2
Crops
1. INTRODUCTION
Crop agriculture is one of America’s success stories. U.S. farmers produce 34.1 percent of the world’s soybeans, 35.5 percent of the world’s corn, 13.4 percent of the world’s cotton, and 7.6 percent of the world’s wheat (USDA-FAS, 2018b). Crop production accounts for approximately $194 billion per year in agricultural cash receipts (USDA-ERS, 2018). In 2016, agricultural domestic exports of crops reached $108 billion ($21 billion in soybeans alone) and created an estimated $171.3 billion in additional economic activity (USTR, 2017; USDA-FAS, 2018a).
Yields of major staple crops grown in the United States are the highest or near to the highest in the world. Since the 1930s, yields of corn have increased more than eightfold, while soybean and cotton yields have increased more than fourfold, due primarily to advances in plant breeding, as well as fertilizer use and equipment efficiency, among other innovations (Fernandez-Cornejo, 2004; Nielsen, 2017). Despite successive years of weather disasters, the yield performance of America’s premier crops is a remarkable testament to the resilience and success of the varieties commercially available today (see Figure 2-1).
With increased support for public-sector agricultural research and public–private partnerships, it will be possible to bring the breeding success in corn to many other crops (such as cover crops, fruits and vegetables, and bioenergy crops) at a much faster pace than is currently possible. Moreover, these efforts can be used to “harden” crops against the effects of extreme weather and increased pest and disease pressure while maximizing yield and
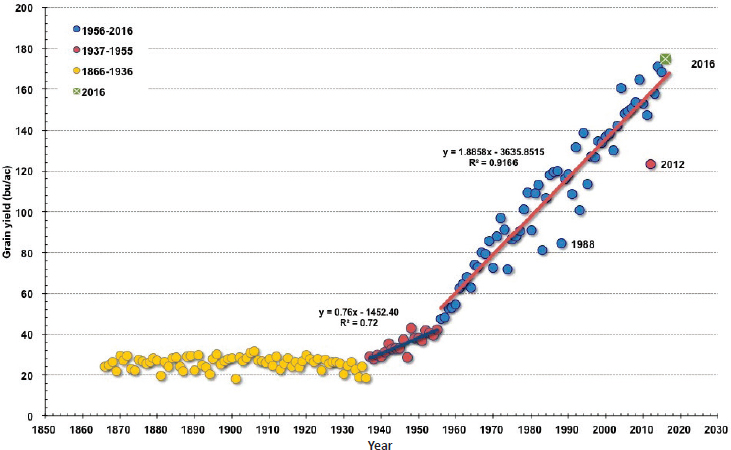
SOURCE: Nielsen, 2017.
increasing nutritional content and flavor. These efforts can also be used to enhance other crop characteristics, such as those that will reduce the need for costly inputs. As stationary organisms, plants exhibit enormous plasticity in the face of adverse and extreme environments. Through the process of natural selection, plant communities adapt to local conditions and provide for their sustenance by interacting with their environments both below and above ground. Plants have evolved to grow and thrive in almost all environments on this planet—from lands with virtually no water to lands that flood routinely, and in all temperature extremes. This evolutionary pressure has given rise to the incredible genetic diversity of plant life that, when coupled with new gene-editing technologies, offers exciting new avenues for solving some of the big challenges facing crop production today.
Other chapters in this report will address soil, fertilizer use (see Chapter 5), and water (see Chapter 6), proper management of which is critical for sustainable, environmentally responsible crop production. There is a pressing need for changes in agronomic practices that improve the stewardship of natural resources and decrease the environmental impacts of crop production. The focus of this chapter is on the crop toolbox and how crop germplasm can be developed over the next two decades—efforts that can be pursued in conjunction with agronomy and crop science to facilitate increased diversity and greater flexibility in cropping systems. The
domestication of wild plant species suited to different environments has commercial value in crop rotations and these species can serve as habitats for beneficial insects, with the potential to expand options for crop management. Similarly, the genetic tools are rapidly becoming available to endow current cultivars with traits that minimize the need for inputs (e.g., water, nitrogen, phosphate, pesticides, and fungicides), maximize yield in changing environments/climatic extremes, resist diseases and pests, and provide improved nutrition, among other characteristics.
2. CHALLENGES
It is no longer safe to assume that U.S. crop production is in a self-sustaining, steady state. First and foremost, the natural resource base for U.S. agriculture is recognized as increasingly fragile. Groundwater and fertile soil are finite resources, and their use and misuse define the boundaries of sustainable production in the long term (Rockström et al., 2017). Aquifers supporting the majority of U.S. production are being drained (Konikow, 2013; Steward et al., 2013), and soil quality in some parts of the country is degrading (Baumhardt et al., 2015). These developments set the stage for lower productivity and the need for crops that can perform well in less than optimal environments.
Second, crop systems are also stressed by changing weather patterns and extreme weather events (Walthall et al., 2012). Prolonged drought over the past decade (Diffenbaugh et al., 2015; Howitt et al., 2015) and flooding more recently (Mallakpour and Villarini, 2015) have been responsible for the largest proportion of U.S. crop disaster payments from abiotic and biotic stresses (see Figure 2-2). The need for crops that are resilient to multiple abiotic stresses, such as drought and flooding, will be a challenging problem for breeders over the next decade. There are success stories in addressing some of the challenges, such as the development of flood-tolerant varieties of rice via introduction of the SUB1A gene into modern cultivars (Bailey-Serres et al., 2012; see Box 2-1).
Third, biotic challenges to crops including pests and diseases are an increasing threat due to agricultural intensification, expanding global trade, and extreme weather. A recent example is Huanglongbing (HLB), also known as citrus greening disease. HLB was detected in Asia more than 100 years ago and was first seen in the United States in Florida orchards in 2005. Within 3 years the vector-transmitted bacterial pathogen spread to the majority of citrus orchards in Florida, leading to losses of more than $1 billion annually (Court et al., 2017). There is no known cure for HLB, which kills trees in 3-5 years and has now spread to California, Georgia, Hawaii, Louisiana, South Carolina, and Texas (NASEM, 2018).
