1
Introduction
Every patient is unique, and the evolving field of precision medicine aims to ensure the delivery of the right treatment to the right patient at the right time. In an era of rapid advances in biomedicine and enhanced understanding of the genetic basis of disease, health care providers increasingly have access to advanced technologies that may identify molecular aberrations specific to an individual patient that subsequently can be targeted for treatment. Known as biomarker tests for molecularly targeted therapies, these complex tests have the potential to enable selection of the most beneficial treatment for the molecular underpinnings of an individual patient’s disease. Such tests are key to unlocking the promise of precision medicine (see Box 1-1).
Further advances in precision medicine, however, require tests that are accurate, reliable, properly validated, and appropriately implemented in clinical practice, as well as the collection and sharing of information on the outcomes of patients whose treatment is guided by these biomarker tests. In other words, precision medicine requires getting the biomarker tests right to optimize the treatment of each patient and improve patient outcomes, while at the same time advancing our understanding of the role of genetics in disease. Getting the biomarker test right is crucial because a bad biomarker test is as problematic as a bad drug (Hayes, 2013).
Patients recognize the promise of molecularly targeted therapies and are looking to the scientific and biomedical communities to provide validated, reliable biomarker tests that accurately direct treatment at an individual level that has the potential to lead to better outcomes with fewer
side effects (IOM, 2012a). Research discoveries have enabled hundreds of investigational targeted agents to enter the cancer drug development pipeline, and several targeted cancer drugs have been approved for clinical use over the past several years. Progress has been uneven, however, because advances have not been consistent across all types of cancers, and meaningful improvements have been slow to materialize in many other disease domains (IOM, 2015c). Timely access to reliable tests that enable health care providers to accurately match therapies to individual patients is critical for patients with cancer and other diseases.
BIOMARKER TESTS
Remarkable scientific and technical advances have occurred over the past decade and a half, including breakthroughs that emerged from the first draft sequence of the human genome in 2001. That landmark achievement, and subsequent discoveries, served to propel biomedical research in genomics and other omics-based fields,1 as well as bioinformatics
___________________
1 “Omics” is a term encompassing multiple molecular disciplines that involve the characterization of global sets of biological molecules such as DNAs, RNAs, proteins, and metabolites. For example, genomics investigates thousands of DNA sequences, transcriptomics investigates all or many gene transcripts, proteomics investigates large numbers of proteins, and metabolomics investigates large sets of metabolites. Omics-based tests can be considered a complex form of a biomarker test. An omics-based test is derived from complex high-dimensional data; these data are often generated through measurement of many more variables per sample than the total number of biological samples used to generate the dataset. These data are used to produce a computational model that can be used to analyze samples from individual patients (IOM, 2012a).
and computational biology. This research has afforded a more profound understanding of the molecular and genetic basis of disease (IOM, 2012a). These biomedical advances have converged in the rapidly evolving field of precision medicine with a proliferation of complex tests identifying biological indicators, or biomarkers. Definitions and terminology are critical to a complex, rapidly evolving field such as biomarker tests for molecularly targeted therapies (see Box 1-2). The committee also provides additional scientific and technical definitions in the Glossary of the report.
Biomarkers are defined as “a characteristic that is objectively measured and evaluated as an indicator of normal biological processes, pathogenic processes, or pharmacologic responses to a therapeutic intervention” (Biomarkers Definitions Working Group, 2001, p. 91). These can be measurements of macromolecules (DNA, RNA, proteins, lipids), cells, or processes that describe a normal or abnormal biological state in an organism (IOM, 2010a).
Biomarker tests have many different uses in clinical practice (see Table 1-1), including disease screening tests (e.g., prostate-specific antigen), diagnostic tests (e.g., pathologic or histologic assessment of a tissue biopsy), treatment and posttreatment monitoring tests (detection of treatment complications or subsequent disease advancement), and prognostic tests for estimating risk or time to clinical outcomes (e.g., aggressive cancers have a poorer prognosis than more indolent cancers). In addition, biomarker tests are used to predict patient response to specific treatments (IOM, 2007, 2010a).
Such predictive biomarker tests are used by health care providers to tailor treatment to an individual patient’s clinical condition and treatment goals. A subset of these tests examines an individual’s ability to metabolize a drug, primarily in the context of treatment-related toxicity. Another subset includes biomarker tests for specific aberrations in biological mechanisms of action that are associated with response or resistance to a specific targeted therapy. The clinical use of these predictive tests, referred to in this report as biomarker tests for molecularly targeted therapies, is the focus of this study.
A number of types of biomarker tests for molecularly targeted therapies are in clinical use (see Figure 1-1), ranging from single-analyte tests to guide the use of a single class of therapy (e.g., human epidermal growth factor receptor 2 [HER2] amplification and trastuzumab) to a suite of multiple, but separate, tests for single analytes to guide the use of multiple therapy options in a specific clinical context (e.g., estrogen receptor/progesterone receptor [ER/PGR] expression and HER2 amplification for guiding treatment for breast cancer). Multiple-analyte panels include additional analytes for other clinical or research purposes, including assessing secondary response or resistance to targeted therapies or
eligibility for enrollment in clinical trials. Finally, the entire genome may be analyzed using next-generation sequencing (NGS) technology (IOM, 2015c; Meric-Bernstam et al., 2015; Yu et al., 2015). Rapid technological advances have decreased the per-analyte cost of testing (Hayden, 2014; Trosman et al., 2015).
However, the unprecedented amount of data available from a single NGS test, resulting from what is essentially a parallel series of hundreds, thousands, or even millions of single-analyte tests performed on a patient specimen, are blurring the line between clinical research and clinical care.
In oncology, for example, such a test result could suggest treatment with a variety of drugs, each with varying levels of evidence supporting their efficacy. The implications of this transition, specifically related to tests guiding the use of molecularly targeted therapies, are one of the central topics addressed throughout the subsequent chapters of this report.
