, and ecosystems are complex and diverse. Referring to the 2018 IPCC report, she highlighted the increased emissions of CO2 and associated warming, and in particular, the impacts on the Arctic regions. As previous speakers have noted, permafrost is located throughout the Arctic, some of which is isolated and sporadic, and
some of which is continuous. Thawing permafrost contributes to a shift in tundra carbon balance over time; more thawing indicates increased levels of carbon released compared to the amount of carbon that is taken in by the system. Dr. Øvreås outlined some of the changes specifically to the system near Svalbard, including rifts in the soil, lake drainage, and other impacts on local settlements. Researchers in the area have discovered that ice wedges are melting quickly, sometimes on the order of months, releasing carbon and other gases to the atmosphere.
Dr. Øvreås noted that the microbial world is a scientific frontier, with untold bacteria and microorganisms to be discovered and explored. She suggested that most have never been cultivated and their functions remain unknown. These microbes are responsible for fundamental life processes, on a global scale, including cycling of carbon, nitrogen, and other nutrients. It is known that permafrost thaw-induced microbial decomposition releases carbon dioxide and methane, and there is potentially hundreds of gigatons of carbon stored in permafrost. To begin to understand the microorganisms in permafrost, DNA is extracted and genome sequences are studied to provide information on metabolic pathways that could mediate biogeochemical cycles. Findings from this research include a stress response and antibiotic resistance in the samples (Xue et al., 2019).
Dr. Sanne Eline Wennerberg, Veterinary and Food Authority of Greenland, shared her thoughts on the need to access data and the difficulties in gathering this information in Greenland. She reiterated the importance of the research and scientific results from previous presentations, and noted that this work could be extremely helpful in determining risks to local populations. However, access to this information can be challenging. Denmark provides some of these data, but it would be especially helpful to have the ability to engage with local knowledge networks and databases when the need arises. Dr. Wennerberg also highlighted the importance of action and information that is accessible and useable within local communities. Changing climate conditions have clear impacts on the daily lives of people living in Arctic regions, and access to information about potential diseases and emerging threats would enable achievable and actionable responses.
Continuing on the themes raised by Dr. Øvreås, Dr. Jessica Ernakovich, University of New Hampshire, discussed some of the key unknowns within the permafrost microbiome and the role of pathogens now and in the future. She noted that several researchers, collectively known as the Permafrost Microbiome Network, are working on a database to collect, curate, and reanalyze sequencing datasets to understand microbial communities in changing permafrost landscapes. Dr. Ernakovich started broadly by discussing the influence of the landscape topography on microbial communities, emphasizing that whether permafrost landscapes are drained or inundated with water influences which microorganisms are present and can thrive. Dr. Ernakovich then highlighted some of the information that is known about permafrost microbiomes at present: Active layer and permafrost communities are not the same, and more is known about the active layer than the permafrost layer microbiome.
Diversity declines with depth (for bacteria and fungi; it is likely that diversity increases with depth for archaea). It is likely that microbes live in brine channels in frozen permafrost. Microbes in permafrost are selected (adapted or well suited) for long-term survival in harsh conditions (Bottos et al., 2018; Mackelprang et al., 2017). Community composition shifts when permafrost thaws, but the controls on community assembly are not yet known. Understanding what factors contribute to composition of microbial communities after permafrost thaw may be helpful in predicting pathogen risks.
In the microbiome following permafrost thaw, there are still several unanswered questions. Some of these raised by Dr. Ernakovich include: What proportion of the permafrost microbes survive the thaw? The sequencing techniques mentioned previously are not quantitative and thus other tools may be needed. Is there a biological or physical mechanism to determine the likelihood of a microorganism surviving permafrost thaw? Is it random? Given that permafrost microbes are selected for survival, can they compete with newly dispersed active layer microbes? Do increasing biotic interactions affect community composition? How do these change over space and time? She also shared ideas on the role of community assembly in permafrost layers after thaw. When permafrost conditions are stable (intact), permafrost acts as a “selective filter” and the community is selected specifically for life in permafrost and there is not much dispersal from the active layer. When permafrost thaws, there is a change in conditions of selective pressures, in which the community is affected by random losses of species and by dispersal from the active layer. Over time, the community continues to respond to changing niche space, but also to dispersal (Graham and Stegen, 2017) and perhaps also diversification. Additional information is needed on the permafrost microbiome before the survival of pathogens can be understood and predicted. According to Dr. Ernakovich, key information gaps include understanding (1) whether pathogens in permafrost are active or dormant, which may reveal clues about their availability to survive and thrive after permafrost thaw, (2) if they are able to thrive and grow to the required population size to be an effective pathogen, (3) whether pathogen populations will be kept in check by the effects of community, or if the low biomass and diversity of other community members allow pathogens to get a foothold, and (4) if pathogen dominance is one of many alternate stable states.