In another example, rising temperatures are leading to migrations of
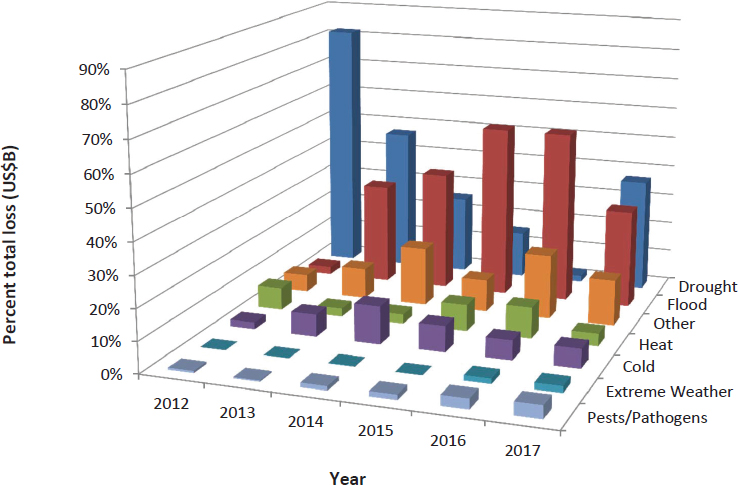
SOURCE: USDA-RMA, 2018.
insect pests to higher elevations, while greater intensification is leading to more rapid evolution and spread of diseases (Katsaruware-Chapoto et al., 2017). Direct yield losses caused by pathogens and pests are predicted to lower global agricultural productivity in the future by 20 to 40 percent (Savary et al., 2012; Baltes et al., 2017). Some natural threats to crop production have been made worse by management practices, such as the emergence of herbicide-resistant weeds following overuse of a single herbicide (e.g., glyphosate) to control weeds. Improved resistance management practices can reduce input costs, improve yields, and increase returns (Livingston et al., 2015).
There is a deep sense of urgency and opportunity to speed up and expand the power of crop improvement to address these challenges. As previously mentioned, global food production will need to increase by at least 50 percent by 2030 to feed a growing global population (UN DESA, 2017). However, the current pace of yield growth worldwide may not be sufficient to meet the predicted need for 2030 and beyond (Ray et al., 2012), and there are even signs that the rate of yield increase in many crops is beginning to plateau (Wei et al., 2015; Andersen et al., 2018). The question of why yield growth is slowing has yet to be answered, with speculation ranging from the decline in agricultural research for basic crop improvement (Pardey and Beddow, 2013; Andersen, 2015), temperature and other effects of climate change (Lobell and Field, 2007), or that some crops are nearing their biological yield potential (Evans and Fischer, 1999; Alston et al., 2009; Ort et al., 2015).
3. OPPORTUNITIES
3.1 Gene-Editing Systems
The discovery of gene-editing systems (such as CRISPR-Cas9) has revolutionized our ability to both understand and genetically modify both plants and animals (Yin et al., 2017). For crops, new alleles can be generated and introduced directly into a cultivar of choice, leaping over the time-consuming process of making multiple crosses to combine desirable traits in the progeny. Gene editing creates the potential to identify and implement new traits in the field on a much faster timescale. Traditional plant breeding is slow and tedious as it can only exploit the limited quantitative trait alleles found in wild relatives, and then it can take between 7 to 12 years to utilize conventional methods to develop a new cultivar (Baenziger et al., 2006). The ability to fine-tune the expression of a quantitative trait locus rather than utilizing only what is available in wild relatives has already shown promise as a way to increase yield. For example, researchers edited genes in three pathways that contribute to productivity in tomato plants—plant architecture, fruit
size, and inflorescence—to rapidly produce alleles that alter their promoters. The resultant plants displayed a series of previously unobserved phenotypes, including several with increased yield (Rodríguez-Leal et al., 2017).
Gene editing has now opened the door for quickly exploring additional changes. Box 2-2 describes a success story of a transgenic approach used in the quest to turn a C3 plant (such as rice) into a C4 plant (such as maize). Gene editing also offers the ability to eliminate linkage drag, a problem that has plagued crop breeding since its inception. Linkage drag occurs when undesirable traits are linked to, and therefore inherited with, desirable traits (Giovannoni, 2018; Zhu et al., 2018b). Using a variety of genomic and associated approaches to characterize a large tomato diversity population, Zhu and colleagues (2018b) were able to “see” the genomic outcomes of tomato domestication, many of which are undesirable. Knowledge of these “hitchhiking genes” is a necessary first step toward their eventual modification or elimination by gene-editing methodologies.
3.2 Development of Dynamic Crops
Agriculture could benefit from crops that provide farmers with the flexibility to change their crop’s physiology in response to unexpected environments. Park and colleagues (2015) designed such a crop whose water use can be controlled by chemical application (see Box 2-3). An even greater
advancement would involve developing crops that can respond in real time to environmental changes (e.g., drought, floods, and temperature extremes) as well as to the appearance of disease organisms and pests. Given the inability to reliably predict the conditions that any particular crop may face over the growing season combined with more extreme weather events predicted over the long term (Walthall et al., 2012), dynamic crops could better respond to such events and become an asset for enhancing food security.
3.3 Mining Plant Diversity
Traditionally, the traits to overcome the many challenges previously noted have come from wild relatives. Although gene editing can eliminate the laborious introgression of traits (at least for crops for which those traits can be successfully introduced; see discussion of transformation later in this chapter), there is a vast array of plant species whose agricultural potential remains untapped (see Box 2-4). There are more than 50,000 edible plants, but only 15 crops are used to meet 90 percent of the world’s energy demands, and 3 commodity crops (rice, maize, and wheat) account for two-thirds of human caloric intake (Gruber, 2017). Additionally, more than 70 percent of wild relatives of domesticated crops are in urgent need of conservation, and are ironically threatened by the expansion of agriculture into natural ecosystems (Castañeda Alvarez et al., 2016). Around the world, gene banks and botanical gardens hold more than 7.4 million seeds or plant tissues from thousands of species (Gruber, 2017). These collections need to be maintained, curated, and explored.
3.4 Taking Advantage of Plant–Microbe Interactions
Billions of microorganisms and macroorganisms (from viruses to nematodes) live on, inside, and near plants, both above and below ground (Leach et al., 2017). Beneficial plant–microbe interactions include direct stimulation of plant growth, protection of plants from pathogens and insect pests through direct production of toxins or through induced resistance in the plant, and improvement of resilience to environmental stress (e.g., drought, salinity) (Parnell et al., 2016; Finkel et al., 2017). Beneficial interactions can occur in the root zone (rhizosphere), on leaf surfaces (phyllosphere), and in internal tissues (endosphere) (APS, 2016).