Regardless of the type of biomarker test for molecularly targeted therapy being performed, the test results that ultimately inform treatment decisions rely on the performance and interpretation of these biomarker tests by anatomic and clinical pathologists (hereafter referred to collec-
TABLE 1-1 Clinical Uses of Biomarkers
| Clinical Biomarker Use | Clinical Objective |
|---|---|
| Screening | Detect and treat early stage disease in the asymptomatic population. |
| Diagnosis/differential diagnosis | Definitively establish the presence and precise description of disease. |
| Classification | Classify patients by disease subset. |
| Prognosis | Estimate the risk of or the time to clinical outcomes. |
| Prediction/treatment stratification | Predict response to particular therapies and choose the drug that is most likely to yield a favorable response in a given patient. |
| Therapy-related risk management | Identify patients with a high probability of adverse effects of a treatment. |
| Therapy monitoring | Determine whether a therapy is having the intended effect on a disease and whether adverse effects arise. |
| Posttreatment monitoring | Provide early detection and treatment of advancing disease or complications. |
SOURCES: Adapted from IOM, 2007, 2010a.
tively as pathologists), clinical laboratory geneticists, and other laboratory health care professionals. These professionals must be aware of existing and emerging uses and limitations of the testing methodologies and the interpretation of test results in order to reliably report the clinical significance of biomarker test results to other health care providers.
PRECISION MEDICINE
Biomarker tests for molecularly targeted therapies are used to select the therapy most likely to result in a favorable response in a given patient (IOM, 2012a). These tests are key to the clinical implementation of precision medicine, which depends on the application of information about molecular mutations or aberrations in an individual patient’s genome or tumor to classify patients into subgroups based on their potential response to a specific treatment. The goal of this stratification is to ensure that patients receive the most beneficial therapies. Accurately matching therapy to the individual patient optimizes treatment selection by focusing specific therapies on those most likely to benefit and decreases treatment harms by avoiding treatment in those unlikely to respond
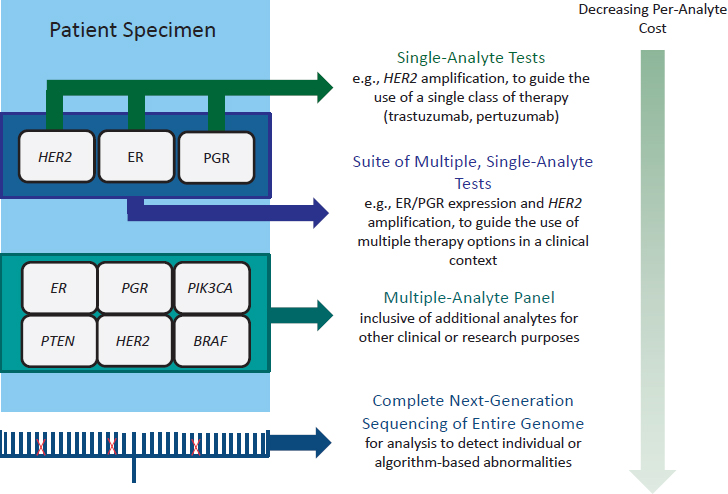
NOTE: BRAF = B-RAF proto-oncogene, serine/threonine kinase, ER = estrogen receptor, HER2 = human epidermal growth factor receptor 2, PGR = progesterone receptor, PIK3CA = phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit alpha, PTEN = phosphatase and tensin homolog; all are analytes potentially detected by biomarker tests in oncology.
SOURCE: Adapted from Yu et al., 2015.
or predicted to have an adverse reaction to treatment. Moreover, biomarker tests for molecularly targeted therapies may have the potential to “bend the health care cost curve” (Armstrong, 2012) through cost savings achieved by avoiding use of nonbeneficial treatments in specific patients (Armstrong, 2012; de Gramont et al., 2015; Jameson and Longo, 2015; NRC, 2011; Schott et al., 2015).
Certain biomarker tests have demonstrated cost-effectiveness, including some gene expression tests to predict risk of cancer recurrence for patients with early-stage breast cancer (Harris et al., 2007). These test results indicate that many women can safely avoid toxic and costly chemotherapy regimens (Jain and Gradishar, 2014; Lyman et al., 2007; Sparano et al., 2015). Studies also have shown that multiplexed testing of tumors for epidermal growth factor receptor (EGFR) and anaplastic lymphoma kinase (ALK) gene rearrangements, followed by biomarker-guided molecularly targeted therapy is cost-effective compared to standard che-
motherapy without any testing in patients with metastatic non-small cell lung cancer (Romanus et al., 2015).
The underlying concept of precision medicine, namely using information about an individual patient’s characteristics to direct treatment, is not a recent innovation. For example, “blood typing has been used to guide transfusions for more than a century” (Collins and Varmus, 2015, p. 793). However, the tremendous scientific advances over the past decade and a half are what have led to a deeper understanding of the molecular underpinnings of complex diseases, enabling researchers to identify the genetic alterations in tumors in great detail—including the specific genetic alterations that drive the growth of individual tumors. Much optimism exists that this newfound ability will lead to more effective treatments and improved outcomes for patients (IOM, 2015c). The recently launched Precision Medicine Initiative calls for significant financial investment in this evolving field in an effort to improve health and disease treatment outcomes (see Box 1-3).
Oncology has been at the forefront of advances in precision medicine, primarily due to the genomic nature of cancer: “Most cancers harbor a cocktail of mutated (or otherwise altered) oncogenes and tumor suppressors that work in concert to specify the molecular pathways that lead to their genesis, maintenance, and progression” (Garraway et al., 2013, p. 1). Treatment previously based on the anatomic origin of cancer (e.g., lung, breast, colon, prostate) is being expanded to include the use of genomic
tests to stratify patients into subsets based on the specific molecular drivers of their individual tumors. The tailoring of treatment to specific molecular targets is being applied to other diseases and conditions, including cystic fibrosis and Duchenne muscular dystrophy (Fairclough et al., 2013; Garraway et al., 2013; Mendelsohn, 2013; Rubin, 2015; Schott et al., 2015; Towse et al., 2013; Trosman et al., 2013).
