1
Introduction and Themes of the Workshop1
The sequencing of the human genome has facilitated a significant increase in our understanding of disease. By using individual genetic information to prevent, diagnose, and treat disease with better precision, genomics-enabled medicine promises health care that is personalized, predictive, proactive, and preventive rather than reactive, said Sam Shekar, chief medical officer at Northrop Grumman Health IT and co-chair of the workshop. On December 8, 2014, the Roundtable on Translating Genomic-Based Research for Health of the Institute of Medicine (IOM) hosted a workshop on Genomics-Enabled Learning Health Care Systems: Gathering and Using Genomic Information to Improve Patient Care and Research in Washington, DC.2
Technological advances have improved the accuracy of genome sequencing while decreasing the cost of obtaining it, and this is allowing for an unprecedented opportunity for practitioners to customize treatment options for patients based on their genetic signatures. As a result of this progress, huge quantities of genomic data are being generated and made available for use in the health care system. These developments—including the rapidly growing number of new technologies, the associated increase in genomes
_________________
1The planning committee’s role was limited to planning the workshop. The workshop summary has been prepared by the rapporteurs as a factual account of what occurred at the workshop. Statements, recommendations, and opinions expressed are those of individual presenters and participants and are not necessarily endorsed or verified by the Institute of Medicine. They should not be construed as reflecting any group consensus.
2The workshop agenda, speaker biographical sketches, full statement of task, and registered attendees can be found in Appendixes A–D respectively. For more information about the workshop, see http://www.iom.edu/Activities/Research/GenomicBasedResearch/2014-DEC-08.aspx (accessed February 9, 2015).
sequenced, and the resulting massive amounts of new data—are enabling the ongoing transformation of the traditional, symptom-based approach to health care and treating disease into a condition-based, personalized-medicine approach, Shekar said. By integrating genomic information into the health system and delivering greater value for clinical diagnosis and treatment, precision medicine will be a disruptive force in health and health care, he said.
The integration of genomic information into clinical practice has led to discussions about how to maximize the benefits of this large amount of data. Electronic health records (EHRs), for example, could be valuable for storing and accessing clinical genomic information, as could other data sources such as self-report databases. However, the current health care system is largely unprepared to handle information on this scale (Starren et al., 2013). There is a lack of standards for such data, and it would also be useful to address such issues as interoperability, scalability, storage, privacy, security, and ethics. While individual efforts may exist to collect and use these data, it would be valuable to have a more coordinated effort that engages the broader stakeholder population in order to improve patient health and maximize the knowledge that would be obtained from integrating genomic information into health care systems.
The inclusion of genomic data in a knowledge-generating health care system3 infrastructure is one promising way to harness the full potential of that information to provide better patient care. In such a system, clinical practice and research inform each other with the goal of improving the efficiency and effectiveness of disease prevention, diagnosis, and treatment (Ginsburg, 2014). To examine pragmatic approaches to incorporating genomics in learning health care systems, the IOM’s Roundtable on Translating Genomic-Based Research for Health hosted a workshop which convened a variety of stakeholder groups, including commercial developers, health information technology professionals, clinical providers, academic researchers, patient groups, and government and health system representatives, to present their perspectives and participate in discussions on maximizing the value that can be obtained from genomic information. The workshop examined how a variety of systems are capturing and making use
_________________
3Knowledge-generating health care system refers to an automated system that relies upon large databases of research and patient information. Information gleaned from patients and clinical research is used in learning networks to inform clinical decisions and create a more efficient way to improve health care for future patients. This concept is also referred to as a learning health care system (IOM, 2012) or a rapid learning health care system (Etheredge, 2014).
BOX 1-1
Objectives of the Workshop
- To explore how key pieces of genetic/genomic information can be effectively and efficiently delivered to patients and clinicians for improving care.
- To discuss how both the health care system and genomic data can be used for evidence generation in research and in patient care.
- To assess current best practices for using knowledge-generating/learning health care systems and which models may provide an opportunity for genomics to be used in the rapid-learning process.
of genomic data to generate knowledge for advancing health care in the 21st century. It also sought to evaluate the challenges, opportunities, and best practices for capturing or using genomic information in knowledge-generating health care systems. The objectives from the workshop are given in Box 1-1.
BUILDING ON THE EXISTING LEARNING HEALTH CARE SYSTEM
The health care system already has made great progress in building a rapid-learning system, even before the widespread incorporation of genomic information in these systems, observed Lynn Etheredge, the director of the Rapid Learning Project. Several entities, including the National Institutes of Health (NIH), the U.S. Food and Drug Administration (FDA), and the Patient-Centered Outcomes Research Institute (PCORI), have been at the forefront of this effort, and billions of dollars already have been invested in laying the foundation for such a system (see Chapter 2 for more information).