The greatest density and diversity of microbial life associated with plants occurs in the soil, in the rhizosphere. In leguminous plants, some species of bacteria (Rhizobium) induce the formation of root nodules in a symbiotic relationship that converts atmospheric and largely inert N2 into ammonia (NH3) and other molecular precursors that the plant uses in the biosynthesis of nucleotides, coenzymes, and amino acids. In many more species of plants, fungal symbionts (called arbuscular mycorrhizal fungi) form hyphae that increase the ability of plant roots to access minerals (particularly phosphorus) and water (Leach et al., 2017). It has been proposed that an opportunity lies in domesticating or improving specific plant-associated microbial communities to use them as soil or seed inoculants to improve plant growth (Parnell et al., 2016).
The characterization of microbial communities in concert with plants, which often cannot be cultured outside the plant environment, has been
enabled by high-throughput gene sequencing of community DNA (metagenomics) and RNA (metatranscriptomics) (Schlatter et al., 2015; Nesme et al., 2016). In addition, progress has been made at the intersection of miniaturized microfluidics and imaging that permits real-time monitoring of root interactions with microbial species (Massalha et al., 2017).
Based on knowledge gained from these technologies, a future opportunity is using synthetic biology tools to design plant–microbe associations that improve crop productivity. Such associations can be approached by using gene editing in plants, microbes, or both. For example, plant genes controlling nodule formation by nitrogen-fixing rhizobacteria might be expanded to non-legume crops to reduce the need for fertilizer application, and microbial consortia present in the root zone could be engineered to produce novel plant growth promoters or protectants (Ahkami et al., 2017).
3.5 Making Food More Nutritious
Relatively little breeding effort has gone into making staple crops better sources of vitamins and minerals. More than 2 billion people suffer from micronutrient deficiencies because their plant-based diets do not provide sufficient nutrition. Factors that drive the modern food system include consistency, predictability, low cost, and high-edible yield, but the nutritional value of food produced for direct human consumption has not been a priority of the market (Dwivedi et al., 2017). There have been concerted efforts to survey cultivars for natural variation in levels of select nutrients. For example, a recent study examined vitamin E levels in maize grain and determined that only two loci are responsible for most of the variation (Diepenbrock et al., 2017). This discovery suggests that targeting these loci in any number of crops may increase vitamin E content. There has also been some success using transgenic approaches to increase nutritional content. For example, biofortified rice that meets dietary targets for iron and zinc and has no yield penalty in the field is a major breakthrough in this area (Trijatmiko et al., 2016). There are also opportunities to increase levels of important phytonutrients that have health benefits, such as polyphenols and carotenoids (Martin and Li, 2017; Yu and Tian, 2017).
Unfortunately, breeding almost exclusively for increased yield has made some crops less nutritious. For example, the concentrations of iron, zinc, and selenium in wheat have dropped by 28, 25, and 18 percent, respectively, all in the period between 1920 and 2000 (Gruber, 2016). Such decreases are thought to be a dilution effect as improvements in the amount of fixed carbon have been made without a proportional increase in mineral content (Marles, 2017). This results in lower mineral concentrations when expressed on a dry weight basis. Furthermore, recent studies are projecting that levels of zinc and iron in grains and legumes will continue to decline as
CO2 levels continue to rise (Myers et al., 2014; Zhu et al., 2018a). Because most of the world relies on plants as their dietary source of these micronutrients, such decreases are a cause for concern. A key scientific question is whether these declines in micronutrients can be reversed with changes in plant traits. Therefore, more attention needs to be paid to the unintended consequences of breeding exclusively for yield.
3.6 Optimizing Crop Production Systems
The gap between actual yield and yield potential can be accounted for by several sources of variance, including genetics (G), environment (E), management practices (M), and socioeconomic factors (S). Numerous enabling technologies can be implemented in crop production systems to achieve both resilience and sustainability and to enable crop yields to reach their full potential. These include the use of precision agriculture to manage water and fertilizer use, and the use of data science to integrate information from field- and plant-based sensors and weather prediction parameters (for a more complete discussion, see Chapter 5, section 3.1, “Leveraging Advances in Microelectronics, Sensing, and Modeling” and Chapter 6, “Water-Use Efficiency and Productivity”). Breakthroughs in genomics, nanotechnology, and robotics along with improvements in computational, statistical, and modeling capabilities will make it possible for scientists and producers to make well-informed, data-driven decisions. For example, as it will be discussed in Chapter 7, development of high-throughput automated phenotyping capabilities can speed the process of breeding via the use of artificial intelligence and machine learning. However, in order to successfully model, manage, and predict crop production in any given location, better information is also needed on how different cropping management systems (e.g., use of cover crops and crop rotation) influence soil properties such as water storage capacity and nutrient availability.
The emerging field of plant nanobiotechnology promises transformative solutions for nondestructive monitoring of plant signaling pathways and metabolism (Kwak et al., 2017). This can increase plant tolerance (e.g., drought, disease, and soil nutrient deficiencies [Elmer and White, 2016; Wu et al., 2017]), alter photosynthesis (Giraldo et al., 2014), and enable plants to communicate their biochemical status (Wong et al., 2017). These discoveries will lead to the creation of “smart” plants that are more resilient to climate-induced stresses. Better understanding of the fundamental biophysical processes controlling nanomaterial–plant interactions will enable delivery of nanomaterials to precise locations in plants where they are needed to be active. The continuous real-time monitoring of plant heat status and the ability to combine these nano-enabled technologies with wireless soil sensors and automated water and nutrient delivery systems can lead to more precise
delivery of nutrients and water, leading to more efficient use of inputs and greater yields.
To ensure that all aspects of GEMS are addressed in a comprehensive manner, there is an opportunity to assemble teams (including geneticists, soil scientists, agronomists, plant pathologists, and entomologists) to work together to evaluate the response of different crop genotypes to environmental stresses and crop management practices (Hatfield and Walthall, 2015), and to factor in socioeconomic issues such as costs, access to small growers, and technology adoption. Such a systems approach can help to identify the optimum combination of G × M × S for the current and anticipated E.