The evolving field of precision medicine has significant potential to improve health care and patient outcomes, but science- and policy-related challenges may constrain further progress. Although significant advances have been made in terms of understanding the biological basis of diseases such as cancer, continued research is required in domains such as molecular biology, cell biology, and biochemistry to further researchers’ and health care providers’ understanding of the biology behind the alterations that drive the progression of cancer and other “omics-based” conditions (Marcus, 2015; Parkinson et al., 2014; Poste, 2011; Sawyers, 2008).
In oncology, such deeper knowledge of the biology of driver mutations must be combined with the ever-increasing volume of genomic data available to expand the range of targeted treatments. It is important to acknowledge significant sentinel treatment successes, such as imatinib (Gleevec®) in the treatment of patients with chronic myeloid leukemia and erlotinib for non-small cell lung cancers with EGFR-activating mutations. However, despite the discovery of RAS mutations in many different tumor types, no successful targeted treatment has yet been developed and implemented in clinical practice, leading one expert to observe: “We are still dealing with a large gap between discoveries at the lab bench and treatments at the bedside” (Marcus, 2015, p. 31). Another daunting challenge to the development of targeted cancer treatments is the variability between cancers that were previously considered to have a more uniform biology, such that a patient with one type of cancer likely has subsets of tumor cells that differ genetically (IOM, 2013c). Such tumor heterogeneity has a critical impact on treatment strategies, as a therapy designed to target a driver mutation in one cell subset may not have an impact on another subset within the same tumor (Marcus, 2015). Moreover, evidence reveals that some targeted therapies can be context specific: colon cancers with BRAF mutations, for example, are largely unresponsive to BRAF inhibition despite therapeutic effectiveness in BRAF-mutant melanoma (Hyman et al., 2015; Prahallad et al., 2012).
The evidence for the clinical use of biomarker tests to direct treatment is constantly evolving; research into therapies thought to target only one variant of a biomarker, for example, may in fact have a more complex mechanism of action and may be found to be effective against other molecular targets (see Box 1-4). Advanced biomarker tests to direct molecularly targeted therapy can be used to characterize a patient’s dis-
ease, and suggest the use of treatments beyond a drug’s FDA-approved intended use (i.e., off-label use). For example, evolving evidence could indicate that a drug originally developed and approved to target one type of tumor may be effective against different types of tumors. For patients faced with few FDA-approved treatment choices, particularly those with rare cancers or other diseases, this off-label use of molecularly targeted therapies has become an important treatment option.
To develop a deeper understanding of the potential benefits and risks of such molecularly targeted therapies, it will be critical to track their impact on patient outcomes, whether the treatment is on- or off-label. These data on biomarker test use and treatment selection, as well as patient outcomes, need to be systematically captured, analyzed, and shared for continuous learning. In this respect the use of molecularly targeted therapies, particularly in patients with an unmet need for effective treatment, represents a blurring of the line between clinical research and clinical care. Traditional clinical trials only enroll a small proportion of patients, and these are often drawn from those patients who are treated at larger medical centers (Murthy et al., 2004; NCI, 2010). The majority of patients with diseases such as cancer, for example, are still treated in smaller community hospital settings (The Moran Company, 2013). Indeed, the process of ongoing evidence development requires effective approaches to handling the large amounts of complex omics-based patient information across various clinical settings, which need to be developed and implemented. Electronic health records (EHRs) must be configured that are capable not only of capturing individuals’ genomic and other omics-based information, but also providing support tools to aid clinical decisions based on that information (Kohane, 2015; Mirnezami et al., 2012).
The focus on a deeper understanding of disease based on molecular phenotyping may lead to reclassification of disease states to incorporate molecular data, potentially through modernization of the World Health Organization’s International Classification of Diseases. The National Research Council proposed a “new taxonomy of disease” and highlighted the need for new disease classifications to reflect both fundamental biology as well as traditional signs and symptoms (Mirnezami et al., 2012; NRC, 2011).
These significant challenges notwithstanding, precision medicine is advancing our understanding of the molecular basis of diseases and leading to new treatment strategies. The advancement of precision medicine needs to balance optimism and enthusiasm about the promising impact of new emerging technologies and targeted treatments with pragmatic approaches to overcoming current challenges (Joyner and Paneth, 2015; Rubin, 2015).
STUDY SCOPE
The advancement of precision medicine depends not only on progress in science and technology, but on the creation of a supportive policy infrastructure to promote and facilitate the adoption of appropriate biomarker tests for molecularly targeted therapies into routine clinical practice. The policy infrastructure encompasses regulatory issues, including the type and level of oversight needed for test development, validation, and use in clinical practice; the type, amount, and quality of evidence required for health plans, health insurers, and other payers to make coverage and payment decisions; and the best methods of disseminating knowledge of new tests and targeted therapies across a range of clinical practice settings to meet the informational needs of patients, families, the public, and health care professionals (Frueh, 2013; Ramsey and Sullivan, 2014; Simonds et al., 2013).
One of the more significant obstacles to creation of a supportive policy infrastructure is the lack of agreement among stakeholders regarding the evidentiary standards for the clinical utility of a biomarker test that directs molecularly targeted therapy. Such a test is deemed to have clinical utility if evidence demonstrates that the test will result in improvement in patient outcomes (IOM, 2012b). Indeed, the increased complexity of biomarker tests and precision medicine more broadly has led many observers to assert that new standards and methods are needed to assess clinical utility as well as to inform regulatory and reimbursement decisions (IOM, 2015c). Moreover, there is concern that payer decisions about coverage and reimbursement for biomarker tests for molecularly targeted therapies are not reflective of their clinical value, leading to reluctance on the part of test developers to invest resources to demonstrate clinical utility to support reimbursement decisions—what Hayes and colleagues characterize as “a vicious cycle” (Hayes et al., 2013).