Dr. Tatiana Vishnivetskaya, University of Tennessee Knoxville, discussed microbial research on permafrost core samples that were approximately two million years old. She noted that permafrost collection techniques may need to be adjusted to ensure that the samples remain uncontaminated. It has been estimated that prokaryotic diversity in soil reaches between 6,400 and 38,000 species per gram (Curtis et al., 2002), but the absolute diversity is unknown. Dr. Vishnivetskaya presented information from a number of published papers on total cell counts, microbial biomass, and viable counts for different permafrost samples (e.g., Gilichinsky, 2005; Hansen, 2007; Rivkina, 1998; Vishnivetskaya, 2000; and others). Only 0.01 to 1 percent of the native microorganisms are able to be cultured; therefore, most of the microbial diversity in permafrost sediments remains unexplored. However, a number of studies have determined the
viable bacterial community structure in different permafrost sediments. She noted that the type of media used by researchers could influence structure of the bacterial community.
Types of microorganisms that have been isolated from Siberian permafrost include prokaryotes (e.g., bacteria, archaea, cyanobacteria) and eukaryotes (e.g., yeasts, fungi, green algae, moss, protista, nematode). Dr. Vishnivetskaya noted that viruses could also be isolated from this permafrost. In all, she identified 26 strains that have been isolated from these permafrost samples (Shatilovich et al., 2018). Database information, based on 37 permafrost metagenomes and 26 active layer metagenomes, shows that the majority of biodiversity exists at levels below 1 percent. Metagenomes show differences in dominant phyla from permafrost versus the active layer, due to their differing functions in the soil.
The final panelist of the session, Dr. David Stanton, Swedish Museum of Natural History, discussed DNA from ice age wildlife and the possibility of pathogens emerging from these samples. A broad range of ancient species has been found in permafrost, with remarkable levels of preservation, even in samples over 50,000 years old. DNA is extracted from the samples, and a library is built and sequenced. Dr. Stanton pointed out that often there are low amounts of the actual species of interest for the research group and the rest are data that could potentially contain bacterial DNA and pathogens. These data could possibly be used to investigate pathogens.
Dr. Stanton shared an example from lion samples and mapping for the anthrax genome. This approach can reveal pathogens present in the carcasses and their dependence on location, age, or other factors. He emphasized that DNA from ice age wildlife is highly damaged and fragmented; even if pathogens are present, they are not necessarily viable. Most of the DNA that the team works with is approximately 70 to 80 base pairs. Another example from mammoth samples revealed an innocuous soil bacterium (Azospirillum brasilense). Some bacteria that produce toxins were discovered as well (Clostridium botulinum), but these would not necessarily pose a threat to researchers working with the samples, unless samples containing the toxin were consumed. He noted that knowledge gaps include an understanding of potential risks posed by pathogens that are present in the samples, and the evolutionary history of parasites as well.
Discussion on Biosafety and Biosecurity Risks
Following breakout group discussions, participants shared ideas and opportunities related to laboratory procedures, biocontainment and engineering controls, as well as risk assessments and impacts on indigenous and local communities.