3.7 Controlled Environment Agriculture (CEA)
Controlled environment agriculture1 (CEA) offers systems-level opportunities to increase the sustainability of some crops (e.g., fruits, vegetables, and herbs) by providing resource-efficient farming systems with respect to water and nutrient use. CEA can close the loop on nitrogen and phosphorus use (Thomaier et al., 2015), reduce food miles (Weber and Matthews, 2008; Cleveland et al., 2011; Nicholson et al., 2015), and lower water exports from water-poor regions to water-rich regions. Locating food production in urban areas could also help to lower food waste by decreasing the amount of time from harvest to consumption. CEA can also address important food safety issues such as Escherichia coli outbreaks (CDC, 2018). CEA can provide year-long growing seasons and protection against pests and diseases. Genetic approaches described in this chapter can be used to provide traits to plants to make them suitable for growth in CEA. The importance of CEA for improving water-use efficiency is discussed in Chapter 6.
4. GAPS
There is growing excitement in the plant sciences and breeding communities that we are on the cusp of a second green revolution. While the first green revolution was facilitated by the introduction of genes from other varieties and wild relatives, this second revolution will be fueled by basic research discoveries with model organisms and analyses of massive datasets that will combine to identify genes and regulatory sequences for targeted editing. However, the following gaps in knowledge and capabili-
___________________
1 Controlled environment agriculture includes indoor production systems that range from low-technology covers for field crops to highly automated greenhouses, vertical farms, and recirculating aquaculture systems (RASs). The latter use the latest advances in hydroponics or aquaponics to increase productivity and improve water- and nutrient-use efficiencies. For example, a RAS is coupled with hydroponic plant production by using waste derived from fish production as fertilizer for plant growth (Badiola et al., 2018; Palm et al., 2018).
ties will have to be addressed before these technologies can be applied to a diversity of crops.
- Knowledge of the genetic basis of the phenotypes and of allelic variants. Although the functions of many plant genes are known from studies in the major model plants, such as Arabidopsis, there is a lack of basic knowledge of the genetic basis of many traits in crop species. It logically follows that understanding gene function is necessary for the use of gene editing to create desirable traits. However, a gene-editing approach can be used as a discovery tool to knock out genes in crop plants to better understand their functional roles. Additionally, because the precision of gene editing permits the modification of as little as a single base pair in the genome, it can be used to create allelic variants, essentially producing a library of diverse forms of the same gene, with which to explore function and phenotype.
- Phenotyping: Connecting genomic variation with phenotypic impact. Identification of agriculturally desirable traits will require the design and construction of above- and belowground phenotyping facilities and the development of data science that can correlate phenotype to genotype from vast visual, physiological, and “omics” resources. There is a need to increase the speed to characterize phenotype, to use new technologies that include overhead imaging (e.g., photo from a drone) and underground imaging to detect morphologies and traits, including chemical exudates (Das et al., 2015; Shi et al., 2016).
- Ability to improve transformation efficiency of more plant species. Gene editing requires an ability to introduce DNA into plant tissue (called genetic transformation) and, in most cases, regenerate the tissues into transformed plants. Unfortunately, despite substantial progress in sequencing, assembling, and annotating genomes from a vast array of plant species, significant bottlenecks exist in the successful genetic transformation of most crops. Research is needed to improve plant cell and tissue culture, identify better methods of introducing genetic material, and modulate the plant development pathway to improve the receptivity, stability, and regrowth of the transformed tissue (Altpeter et al., 2016). Studies of how plants regenerate themselves from cuttings or after injury, for example, have begun to provide insight into the genetic network that controls cellular proliferation and reprogramming. Such insights could lead to better methods to increase the chances of growing whole plants from transformed cells and tissues (Ikeuchi et al., 2018). A second green revolution powered by gene editing will not be pos-
-
sible unless facile transformation protocols are developed for use on any strain of any crop in any laboratory. Similarly, the inability to transform wild plant species will prevent the use of gene editing to accelerate their domestication.
- Workforce, education, and training (and funding). Advances arising out of basic science and technology remain relatively new and underexploited for germplasm improvement for most crops. This is due in part to the relatively small community of plant scientists educated to use science and technology, and the limited public and private investments for plant trait discovery and for translating such knowledge to enable crop discoveries. Investment would be necessary to maintain a pipeline of investigators who can push the limits of scientific inquiry and train the next generation of advanced plant scientists (USDA, 2015).
5. RECOMMENDATIONS FOR NEXT STEPS
Despite steady decreases in the funding of both basic and applied crop science research, U.S. scientists have continued to innovate and make cutting-edge discoveries. However, without sufficient support, U.S. plant scientists are likely to fall behind their counterparts in other countries in fully realizing gene editing as both a basic discovery and crop improvement tool. Since 2009, plant science funding in China has quadrupled and is supported with infrastructure, eclipsing the U.S. investment. Notable advancements in all aspects of plant science have been forthcoming from the Chinese (Chong and Xu, 2014). Recognizing that food security and international competitiveness are critical components of national security, the following steps are recommended to secure U.S. leadership in crop improvement:
- Continue to genetically dissect and then introduce desirable traits (increased photosynthetic efficiency; drought and flood tolerance; temperature extremes tolerance; disease and pest resistance; and improved taste, aroma, and nutrition) and remove undesirable traits from crop plants through the use of both traditional genetic approaches and targeted gene editing. The goal will be to
- expand the number of alleles of known breeding improvement genes from what have been introgressed from wild relatives;
- engineer below- and aboveground plant architecture, by obtaining and applying basic knowledge of plant root, shoot, and influorescence development;
- modify the plant microbiome to enhance desirable crop traits, including resistance to disease and increased nutrient-use efficiency; and
-
- remove undesirable traits (due to negative epistasis) that are tightly linked to alleles selected during domestication.
- Enable routine genome modification of all crop plants through the development of facile transformation and regeneration technologies. Achieving this goal will require research into improved plant cell and tissue culture, better methods of introducing genetic material, and the ability to modulate plant development to improve the regeneration of the transformed tissue into whole plants. It will also require development of facilities for more rapid phenotype detection and analysis under different environmental (soil, climate, and moisture) conditions and management regimes.
- Monitor plant stress and nutrients through the development of novel sensing technologies, and allow plants to better respond to environmental challenges (heat, drought, flood, pests, and nutrient requirements) by exploring the use of nanotechnology, synthetic biology, and the plant microbiome to develop dynamic crops that can turn certain functions on or off only when needed. Developing dynamic crops will require complementary approaches to gene editing, including harnessing beneficial plant–root–soil–microbe interactions to enhance desirable crop traits (such as drought resistance, disease resistance, and nutrient-use efficiency), using novel sensing technologies to sense plant nutrient status and stress, and using nanotechnologies for delivering nutrients and managing plant stress.