The core element of a rapid-learning health system, Etheredge said, is a computerized health system designed to use large, distributed databases and learning networks with tens of millions of privacy-protected patient records. “The goal is to learn as quickly as possible about the best medical care for each person—and to deliver it,” he said.
The model that has been emerging is a common data model for data drawn largely from EHRs and claim information. This model uses distributed but networked databases, national coordinated centers, automated study designs, data quality checks, and analysis and reporting tools for
computerized studies and clinical trials. The data from EHRs and claims are not necessarily as accurate as those from clinical trials, Etheredge acknowledged, but they contain valuable information that often is accurate and is representative of diagnoses from the larger population.
This system offers tremendous performance gains over what has been accomplished in the past, Etheredge said. The rapid-learning model has enabled the production of an abundance of data because many studies can be done and results can be produced much faster than before. More patients and subgroups can be studied, including seniors, children, patients with multiple diseases, minorities, and people with rare diseases. Database studies that used to take 2 years can be completed in weeks and can encompass millions of patients. For example, Etheredge said, in a study of the risk of angioedema associated with patients who take drugs to treat hypertension, 3.9 million individuals from 17 health plans that participated in the Mini-Sentinel program were examined quite quickly because the information was accessible in a database (Toh et al., 2012). Studies performed as randomized clinical trials costing millions of dollars can be completed quickly as registry trials for a fraction of the cost, as was demonstrated with the Thrombus Aspiration in ST-Elevation Myocardial Infarction (TASTE) trial in Scandinavia (Lauer and Bonds, 2014; Lauer and D’Agostino, 2013). With accessible databases, a drug safety trial that used to take months can be done in 24 hours, and knowledge that used to take decades to acquire can be generated in less than 1 year, Etheredge said.
There are other advantages to using a rapid-learning system. More questions can be answered with the data, yielding additional information that is useful to providers and patients. A larger number of researchers and learning networks can be involved, which can lead to both informal and formal collaborations. And more groups are interested in funding the research, including specialty societies, patient groups, health plans, hospital groups, accountable care organizations, and foundations. This new model, Etheredge said, has been “laying the foundation for 21st-century biomedicine as a digital science and as a system that is optimized for discovery science.”
A rapid-learning system also takes advantage of the positive economics of data sharing. If each of 10 institutions shares 100 cases, then each institution gets 900 added cases in return for its sharing of 100, a return of 9 to 1. If each of 100 institutions shares 1,000 cases, the return is 99 to 1. Data sharing is a high-payoff strategy, and more data sharing multiplies benefits, Etheredge said.
Challenges to Integrating Genomics
Moving into the genomics era poses a challenge for the rapid-learning system. The current learning health care system does not integrate genomic information, and part of the challenge of such integration will be to encourage the developers of these systems to incorporate genomic data. The logical way to approach this task, Etheredge said, will be to begin to add genomic data to the common data model4 so that genomic data can be accessed through the high-speed, high-performance research system. Key questions concerning this process are what data to add, from how many patients, and from which population cohorts. The NIH should play a significant role in answering these questions, Etheredge said, but a genomics-enabled rapid-learning health system will require collaboration among multiple government agencies and the health sector as well, including physicians, other providers, and patients.
The investments that have already been made in a pre-genomics rapid-learning health care system have created the foundations and opportunities for a genomics-enabled rapid-learning system, Etheredge concluded. Failure to act now may lead to massive amounts of genomic data being paid for by health systems but not being available for learning.
Health Information Technology Infrastructure
One of the challenges of integrating genomic information into the health care system, Shekar said, is that the massive amount of data requires a supportive information technology infrastructure for the assessment of the data. According to a 2014 report to the Agency for Healthcare Research and Quality,5 a robust data infrastructure that can enable a learning health care system must have several features (see Figure 1-1). One is the ability to integrate various sources of information, including clinical data, genomic data, and laboratory data. The integrated data are analyzed with tools for sequence processing, managing big data analytics, and using the cloud, all with data security and safeguarded access. Findings from these analyses must be visualized in a way that makes sense and that can be communicated at the point of care and in a
_________________
4Distributed Database and Common Data Model, http://mini-sentinel.org/date_activities/distributed_db_and_data/details.aspx?ID=105 (accessed March 8, 2015).