In an effort to explore these opportunities and challenges, the Institute of Medicine (IOM) appointed an independent committee that was charged with examining policy issues related to the clinical development and use of biomarker tests for molecularly targeted therapies. The committee views this study report as building on the 2012 IOM consensus report Evolution of Translational Omics: Lessons Learned and the Path Forward (IOM, 2012a). The Omics report examined key issues in the proper development and validation of complex omics-based biomarker tests and recommended a three-step framework for evaluation that includes the discovery phase, the test validation phase, and the evaluation for clinical utility and use stage. The first stage of omics-based test development, as shown in Figure 1-2, includes two phases: discovery and test validation.
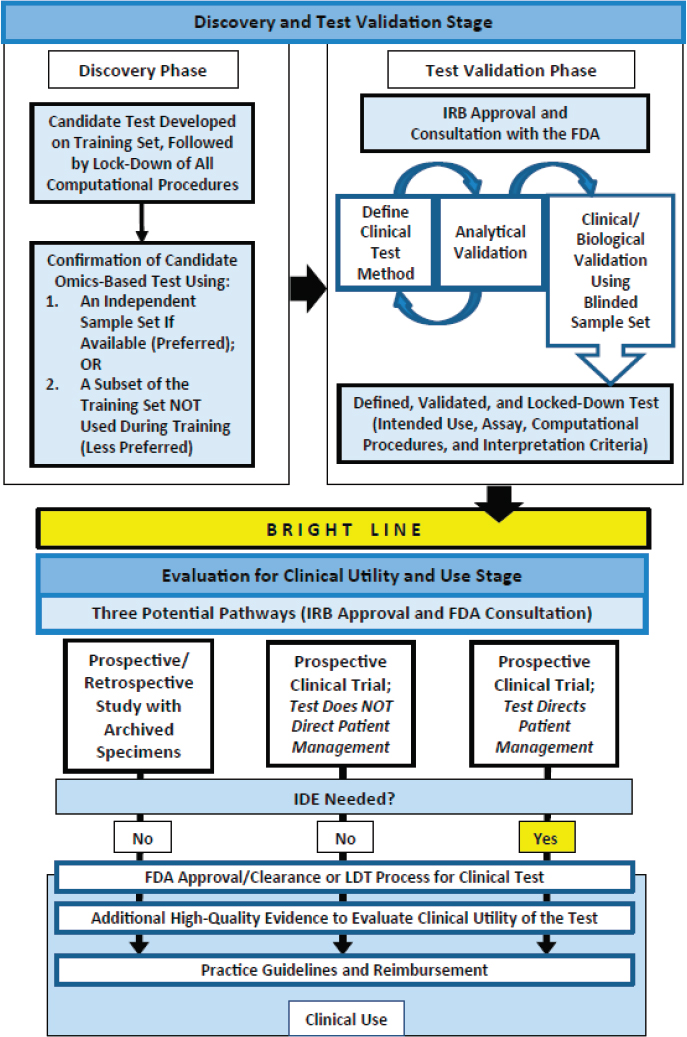
NOTE: FDA = Food and Drug Administration; IDE = investigational device exemption; IRB = institutional review board; LDT = laboratory-developed test.
SOURCE: IOM, 2012a.
In the discovery phase, a candidate test is developed and confirmed. The fully specified computational procedures are locked down in the discovery phase and should remain unchanged in all subsequent development steps. . . . In the test validation phase, the omics-based test undergoes analytical and clinical/biological validation. The bright line signifies the point in test development where a fully defined, validated, and locked-down clinical test (analytical and clinical/biological validation) is necessary. . . . In the second stage of test development, the fully defined, validated, and locked-down omics-based test undergoes evaluation for its intended clinical use. . . . Statistics and bioinformatics validation occurs throughout the discovery and test validation stage as well as the stage of evaluation for clinical utility and use. (IOM, 2012a, p. 7)
A National Cancer Institute (NCI) working group of scientists and other stakeholders was formed to operationalize the principles set forth in the Omics report. The group created a checklist of 30 points to determine whether an omics test is ready for use in a prospective clinical trial involving patient-care decisions, such as the selection of therapy. The checklist will be used to evaluate NCI-sponsored clinical trials in which selection of patient therapy will be based upon the results of omics tests (McShane et al., 2013).
This committee viewed the 2012 report as foundational to its work and reasoned that this current study begins where the Omics study ended: at clinical use of biomarker tests for guiding molecularly targeted therapy. In light of the 2012 report’s thorough treatment of the clinical development of complex omics-based biomarker tests, the committee did not believe it could improve upon or expand on the report’s comprehensive treatment of the topic.
This committee’s statement of task was to examine the interconnected regulatory, reimbursement, and clinical practice policy issues related to the use of biomarker tests for molecularly targeted therapies (see Box 1-5). Given this committee’s charge was to examine biomarkers for molecularly targeted therapies, this report is focused exclusively on predictive biomarker tests to direct molecularly targeted therapy and the content of this report and the committee’s recommendations reflect that focus. Thus, prognostic, screening, monitoring, and drug metabolism pharmacogenomic biomarker tests are outside the scope of this study. The committee took a broad view of molecularly targeted therapies, but many examples in the report pertain to oncology given its position at the leading edge of available targeted therapies.
Support for this study was provided by a broad coalition of public and private sponsors, including the American Society for Radiation Oncology, American Society of Clinical Oncology, Breast Cancer Research Foundation, Centers for Disease Control and Prevention, College of American
Pathologists, Gilead Sciences, Janssen Diagnostics, National Cancer Institute, Novartis, Pfizer Inc., Quest Diagnostics, and Susan G. Komen.
The committee membership reflects a broad range of expertise, including genomic medicine, biostatistics, bioinformatics, test development and translational research, outcomes research and health economics, academic clinical laboratories, pharmaceutical, molecular diagnostics and clinical laboratory industries, test coverage and reimbursement, bioethics, medical education, community practice, and patient advocacy. Brief biographies of the 15 committee members are available in Appendix A.