Using the genetic diversity of plant and microbial life available together with new molecular and other tools is key for unlocking many opportunities for crop improvement. The process of prioritizing the most important opportunity to pursue would be best informed by thoughtful analyses of the targeted crops and their prospective new traits from a systems perspective, as described in Chapter 9. This includes envisioning their performance and impact in the context of the agroecological system interfacing with humans and socioeconomic considerations that play a role.
REFERENCES
Ahkami, A. H., R. A. White III, P. P. Handakumbura, and C. Jansson. 2017. Rhizosphere engineering: Enhancing sustainable plant ecosystem productivity. Rhizosphere 3(2):233-243.
Alston, J. M., J. M. Beddow, and P. G. Pardey. 2009. Agricultural research, productivity, and food prices in the long run. Science 325(5945):1209-1210.
Altpeter, F., N. M. Springer, L. E. Bartley, A. E. Blechl, T. P. Brutnell, V. Citovsky, L. J. Conrad, S. B. Gelvin, D. P. Jackson, A. P. Kausch, P. G. Lemaux, J. I. Medford, M. L. Orozco-Cárdenas, D. M. Tricoli, J. Van Eck, D. F. Voytas, V. Walbot, K. Wang, Z. J. Zhang, and C. N. Stewart. 2016. Advancing crop transformation in the era of genome editing. Plant Cell 28(7):1510-1520.
Andersen, M. A. 2015. Public investment in U.S. agricultural R&D and the economic benefits. Food Policy 51:38-43. Available at https://doi.org/10.1016/j.foodpol.2014.12.005 (accessed May 8, 2018).
Andersen, M. A., J. M. Alston, P. G. Pardey, and A. Smith. 2018. A century of U.S. farm productivity growth: A surge then a slowdown. American Journal of Agricultural Economics 100(4):1072-1090. Available at https://doi.org/10.1093/ajae/aay023 (accessed May 31, 2018).
APS (American Phytopathological Society). 2016. Phytobiomes: A Roadmap for Research and Translation. St. Paul, MN: American Phytopathological Society. Available at https://www.apsnet.org/members/outreach/ppb/Documents/PhytobiomesRoadmap.pdf (accessed May 8, 2018).
Badiola, M., O. Basurko, R. Piedrahita, P. Hundley, and D. Mendiola. 2018. Energy use in recirculating aquaculture systems (RAS): A review. Aquacultural Engineering 81. doi: 10.1016/j.aquaeng.2018.03.003.
Baenziger, P. S., W. K. Russell, G. L. Graef, and B. T. Campbell. 2006. Improving lives: 50 years of crop breeding, genetics and cytology (C-1). Crop Science 46:2230-2244.
Bailey-Serres, J., S. C. Lee, and E. Brinton. 2012. Waterproofing crops: Effective flooding survival strategies. Plant Physiology 160(4):1698-1709.
Baltes, N. J., J. Gil-Humanes, and D. F. Voytas. 2017. Chapter one–genome engineering and agriculture: Opportunities and challenges. Progress in Molecular Biology and Translational Science 149:1-26.
Baumhardt, R. L., B. A. Stewart, and U. M. Sainju, 2015. North American soil degradation: Processes, practices, and mitigating strategies: A review. Sustainability 7(3):2936-2960.
Blakenship, R. E., and M. Chen. 2013. Spectral expansion and antenna reduction can enhance photosynthesis for energy production. Current Opinion in Chemical Biology 17:457-461.
Castañeda Álvarez, N., C. K. Khoury, H. A. Achicanoy, V. Bernau, H. Dempewolf, R. J. Eastwood, L. Guarino, R. H. Harker, A. Jarvis, N. Maxted, J. V. Müller, J. RamirezVillegas, C. C. Sosa, P. C. Struik, H. Vincent, and J. Toll. 2016. Global conservation priorities for crop wild relatives. Nature Plants 2:16022. doi: 10.1038/nplants. 2016.22.
CDC (Centers for Disease Control and Prevention). 2018. Multistate Outbreak of E. coli O157:H7 Infections Linked to Romaine Lettuce, Investigation Notice: Multistate Outbreak of E. coli O157:H7 Infections April 2018. Available at https://www.cdc.gov/ecoli/2018/o157h7-04-18/index.html (accessed June 4, 2018).
Chong, K. and Z. Xu. 2014. Investment in plant research and development bears fruit in China. Plant Cell Reports 33(4):541-550. doi: 10.1007/s00299-014-1587-6.
Cleveland, D. A., C. N. Radka, N. M. Müller, T. D. Watson, N. J. Rekstein, H. V. Wright, and S. E. Hollingshead. 2011. Effect of localizing fruit and vegetable consumption on greenhouse gas emissions and nutrition, Santa Barbara County. Environmental Science & Technology 45(10):4555-4562.
Court, C. D., A. W. Hodges, M. Ramani, and T. H. Spleen. 2017. Economic Contributions of the Florida Citrus Industry: 2015-2016. Gainsville: University of Florida Food and Economics Department.
Das, A., H. Schneider, J. Burridge, A. K. M. Ascanio, T. Wojciechowski, C. N. Topp, J. P. Lynch, J. S. Weitz, and A. Bucksch. 2015. Digital imaging of root traits (DIRT): A high-throughput computing and collaboration platform for field-based root phenomics. Plant Methods 11:51. Available at http://doi.org/10.1186/s13007-015-0093-3 (accessed June 4, 2018).
Diepenbrock, C. H., C. B. Kandianis, A. E. Lipka, M. Magallanes-Lundback, B. Vaillancourt, E. Góngora-Castillo, J. G. Wallace, J. Cepela, A. Mesberg, P. J. Bradbury, D. C. Ilut, M. Mateos-Hernandez, J. Hamilton, B. F. Owens, T. Tiede, E. S. Buckler, T. Rocheford, C. R. Buell, M. A. Gore, and D. DellaPenna. 2017. Novel loci underlie natural variation in vitamin E levels in maize grain. Plant Cell 29(10):2374-2392.
Diffenbaugh, N. S., D. L. Swain, and D. Touma. 2015. Anthropogenic warming has increased drought risk in California. Proceedings of the National Academy of Sciences of the United States of America 112(13):3931-3936.