Laboratory Procedures and Levels of Risk
The first group specifically considered laboratory procedures and techniques that would be helpful when working with permafrost samples. The current method most commonly used is DNA sequencing and identifying known pathogens. There are
other approaches such as laboratory-based culturing, but there is a great deal of unknown diversity in the samples and these techniques likely could be improved. The group noted that there is a list of known pathogens, and it may be most efficient to identify sequences from that list. Proteomics could be used to confirm pathogenicity, though this is still quite challenging and could be an area for future improvement. Participants noted a distinction between environmental pathogens and pathogens from animal carcasses; in the case of animal carcasses, there will likely be visual indications of risk. Although indications of risk may not be quite as clear in the environment, there are “hot spots” of risk that could be identified. Therefore, a systematic approach for sampling in hot spot areas would be useful to understand the dispersal of those organisms for predictive purposes. Group members highlighted their discussion on a potential moral obligation to preserve both the environmental and clinical samples that are collected, though there is no clear method or resources to do this. Potential discord between the environmental microbiome versus the clinical (as well as the techniques that are used), indicates that it may be useful to culture the organisms and use information from genomes to try to predict substrate utilization.
Biocontainment, Engineering, and Safety Controls
A second breakout group covered topics related to biocontainment and engineering controls. They considered that people working or living in the Arctic might be unaware that samples may contain pathogens. There is also very limited biosafety in the North. Given that context, the group discussed how to regulate effectively without curtailing valid research, or if capacity building would be a better option rather than regulation. In addition, the potential for harmful or potentially hostile activities was noted, including the use of pathogens for nefarious purposes. There are opportunities for increased international data sharing and dissemination, especially with new technologies and citizen science approaches, though biosafety and biosecurity2 concerns will still need to be taken into consideration. International frameworks, codes of ethics, or best practices may be used to ensure affected communities are kept informed and to ensure that sovereignty concerns are addressed.
Risk Assessment Approaches
During a discussion on approaches to risk assessment, participants noted that the isolates and samples themselves are not considered a risk, but risk emerges when the samples are combined with human activity and exposure. There is still a great deal of uncertainty about what the risks are and there are challenges to conducting foundational research and achieving higher levels of certainty. Therefore, questions
___________________
2 “The analysis of ways and development of strategies to minimize the likelihood of the occurrence of biorisks (i.e. the probability or chance that a particular adverse event, including accidental infection or unauthorized access, loss, theft, misuse, diversion or intentional release, possibly leading to harm, will occur).” Source: https://www.cdc.gov/globalhealth/security/actionpackages/biosafety_and_biosecurity.htm.
remain about the probability of pathogen survival in permafrost, the probability of transmission, and the severity of outcome. There was some discussion about pathogens in human remains versus plant and animal matter, and the group noted that there are more plant and animal materials in the permafrost versus human remains; approximately 75 percent of emerging infectious diseases are zoonotic. Effective risk assessments begin with an understanding of whose risk is being assessed and for what purpose. It may be beneficial to start with occupational safety of researchers and capacity building for sustainability. Some participants raised additional questions for consideration: Is there a need to focus on evolutionary history studies versus present-day risk assessments? Should researchers start with a small number of current pathogens of concern that are hardy enough to survive in permafrost? Would it be most effective to focus efforts on highly exposed populations (e.g., miners, tusk miners) or certain sentinel species (e.g., Arctic fox or shellfish)? The group noted that starting with occupational risk and conducting this research in a safe way would be key.
Impacts of Potential Exposure on Indigenous and Local Communities
The final group shared thoughts on impacts for indigenous and local communities, and noted that impacts largely depend on the type of thawing permafrost and its contents. People will be exposed through pathogens in soil, as the ground is uncovered. For example, searching for for animal carcasses can be economically beneficial in some areas and the people revealing these materials may be sentinels in understanding exposure and associated impacts. An early indicator of concern may be living animals that are exposed and exhibit disease symptoms. The group noted that there are also spiritual factors that need to be taken into consideration; some communities may be wary of approaching permafrost for fear of mishandling human remains, and this can lead to stress and behavioral concerns within the communities. The State of Alaska has a fact sheet on best practices for handling human remains, even though the cause of death is not always clear. Eroding gravesites should be reported to appropriate authorities, however communities need to be involved with any decision making regarding these sites.
Another concern for local communities is food safety and the development of best practice materials as well as a common vocabulary to ensure understanding. Guidance from trusted authorities could be developed for both the public and officials. Participants pointed out that it is not practical for hunters and fishers to have personal protective equipment at all times. Local public health departments can be a helpful first contact, but coherence between police and public health entities is probably desirable.
This page intentionally left blank.