CONTEXT OF THE STUDY
Biomarker tests for molecularly targeted therapies do not exist in a vacuum, and must be viewed within the context of the broader health care system. The U.S. health care system is undergoing rapid and far-reaching changes, from the proliferation of cutting-edge technological advances and the growing influence of precision medicine to innovative care delivery and payment reforms. The accelerating pace and significant scope of change raises considerable challenges for all health care stakeholders—
patients, health care professionals, health plans and insurers, regulatory agencies, researchers, pathologists, geneticists, and in vitro diagnostic and pharmaceutical manufacturers—while also offering new opportunities to improve patient outcomes and cost-effectiveness of care. The ongoing transformation of health care includes the increased use of EHRs, linking data on health care quality and outcomes; growing consolidation and coordination among care providers, including hospital systems and physician practice groups; and intensified focus on clinical outcomes as risk is shifted from payers to health care providers, with payment more closely tied to the value of health care rather than the volume of services. Defining value in health care is challenging as is evidenced by multiple definitions, and varying perspectives on value that exist. Essentially, health care value is premised upon an assessment of the quality of care relative to its cost. Value is seen to be created when health care outcomes improve while costs remain stable, or when costs decrease without an adverse impact on health outcomes (IOM, 2013a).
One important factor influencing the changing health care landscape is the implementation of the Affordable Care Act of 2010, which continues to affect the nearly $3 trillion U.S. health care system through expanded insurance coverage, reform of health care delivery and payment systems, and new measures that transfer more responsibility for cost and quality from payers to health care providers, with a renewed focus on value (Blumenthal et al., 2015). Given that overall health care expenditures represent 17 percent of the nation’s gross domestic product, and that government pays for 43 percent of U.S. health care costs, efforts to control costs and improve quality, thereby enhancing the value of health care, are critical (Blumenthal et al., 2015; CMS, 2014).
Proof that health care reform efforts are ongoing is evidenced by two significant pieces of draft legislation currently in discussion in Congress. In July 2015, the House Energy and Commerce Subcommittee voted in a bipartisan fashion to move HR 6, known as the 21st Century Cures Act,2 out of committee. The bill aims to accelerate the availability of safe and effective treatments and contains a number of provisions related to biomedical research generally and precision medicine and biomarkers in particular. Although the bill has bipartisan support, and has been praised by many stakeholders in health research and care, concerns have been raised regarding some of the draft provisions, particularly those that may affect FDA’s ability to regulate medical devices (Redberg and Dhruva, 2015).
In contrast to the comprehensive approach taken in the House of Representatives, the Senate Health Education, Labor and Pensions (HELP)
___________________
2 See http://energycommerce.house.gov/cures (accessed June 6, 2016) for extensive background information.
Committee is drafting a series of bills focused on a number of issues related to electronic health records, medical device regulation, targeted therapies and elements of the president’s Precision Medicine Initiative (Alexander, 2016). At the time of this writing, the Senate bills are entering into the markup phase and it is not clear what shape the legislation ultimately will take.
Previous IOM Work
In addition to Evolution of Translational Omics (IOM, 2012a) noted earlier, previous related IOM work includes Toward Precision Medicine (NRC, 2011), Evaluation of Biomarkers and Surrogate Endpoints in Chronic Disease (IOM, 2010a), and Cancer Biomarkers (IOM, 2007). The Surrogate Endpoints and Cancer Biomarkers reports examined the processes for validation, development, and use of biomarkers, in general, and provided an additional basis from which the committee could explore the policy issues related to the clinical use of biomarker tests to guide molecularly targeted therapy. Other related consensus reports provided important background and context to the committee’s work, including Delivering High-Quality Cancer Care (IOM, 2013c) as well as Best Care at Lower Cost (IOM, 2013a). In addition, the National Academies of Sciences, Engineering, and Medicine’s National Cancer Policy Forum and the Roundtable on Translating Genomics-Based Research for Health have produced an extensive number of workshop summaries on a broad range of topics in the fields of cancer and genomics, respectively, which served as a springboard for this committee’s examination of policy issues related to biomarkers for molecularly targeted therapies (IOM, 2009, 2010b, 2012b,c, 2013b,d,e, 2014a,b, 2015a,b).3
METHODS OF THE STUDY
The committee sought to expand its understanding of the full range of challenges and opportunities facing biomarker tests for molecularly targeted therapies. A diverse range of sources informed the committee’s work, including published literature and expert testimony. The committee deliberated during four in-person meetings, as well as numerous conference calls and email exchanges, between January and September 2015.
The committee invited a number of external experts to inform its deliberations during its first three committee meetings (January, April,
___________________
3 See https://www.nationalacademies.org/hmd/Activities/Disease/NCPF.aspx for a complete list of National Cancer Policy Forum Workshop Summaries. See https://www.nationalacademies.org/hmd/Activities/Research/GenomicBasedResearch.aspx for a complete list of Genomics Roundtable Workshop Summaries.
and June 2015). These speakers provided valuable input to the committee on a broad range of issues, including biomarker development, evaluation, and implementation; applications of biomarkers and molecularly targeted therapies in clinical practice; and payment and regulatory issues affecting biomarker tests for molecularly targeted therapies. In addition to in-person testimony, the committee heard from experts in reimbursement, payment, and coverage policy related to biomarker tests for molecularly targeted therapies via webinar (see Appendix C). Moreover, a number of experts provided written input to the committee in areas related to ethics, genomic literacy, and genomic data collection and analysis. Finally, in addition to benefiting from a range of expert oral and written input, the committee reviewed an extensive body of literature on biomarker tests for molecularly targeted therapies to inform its deliberations.