Dwivedi, S. L., E. T. Lammerts van Bueren, S. Ceccarelli, S. Grando, H. D. Upadhyaya, and R. Ortiz. 2017. Diversifying food systems in the pursuit of sustainable food production and healthy diets. Trends in Plant Science 22(10):842-856.
Elmer, W. H., and J. C. White. 2016. The use of metallic oxide nanoparticles to enhance growth of tomatoes and eggplants in disease infested soil or soilless medium. Environmental Science: Nano 3:1072-1079.
Evans, L. T., and R. A. Fischer. 1999. Yield potential: Its definition, measurement, and significance. Crop Science 39(6):1544-1551.
Fernandez-Cornejo, J. 2004. The Seed Industry in U.S. Agriculture: An Exploration of Data and Information on Crop Seed Markets, Regulation, Industry Structure, and Research and Development. Washington, DC: USDA Economic Research Service. Available at https://www.ers.usda.gov/publications/pub-details/?pubid=42531 (accessed May 10, 2018).
Finkel, O. M., G. Castrillo, S. Herrera Paredes, I. Salas González, and J. L. Dangl. 2017. Understanding and exploiting plant beneficial microbes. Current Opinion in Plant Biology 38:155-163.
Giovannoni, J. 2018. Tomato multiomics reveals consequences of crop domestication and improvement. Cell 172(1):6-8.
Giraldo, J. P., M. P. Landry, S. M. Faltermeier, T. P. McNicholas, N. M. Iverson, A. A. Boghossian, N. F. Reuel, A. J. Hilmer, F. Sen, J. A. Brew, and M. S. Strano. 2014. Plant nanobionics approach to augment photosynthesis and biochemical sensing. Nature Materials 13:400-408.
Gruber, K. 2016. Re-igniting the green revolution with wild crops. Nature Plants 2:16048.
Gruber, K. 2017. Agrobiodiversity: The living library. Nature 544(7651):S8-S10.
Hatfield, J. L., and C. L. Walthall. 2015. Meeting global food needs: Realizing the potential via genetics × environment × management interactions. Agronomy Journal 107:1215.
Howitt, R., D. MacEwan, J. Medellin-Azuara, J. Lund, and D. Sumner. 2015. Economic Analysis of the 2015 Drought for California Agriculture. Available at https://watershed.ucdavis.edu/files/biblio/Final_Drought%20Report_08182015_Full_Report_WithAppendices.pdf (accessed May 8, 2018).
Ikeuchi, M., M. Shibata, B. Rymen, A. Iwase, A. M. Bågman, L. Watt, D. Coleman, D. S. Favero, T. Takahashi, S. E. Ahnert, S. M. Brady, and K. Sugimoto. 2018. A gene regulatory network for cellular reprogramming in plant regeneration. Plant Cell Physiology 59(4):765-777. doi: 10.1093/pcp/pcy013.
Katsaruware-Chapoto, R. D., P. L. Mafongoya, and A. Gubba. 2017. Responses of insect pests and plant diseases to changing and variable climate: A review. Journal of Agricultural Science 9(12):160.
Konikow, L. F. 2013. Groundwater Depletion in the United States (1900-2008). Scientific Investigations Report 2013-5079. Reston, VA: U.S. Geological Survey. Available at http://pubs.usgs.gov/sir/2013/5079 (accessed May 10, 2018).
Kwak, S.-Y., M. H. Wong, T. T. S. Lew, G. Bisker, M. A. Lee, A. Kaplan, J. Dong, A. T. Liu, V. B. Koman, R. Sinclair, C. Hamann, and M. S. Strano. 2017. Nanosensor technology applied to living plant systems. Annual Review of Analytical Chemistry 10:113-140.
Leach, J. E., L. R. Triplett, C. T. Argueso, and P. Trivedi. 2017. Communication in the phytobiome. Cell 169(4):587-596.
Livingston, M., J. Fernandez-Cornejo, J. Unger, C. Osteen, D. Schimmelpfennig, T. Park, and D. Lambert. 2015. The Economics of Glyphosate Resistance Management in Corn and Soybean Production. Washington, DC: U.S. Department of Agriculture’s Economic Research Service.
Lobell, D. B., and C. B. Field. 2007. Global scale climate–crop yield relationships and the impacts of recent warming. Environmental Research Letters 2(1):014002.
Long, S. 2015. Meeting the global food demand of the future by engineering crop photosynthesis and yield potential. Cell 161(1):56-66.
Mallakpour, I., and G. Villarini. 2015. The changing nature of flooding across the central United States. Nature Climate Change 5(3):250.
Marles, R. 2017. Mineral nutrient composition of vegetables, fruits and grains: The context of reports of apparent historical declines. Journal of Food Composition Analysis 56:93-103.
Martin, C., and J. Li. 2017. Medicine is not health care, food is health care: Plant metabolic engineering, diet and human health. New Phytologist 216(3):699-719.
Massalha, H., E. Korenblum, S. Malitsky, O. H. Shapiro, and A. Aharoni. 2017. Live imaging of root–bacteria interactions in a microfluidics setup. Proceedings of the National Academy of Sciences of the United States of America 114(17):4549-4554.
Myers, S. S., A. Zanobetti, I. Kloog, P. Huybers, A. D. B. Leakey, A. J. Bloom, E. Carlisle, L. H. Dietterich, G. Fitzgerald, T. Hasegawa, N. M. Holbrook, R. L. Nelson, M. J. Ottman, V. Raboy, H. Sakai, K. A. Sartor, J. Schwartz, S. Seneweera, M. Tausz, and Y. Usui. 2014. Increasing CO2 threatens human nutrition. Nature 510(7503):139.
NASEM (National Academies of Sciences, Engineering, and Medicine). 2018. A Review of the Citrus Greening Research and Development Efforts Supported by the Citrus Research and Development Foundation: Fighting a Ravaging Disease. Washington, DC: The National Academies Press.