5Data for Individual Health, http://healthit.ahrq.gov/sites/default/files/docs/publication/2014-jason-data-for-individual-health.pdf (accessed February 9, 2015).
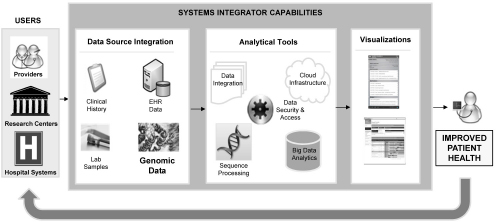
FIGURE 1-1 The deployment of information technology within a knowledge-generating health care system can advance clinical research and care.
SOURCE: Shekar, IOM workshop presentation on December 8, 2014.
brief patient encounter, Shekar said. It would be useful to measure the resulting improvements in patient health, with the results fed back into the system to improve the value and effectiveness of patient care.
A knowledge-generating health care system enabled for genomics will not be a separate system, but rather will be an extension of the current system, said Geoffrey Ginsburg, director of the Center for Applied Genomics and Precision Medicine and professor of medicine and pathology and of biomedical engineering at Duke University and co-chair of the workshop. As such, all stakeholders will be involved in shaping that system, including providers, insurers, patients, researchers, policy makers, and the health information technology community. Ginsburg offered the questions in Box 1-2 to help frame the workshop discussions about the use of genomic data within health systems.
Stakeholders will need to decide how to use the data, Ginsburg said. Health care providers will need information at the point of decision so that they are able to use it in the context of their clinical workflow, he said, and patients will need to define preferences about the use and sharing of their genomic information. All members of the health care workforce and the public will need sufficient genomic literacy to make use of new information. Researchers will need to identify and adopt best practices for
- How can health systems engage individuals to achieve health using genomic and other technologies?
- How can systems, providers, and patients learn from failed efforts to continuously improve health and treatments?
- How can genomic data be used to support patient-centered care?
- How can health systems help research and care teams have access to all the data?
research using EHR-linked genomic information. The EHR vendor community will work separately and collaboratively to offer providers systems that will enable them to make more informed decisions, Ginsburg said. The health information technology community will need to design secure and interoperable genomics-enabled systems for actionable use in both health care and community settings. And policy makers will need to address the return of results, privacy, confidentiality, and education while developing regulations and economic incentives that can align all stakeholders toward the same outcomes. Health care providers will need to learn to apply genomic information to clinical decisions.
ORGANIZATION OF THE WORKSHOP SUMMARY
Following this introductory chapter, Chapter 2 considers the types and quality of genomic data to be handled by a knowledge-generating health care system. Examples are provided of pharmacogenomics data used for research as well as genetic and environmental information used to study common diseases within a learning health care system. The chapter describes efforts by U.S. government agencies and UK agencies to explore the use of large-scale genomic data to inform both research and patient care. The issue of standardizing data so that it could be reused to maximize its potential is also addressed.
Chapter 3 considers the use of genomic information in patient care and research from different perspectives within the health care system. A foundational concept that was presented is the closed, platform-supported rapid learning cycles that provide bidirectional feedback of the analysis of data to produce results and the use of those results to change practices. It is explained that a key element of a functional rapid learning
system is engaging patients and understanding their preferences for access and sharing of the data. Educating patients and physicians about the effective use of genomic data in the clinic and addressing health care disparity issues that may arise from introducing genomic information into the health care system is also discussed.
Chapter 4 examines both the potential and the challenges of integrating genomics into the EHR. Effective incorporation of the data into EHR platforms will require establishing data standards so that information is transmitted among interoperable systems and shared easily. The chapter also addresses challenges to a genomics-enabled EHR, such as determining how and what information will be shared, ensuring access to the information, establishing clinical decision support and guidance for clinicians, and improving the management of big data for enhancing clinical care.
Chapter 5 reviews the recent activities and future plans of the Displaying and Integrating Genetic Information Through the EHR (DIGITizE) Action Collaborative. The idea for this group originated with the Roundtable on Translating Genomic-Based Research for Health to engage key stakeholders in establishing a framework for data standards. The action collaborative is focusing on pharmacogenomics use cases to establish a pilot project for representing genetic test information in a structured format that can reside in the EHR ecosystem.
Finally, Chapter 6 presents possible next steps identified during workshop discussion sessions for integrating genomic data into the health care system. Several workshop participants discussed how it would be useful to make EHRs fully interoperable for genomic and other clinical information. The social science and behavioral aspect of implementing genomic information in the clinic is discussed, as is the inclusion of consumer data.