Framework for the Study
The successful adoption of biomarker tests for molecularly targeted therapies into routine clinical practice to improve patient outcomes depends on a number of interrelated factors: ongoing research and development of targeted therapies and associated biomarker tests with a changing body of evidence over time, a responsive regulatory and reimbursement process capable of keeping pace with rapid technological developments, health care providers trained in and knowledgeable about which test(s) to order and how to act on the test results, insurers and other payers who recognize the value of biomarker tests and targeted therapies by coverage and reimbursement (Agarwal et al., 2015), and patients who understand both the potential and current limitations of precision medicine. These complex and interrelated factors are currently affecting the potential of biomarker tests for targeted therapies to improve patient outcomes. The committee emphasizes the interconnected nature of these challenges and the need for an integrated approach to address them—that is, an interdisciplinary perspective that considers all the components in the process and their interactions (IOM, 2012d). An engaged collaboration across stakeholders—including patients, health care providers, payers, health insurers, federal agencies, professional organizations, researchers, and academic as well as community-based health centers—is required for biomarker tests and molecularly targeted therapies to transform promise into the reality of improved patient care and greater cost-effectiveness.
The committee recognizes that a systems approach is required to allow the most effective use of biomarker tests to fully realize the promise of molecularly targeted therapies. For this reason, the committee envisions the creation of a rapid learning system for biomarker tests for molecularly targeted therapies. Such a system has as its core the use
of various types of clinical care data to generate knowledge to improve patient health care and outcomes. Currently, the opportunity for learning about the “real-world” clinical use and treatment outcomes of biomarker tests for molecularly targeted therapies is not being realized, and there is an urgent need for a framework to capture this critical information. The learning health care system concept, and the committee’s adaptation of the approach to create a rapid learning system to improve the development and use of biomarker tests for molecularly targeted therapies, is described further in Chapter 2 of this report.
ORGANIZATION OF THE REPORT
This report reviews the literature on biomarker tests for molecularly targeted therapies, presents the committee’s findings, and offers recommendations to federal agencies and private organizations, health care providers, EHR developers and vendors, and professional societies. The rapid learning system framework emphasizes the interrelated nature of the policy issues affecting the clinical development and use of biomarkers for molecularly targeted therapies. This study report is organized around the committee’s rapid learning system framework and contains five chapters.
This introductory chapter describes the context of the study and discusses the committee’s charge and the scope, definition of terms, conceptual framework, and methods of the study. Chapter 2 provides an overview of the concept of a learning health care system and offers illustrative examples of such systems that inform the committee’s vision of a rapid learning system for biomarker tests for molecularly targeted therapies. The three components of this rapid learning system are policy environment, data infrastructure, and processes to improve patient care, and are discussed in the Chapters 3, 4, and 5, respectively. Chapter 3 explores the regulatory and reimbursement policy environment influencing the use of biomarker tests for molecularly targeted therapies. Chapter 4 discusses the challenges involved in the development and implementation of a supportive data infrastructure. Chapter 5 focuses on the final component of the committee’s vision of a rapid learning health system for biomarker tests for molecularly targeted therapies: processes to improve patient care. Chapters 3 through 5 of the report contain discussion of the challenges facing the effective clinical use of biomarker tests for molecularly targeted therapies, as well as the committee’s recommended approach to those challenges.
This report includes three appendixes. Appendix A contains biographical sketches of the committee members and the IOM project staff. Appendix B contains an overview of coding issues related to biomarker
tests for molecularly targeted therapies. Appendix C contains a list of speakers at the committee’s public information-gathering sessions.
REFERENCES
Agarwal, A., D. Ressler, and G. Snyder. 2015. The current and future state of companion diagnostics. Pharmacogenomics and Personalized Medicine 8:99-110.
Alexander, L. 2016. Chairman Alexander announces committee schedule for “step by step” consideration of biomedical innovation bills. https://www.help.senate.gov/chair/newsroom/press/chairman-alexander-announces-committee-schedule-for-step-by-stepconsideration-of-biomedical-innovation-bills (accessed January 20, 2016).
Armstrong, K. 2012. Can genomics bend the cost curve? Journal of the American Medical Association 307(10):1031-1032.
Biomarkers Definitions Working Group. 2001. Biomarkers and surrogate endpoints: Preferred definitions and conceptual framework. Clinical Pharmacology & Therapeutics 69(3): 89-95.
Blumenthal, D., M. Abrams, and R. Nuzum. 2015. The Affordable Care Act at 5 years. New England Journal of Medicine 372(25):2451-2458.
Chong, C. R., and P. A. Janne. 2013. The quest to overcome resistance to EGFR-targeted therapies in cancer. Nature Medicine 19(11):1389-1400.
CMS (Centers for Medicare & Medicaid Services). 2014. The nation’s health dollar ($2.9 trillion), calendar year 2013. https://www.cms.gov/Research-Statistics-Data-and-Systems/Statistics-Trends-and-Reports/NationalHealthExpendData/Downloads/PieChartSourcesExpenditures2013.pdf (accessed September 25, 2015).
Collins, F. S., and H. Varmus. 2015. A new initiative on precision medicine. New England Journal of Medicine 372(9):793-795.
de Gramont, A., S. Watson, L. M. Ellis, J. Rodon, J. Tabernero, A. de Gramont, and S. R. Hamilton. 2015. Pragmatic issues in biomarker evaluation for targeted therapies in cancer. Nature Reviews Clinical Oncology 12(4):197-212.
Fairclough, R. J., M. J. Wood, and K. E. Davies. 2013. Therapy for Duchenne muscular dystrophy: Renewed optimism from genetic approaches. Nature Reviews Genetics 14(6):373-378.
FDA (Food and Drug Administration). 2014. In vitro companion diagnostic devices guidance: Draft guidance for industry and Food and Drug Administration staff. http://www.fda.gov/downloads/medicaldevices/deviceregulationandguidance/guidancedocuments/ucm262327.pdf (accessed June 14, 2015).
Frueh, F. W. 2013. Regulation, reimbursement, and the long road of implementation of personalized medicine—a perspective from the United States. Value Health 16(6 Suppl):S27-S31.