Nesme, J., W. Achouak, S. N. Agathos, M. Bailey, P. Baldrian, D. Brunel, Å. Frostegård, T. Heulin, J. K. Jansson, E. Jurkevitch, K. L. Kruus, G. A. Kowalchuk, A. Lagares, H. M. Lappin-Scott, P. Lemanceau, D. Le Paslier, I. Mandic-Mulec, J. C. Murrell, D. D. Myrold, R. Nalin, P. Nannipieri, J. D. Neufeld, F. O’Gara, J. J. Parnell, A. Pühler, V. Pylro, J. L. Ramos, L. F. W. Roesch, M. Schloter, C. Schleper, A. Sczyrba, A. Sessitsch, S. Sjöling, J. Sørensen, S. J. Sørensen, C. C. Tebbe, E. Topp, G. Tsiamis, J. D. van Elsas, G. van Keulen, F. Widmer, M. Wagner, T. Zhang, X. Zhang, L. Zhao, Y.-G. Zhu, T. M. Vogel, and P. Simonet. 2016. Back to the future of soil metagenomics. Frontiers in Microbiology 7:73.
Nicholson, C. F., X. He, M. I. Gómez, H. O. Gao, and E. Hill. 2015. Environmental and economic impacts of localizing food systems: The case of dairy supply chains in the northeastern United States. Environmental Science & Technology 49(20):12005-12014.
Nielsen, R. L. 2017. Historical Corn Grain Yields for the U.S. Web publication, Purdue University Agricultural Extension. Available at https://www.agry.purdue.edu/ext/corn/news/timeless/yieldtrends.html (accessed June 13, 2018).
Ort, D. R., S. S. Merchant, J. Alric, A. Barkan, R. E. Blankenship, R. Bock, R. Croce, M. R. Hanson, J. M. Hibberd, S. P. Long, T. A. Moore, J. Moroney, K. K. Niyogi, M. A. J. Parry, P. P. Peralta-Yahya, R. C. Prince, K. E. Redding, M. H. Spalding, K. J. van Wijk, W. F. J. Vermaas, S. von Caemmerer, A. P. M. Weber, T. O. Yeates, J. S. Yuan, and X. G. Zhu. 2015. Redesigning photosynthesis to sustainably meet global food and bioenergy demand. Proceedings of the National Academy of Sciences of the United States of America 112(28):8529-8536.
Palm, H. W., U. Knaus, S. Appelbaum, S. Goddek, S. M. Strauch, T. Vermeulen, M. H. Jijakli, and B. Kotzen. 2018. Towards commercial aquaponics: A review of systems, designs, scales and nomenclature. Aquaculture International 26:813-842.
Pardey, P. G., and J. M. Beddow. 2013. Agricultural Innovation: The United States in a Changing Global Reality. Chicago: Chicago Council on Global Affairs. Available at https://www.thechicagocouncil.org/sites/default/files/Agricultural_Innovation_Final%281%29.pdf (accessed May 8, 2018).
Park, S., F. C. Peterson, A. Mosquna, J. Yao, B. F. Volkman, and S. R. Cutler. 2015. Agrochemical control of plant water use using engineered abscisic acid receptors. Nature 520(7548):545.
Parnell, J. J., R. Berka, H. A. Young, J. M. Sturino, Y. Kang, D. M. Barnhart, and M. V. DiLeo. 2016. From the lab to the farm: An industrial perspective of plant beneficial microorganisms. Frontiers in Plant Science 7:1110.
Ray, D. K., N. Ramankutty, N. D. Mueller, P. C. West, and J. A. Foley. 2012. Recent patterns of crop yield growth and stagnation. Nature Communications 3:1293.
Rockström, J., J. Williams, G. Daily, A. Noble, N. Matthews, L. Gordon, H. Wetterstrand, F. DeClerck, M. Shah, P. Steduto, C. de Fraiture, N. Hatibu, O. Unver, J. Bird, L. Sibanda, and J. Smith 2017. Sustainable intensification of agriculture for human prosperity and global sustainability. AMBIO 46(1):4-17.
Rodríguez-Leal, D., Z. H. Lemmon, J. Man, M. E. Bartlett, and Z. B. Lippman. 2017. Engineering quantitative trait variation for crop improvement by genome editing. Cell 171(2):470-480.
Savary, S., A. Ficke, J. Aubertot, and C. Hollier. 2012. Crop losses due to diseases and their implications for global food production losses and food security. Food Security 4(4):519-537.
Schlatter, D. C., M. G. Bakker, J. M. Bradeen, and L. L. Kinkel. 2015. Plant community richness and microbial interactions structure bacterial communities in soil. Ecology 96(1):134-142.
Schuler, M. L., O. Mantegazza, and A. P. M. Weber. 2016. Engineering C4 photosynthesis into C3 chassis in the synthetic biology age. Plant Journal 87(1):51-65.
Shi, Y., J. A. Thomasson, S. C. Murray, N. A. Pugh, W. L. Rooney, S. Shafian, N. Rajan. G. Rouze, C. L. Morgan, H. L. Neely, A. Rana, M. V. Bagavathiannan, J. Henrickson, E. Bowden, J. Valasek, J. Olsenholler, M. P. Bishop, R. Sheridan, E. B. Putman, S. Popescu, T. Burks, D. Cope, A. Ibrahim, B. F. McCutchen, D. D. Baltensperger, R. V. Avant, M. Vidrine, and C. Yang. 2016. Unmanned aerial vehicles for high-throughput phenotyping and agronomic research. PLoS ONE 11(7):e0159781.
Steward, D. R., P. J. Bruss, X. Yang, S. A. Staggenborg, S. M. Welch, and M. D. Apley. 2013. Tapping unsustainable groundwater stores for agricultural production in the High Plains Aquifer of Kansas, projections to 2110. Proceedings of the National Academy of Sciences of the United States of America 110(37):E3477-E3486.
Thomaier, S., K. Specht, D. Henckel, and A. Dierich. 2015. Farming in and on urban buildings: Present practice and specific novelties of Zero-Acreage Farming (ZFarming). Renewable Agriculture and Food Systems 30:43-54.
Trijatmiko, K. R., C. Dueñas, N. Tsakirpaloglou, L. Torrizo, F. M. Arines, C. Adeva, J. Balindong, N. Oliva, M. V. Sapasap, J. Borrero, J. Rey, P. Francisco, A. Nelson, H. Nakanishi, E. Lombi, E. Tako, R. P. Glahn, J. Stangoulis, P. Chadha-Mohanty, A. A. T. Johnson, J. Tohme, G. Barry, and I. H. Slamet-Loedin. 2016. Biofortified indica rice attains iron and zinc nutrition dietary targets in the field. Scientific Reports 6:19792.
UN DESA (United Nations Department of Economic and Social Affairs). 2017. World Population Prospects: The 2017 Revision, Key Findings and Advance Tables. ESA/P/WP/248. Available at https://esa.un.org/unpd/wpp/publications/Files/WPP2017_KeyFindings.pdf (accessed May 10, 2018).
USDA (U.S. Department of Agriculture). 2015. USDA Roadmap for Plant Breeding. Available at https://www.usda.gov/sites/default/files/documents/usda-roadmap-plant-breeding.pdf (accessed May 8, 2018).
USDA-ERS (U.S. Department of Agriculture’s Economic Research Service). 2018. Farm Income and Wealth Statistics: Cash Receipts by State. Available at https://data.ers.usda.gov/reports.aspx?ID=17843 (accessed June 18, 2018).
USDA-FAS (U.S. Department of Agriculture’s Foreign Agricultural Service). 2018a. Global Agricultural Trade System (GATS) Data. Available at https://apps.fas.usda.gov/gats/detectscreen.aspx?returnpage=default.aspx (accessed May 8, 2018).
USDA-FAS. 2018b. World Agricultural Production. Available at https://apps.fas.usda.gov/psdonline/circulars/production.pdf (accessed May 8, 2018).
USDA-RMA (U.S. Department of Agriculture’s Risk Management Agency). 2018. Cause of loss historical data files. Available at https://www.rma.usda.gov/SummaryOfBusiness/CauseOfLoss (accessed July 11, 2018).
USTR (Office of the U.S. Trade Representative). 2017. USTR Success Stories: Opening Markets for U.S. Agricultural Exports. Available at https://ustr.gov/about-us/policy-offices/press-office/fact-sheets/2017/march/ustr-success-stories-opening-markets-us (accessed May 8, 2018).
Van Eck, J., K. Swartwood, Z. Lemmone, J. Dalrymple, and Z. B. Lippman. 2017. Development of Agrobacterium-mediated transformation of Physalis peruviana and application of CRISPR/Cas9 genome editing. In Vitro Cellular & Developmental Biology—Animal 53(Suppl.):S34-S35.
Walthall, C. L., J. Hatfield, P. Backlund, L. Lengnick, E. Marshall, M. Walsh, S. Adkins, M. Aillery, E. A. Ainsworth, C. Ammann, C. J. Anderson, I. Bartomeus, L. H. Baumgard, F. Booker, B. Bradley, D. M. Blumenthal, J. Bunce, K. Burkey, S. M. Dabney, J. A. Delgado, J. Dukes, A. Funk, K. Garrett, M. Glenn, D. A. Grantz, D. Goodrich, S. Hu, R. C. Izaurralde, R. A. C. Jones, S.-H. Kim, A. D. B. Leaky, K. Lewers, T. L. Mader, A. McClung, J. Morgan, D. J. Muth, M. Nearing, D. M. Oosterhuis, D. Ort, C. Parmesan, W. T. Pettigrew, W. Polley, R. Rader, C. Rice, M. Rivington, E. Rosskopf, W. A. Salas, L. E. Sollenberger, R. Srygley, C. Stöckle, E. S. Takle, D. Timlin, J. W. White, R. Winfree, L. Wright-Morton, and L. H. Ziska. 2012. Climate Change and Agriculture in the United States: Effects and Adaptation. USDA Technical Bulletin 1935. Washington, DC: U.S. Department of Agriculture’s Agricultural Research Service.
Wang, P., R. Khoshravesh, S. Karki, R. Tapia, C. P. Balahadia, A. Bandyopadhyay, W. Paul Quick, R. Furbank, T. L. Sage, and J. A. Langdale. 2017. Recreation of a key step in the evolutionary switch from C3 to C4 leaf anatomy. Current Biology 27(21):3278-3287.
Weber, C. L., and H. S. Matthews. 2008. Food-miles and the relative climate impacts of food choices in the United States. Environmental Science & Technology 42(10):3508-3513.
Wei, X., Z. Zhang, P. Shi, P. Wang, Y. Chen, X. Song, and F. Tao. 2015. Is yield increase sufficient to achieve food security in China? PLoS ONE 10(2):e0116430.
Wong, M. H., J. P. Giraldo, S.-Y. Kwak, V. B. Koman, R. Sinclair, T. T. Salim Lew, G. Bisker, P. Liu, and M. S. Strano. 2017. Nitroaromatic detection and infrared communication from wild-type plants using plant nanobionics. Nature Materials 16(2):264-272.
Wu, H., N. Tito, and J. P. Giraldo. 2017. Anionic cerium oxide nanoparticles protect plant photosynthesis from abiotic stress by scavenging reactive oxygen species. ACS Nano 11(11):11283-11297.
Yang, S., B. Vanderbeld, J. Wan, and Y. Huang. 2010. Narrowing down the targets: Towards successful genetic engineering of drought-tolerant crops. Molecular Plant 3(3):469-490.
Yin, X., A. K. Biswal, J. Dionora, K. M. Perdigon, C. P. Balahadia, S. Mazumdar, C. Chater, H.-C. Lin, R. A. Coe, T. Kretzschmar, J. E. Gray, P. W. Quick, and A. Bandyopadhyay. 2017. CRISPR-Cas9 and CRISPR-Cpf1 mediated targeting of a stomatal developmental gene EPFL9 in rice. Plant Cell Reports 36(5):745-757.
Yu, S., and L. Tian. 2017. Breeding major cereal grains through the lens of nutrition-sensitivity. Molecular Plant 11:23-30.
Zhu, C., K. Kobayashi, I. Loladze, J. Zhu, Q. Jiang, X. Yu, G. Liu, S. Seneweera, K. L. Ebi, A. Drewnowski, N. K. Fukagawa, and L. H. Ziska. 2018a. Carbon dioxide (CO2) levels this century will alter the protein, micronutrients, and vitamin content of rice grins with potential health consequences for the poorest rice-dependent countries. Science Advances 4(5):eaaq1012.
Zhu, G., S. Wang, Z. Huang, S. Zhang, Q. Liao, C. Zhang, T. Lin, M. Qin, M. Peng, C. Yang, X. Cao, X. Han, X. Wang, E. van der Knaap, Z. Zhang, X. Cui, H. Klee, A. R. Fernie, and J. Luo. 2018b. Rewiring of the fruit metabolome in tomato breeding. Cell 172(1):249-261.




