Garraway, L. A., J. Verweij, and K. V. Ballman. 2013. Precision oncology: An overview. Journal of Clinical Oncology 31(15):1803-1805.
Harris, L., H. Fritsche, R. Mennel, L. Norton, P. Ravdin, S. Taube, M. R. Somerfield, D. F. Hayes, R. C. Bast, Jr., and American Society of Clinical Oncology. 2007. American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer. Journal of Clinical Oncology 25(33):5287-5312.
Hayden, E. C. 2014. Technology: The $1,000 genome. Nature 507(7492):294-295.
Hayes, D. F. 2013. From genome to bedside: Are we lost in translation? Breast 22(Suppl 2):S22-S26.
Hayes, D. F., J. Allen, C. Compton, G. Gustavsen, D. G. Leonard, R. McCormack, L. Newcomer, K. Pothier, D. Ransohoff, R. L. Schilsky, E. Sigal, S. E. Taube, and S. R. Tunis. 2013. Breaking a vicious cycle. Science Translational Medicine 5(196):196cm.
HGP (Human Genome Project). 2012. Human genome project information archive: Glossary. http://web.ornl.gov/sci/techresources/Human_Genome/glossary.shtml (accessed August 31, 2015).
Hyman, D. M., I. Puzanov, V. Subbiah, J. E. Faris, I. Chau, J. Y. Blay, J. Wolf, N. S. Raje, E. L. Diamond, A. Hollebecque, R. Gervais, M. E. Elez-Fernandez, A. Italiano, R. D. Hofheinz, M. Hidalgo, E. Chan, M. Schuler, S. F. Lasserre, M. Makrutzki, F. Sirzen, M. L. Veronese, J. Tabernero, and J. Baselga. 2015. Vemurafenib in multiple nonmelanoma cancers with BRAF V600 mutations. New England Journal of Medicine 373(8):726-736.
IOM (Institute of Medicine). 2007. Cancer biomarkers: The promises and challenges of improving detection and treatment. Washington, DC: The National Academies Press.
IOM. 2009. Assessing and improving value in cancer care: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2010a. Evaluation of biomarkers and surrogate endpoints in chronic disease. Washington, DC: The National Academies Press.
IOM. 2010b. Policy issues in the development of personalized medicine in oncology: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2012a. Evolution of translational omics: Lessons learned and the path forward. Washington, DC: The National Academies Press.
IOM. 2012b. Genome-based diagnostics: Clarifying pathways to clinical use: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2012c. Genome-based therapeutics: Targeted drug discovery and development: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2012d. Integrating large-scale genomic information into clinical practice: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2013a. Best care at lower cost: The path to continuous learning health care in America. Washington, DC: The National Academies Press.
IOM. 2013b. Delivering affordable cancer care in the 21st century: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2013c. Delivering high-quality cancer care: Charting a new course for a system in crisis. Washington, DC: The National Academies Press.
IOM. 2013d. The economics of genomic medicine: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2013e. Genome-based diagnostics: Demonstrating clinical utility in oncology: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2014a. Assessing genomic sequencing information for health care decisions: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2014b. Refining processes for co-development of companion diagnostic tests: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2015a. Genomics-enabled learning health care systems: Gathering and using genomic information to improve patient care and research: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2015b. Improving genetics education in graduate and continuing health professional education: Workshop summary. Washington, DC: The National Academies Press.
IOM. 2015c. Policy issues in the development and adoption of biomarkers for molecularly targeted cancer therapies: Workshop summary. Washington, DC: The National Academies Press.
Jain, S., and W. J. Gradishar. 2014. The application of Oncotype DX in early-stage lymph-node-positive disease. Current Oncology Reports 16(1):360.
Jameson, J. L., and D. L. Longo. 2015. Precision medicine—personalized, problematic, and promising. New England Journal of Medicine 372(23):2229-2234.
Joyner, M. J., and N. Paneth. 2015. Seven questions for personalized medicine. Journal of the American Medical Association 314(10):999-1000.
Kohane, I. S. 2015. Health care policy. Ten things we have to do to achieve precision medicine. Science 349(6243):37-38.
Kuykendall, A., and A. Chiappori. 2014. Advanced EGFR mutation-positive non-small-cell lung cancer: Case report, literature review, and treatment recommendations. Cancer Control 21(1):67-73.
Lyman, G. H., L. E. Cosler, N. M. Kuderer, and J. Hornberger. 2007. Impact of a 21-gene rt-PCR assay on treatment decisions in early-stage breast cancer: An economic analysis based on prognostic and predictive validation studies. Cancer 109(6):1011-1018.
Marcus, A. 2015. The challenges of precision. The Scientist Magazine, April 1, 2015.
McShane, L. M., M. M. Cavenagh, T. G. Lively, D. A. Eberhard, W. L. Bigbee, P. M. Williams, J. P. Mesirov, M. Y. C. Polley, K. Y. Kim, J. V. Tricoli, J. M. G. Taylor, D. J. Shuman, R. M. Simon, J. H. Doroshow, and B. A. Conley. 2013. Criteria for the use of omics-based predictors in clinical trials: Explanation and elaboration. BMC Medicine 11(1):220.
Mendelsohn, J. 2013. Personalizing oncology: Perspectives and prospects. Journal of Clinical Oncology 31(15):1904-1911.
Meric-Bernstam, F., L. Brusco, K. Shaw, C. Horombe, S. Kopetz, M. A. Davies, M. Routbort, S. A. Piha-Paul, F. Janku, N. Ueno, D. Hong, J. De Groot, V. Ravi, Y. Li, R. Luthra, K. Patel, R. Broaddus, J. Mendelsohn, and G. B. Mills. 2015. Feasibility of large-scale genomic testing to facilitate enrollment onto genomically matched clinical trials. Journal of Clinical Oncology 33(25):2753-2762.
Mirnezami, R., J. Nicholson, and A. Darzi. 2012. Preparing for precision medicine. New England Journal of Medicine 366(6):489-491.
The Moran Company. 2013. Results of analyses for chemotherapy administration utilization and chemotherapy drug utilization, 2005-2011 for Medicare fee-for-service beneficiaries. The Moran Group.
Murthy, V. H., H. M. Krumholz, and C. P. Gross. 2004. Participation in cancer clinical trials: Race-, sex-, and age-based disparities. Journal of the American Medical Association 291(22):2720-2726.
NCI (National Cancer Institute). 2010. NCI community cancer centers program pilot: 2007-2010. http://ncccp.cancer.gov/Media/FactSheet.htm (accessed January 20, 2016).
NRC (National Research Council). 2011. Toward precision medicine: Building a knowledge network for biomedical research and a new taxonomy of disease. Washington, DC: The National Academies Press.
OPS (Office of the Press Secretary). 2015. Fact sheet: President Obama’s Precision Medicine Initiative. https://www.whitehouse.gov/the-press-office/2015/01/30/fact-sheet-president-obama-s-precision-medicine-initiative (accessed August 10, 2015).
Parkinson, D. R., R. T. McCormack, S. M. Keating, S. I. Gutman, S. R. Hamilton, E. A. Mansfield, M. A. Piper, P. DeVerka, F. W. Frueh, J. M. Jessup, L. M. McShane, S. R. Tunis, C. C. Sigman, and G. J. Kelloff. 2014. Evidence of clinical utility: An unmet need in molecular diagnostics for patients with cancer. Clinical Cancer Research 20(6):1428-1444.
PCAST (President’s Council of Advisors on Science and Technology). 2008. Priorities for personalized medicine. https://www.whitehouse.gov/files/documents/ostp/PCAST/pcast_report_v2.pdf (accessed April 21, 2015).
Poste, G. 2011. Bring on the biomarkers. Nature 469(7329):156-157.
Prahallad, A., C. Sun, S. Huang, F. Di Nicolantonio, R. Salazar, D. Zecchin, R. L. Beijersbergen, A. Bardelli, and R. Bernards. 2012. Unresponsiveness of colon cancer to BRAF(V600E) inhibition through feedback activation of EGFR. Nature 483(7387):100-103.
Ramsey, S. D., and S. D. Sullivan. 2014. A new model for reimbursing genome-based cancer care. Oncologist 19(1):1-4.
Redberg, R. F., and S. S. Dhruva. 2015. The FDA’s medical device problem. The New York Times, July 17, 2015.
Romanus, D., S. Cardarella, D. Cutler, M. B. Landrum, N. I. Lindeman, and G. S. Gazelle. 2015. Cost-effectiveness of multiplexed predictive biomarker screening in non-small-cell lung cancer. Journal of Thoracic Oncology 10(4):586-594.
Rubin, R. 2015. Precision medicine: The future or simply politics? Journal of the American Medical Association 313(11):1089-1091.
Sawyers, C. L. 2008. The cancer biomarker problem. Nature 452(7187):548-552.
Schott, A. F., C. M. Perou, and D. F. Hayes. 2015. Genome medicine in cancer: What’s in a name? Cancer Research 75(10):1930-1935.
Simonds, N. I., M. J. Khoury, S. D. Schully, K. Armstrong, W. F. Cohn, D. A. Fenstermacher, G. S. Ginsburg, K. A. Goddard, W. A. Knaus, G. H. Lyman, S. D. Ramsey, J. Xu, and A. N. Freedman. 2013. Comparative effectiveness research in cancer genomics and precision medicine: Current landscape and future prospects. Journal of the National Cancer Institute 105(13):929-936.
Sparano, J. A., R. J. Gray, D. F. Makower, K. I. Pritchard, K. S. Albain, D. F. Hayes, C. E. Geyer, Jr., E. C. Dees, E. A. Perez, J. A. Olson, Jr., J. Zujewski, T. Lively, S. S. Badve, T. J. Saphner, L. I. Wagner, T. J. Whelan, M. J. Ellis, S. Paik, W. C. Wood, P. Ravdin, M. M. Keane, H. L. Gomez Moreno, P. S. Reddy, T. F. Goggins, I. A. Mayer, A. M. Brufsky, D. L. Toppmeyer, V. G. Kaklamani, J. N. Atkins, J. L. Berenberg, and G. W. Sledge. 2015. Prospective validation of a 21-gene expression assay in breast cancer. New England Journal of Medicine 373(21):2005-2014.
Teutsch, S. M., L. A. Bradley, G. E. Palomaki, J. E. Haddow, M. Piper, N. Calonge, W. D. Dotson, M. P. Douglas, A. O. Berg, and E. W. Group. 2009. The evaluation of genomic applications in practice and prevention (EGAPP) initiative: Methods of the EGAPP working group. Genetics in Medicine 11(1):3-14.
Towse, A., D. Ossa, D. Veenstra, J. Carlson, and L. Garrison. 2013. Understanding the economic value of molecular diagnostic tests: Case studies and lessons learned. Journal of Personalized Medicine 3(4):288-305.
Trosman, J. R., C. B. Weldon, J. C. Schink, W. J. Gradishar, and A. B. Benson, 3rd. 2013. What do providers, payers and patients need from comparative effectiveness research on diagnostics? The case of HER2/Neu testing in breast cancer. Journal of Comparative Effectiveness Research 2(4):461-477.
Trosman, J. R., K. A. Phillips, C. B. Weldon, and R. K. Kelley. 2015. Challenges of coverage policy development for next-generation tumor sequencing panels: Experts and payers weigh in. Journal of the National Comprehensive Cancer Network 13(3):311-318.
Yu, P. P., M. A. Hoffman, and D. F. Hayes. 2015. Biomarkers and oncology: The path forward to a learning health system. Archives of Pathology & Laboratory Medicine 139(4):451-456.
This page intentionally left blank.
























