Wilcox, B. A., and R. R. Colwell. 2005. Emerging and reemerging infectious diseases: biocomplexity as an interdisciplinary paradigm. EcoHealth 2(4):244-257.
Woolhouse, M. E., and S. Gowtage-Sequeria. 2005. Host range and emerging and reemerging pathogens. Emerging Infectious Diseases 11(12):1842-1847.
Yob, J. M., H. Field, A. M. Rashdi, C. Morrissy, B. van der Heide, P. Rota, A. bin Adzhar, J. White, P. Daniels, A. Jamaluddin, and T. Ksiazek. 2001. Nipah virus infection in bats (order Chiroptera) in peninsular Malaysia. Emerging Infectious Diseases 7(3):439-441.
Author contributions: D.G., R.K., S.A., J.R., M.Y.S., D.M., J.M., and D.U.P. designed research; B.A.J., D.G., R.K., S.A., J.R., M.Y.S., D.M., F.M., J.Y., and D.U.P. performed research; B.A.J., D.G., R.K., S.A., J.R., M.Y.S., and D.U.P. analyzed data; and B.A.J., D.G., R.K., S.A., J.R., D.M., and D.U.P. wrote the paper.
The authors declare no conflict of interest.
This article is a PNAS Direct Submission. Jeffrey Sayer (James Cook University, Cairns, QLD) is a guest editor invited by the Editorial Board.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1208059110/-/DCSupplemental.
GLOBAL TRENDS IN EMERGING VIRAL DISEASES OF WILDLIFE ORIGIN
Jonathan Sleeman31and Hon Ip31
Introduction
Fifty years ago, infectious diseases were rarely considered threats to wildlife populations, and the study of wildlife diseases was largely a neglected endeavor. Furthermore, public health leaders at that time had declared that “it is time to close the book on infectious diseases and the war against pestilence won,” a quote attributed to Dr. William H. Stewart in 1967. There is some debate whether he actually said these words; however, they reflect the widespread belief at that time (Spellberg, 2008). Leap forward to today, and the book on infectious diseases has been dusted off. There is general consensus that the global environment favors the emergence of infectious diseases, and in particular, diseases of wildlife origin (Taylor et al., 2001). Examples of drivers of these infectious diseases include climate and landscape changes, human demographic and behavior changes, global travel and trade, microbial adaptation, and lack of appropriate infrastructure for wildlife disease control and prevention (Daszak et al., 2001). The consequences of these emerging diseases are global and profound with increased burden on the public health system, negative impacts on the global economy and food security, declines and extinctions of wildlife species, and subsequent loss of ecosystem integrity. For example, 35 million people are currently living with HIV infection globally (http://www.who.int/gho/hiv/en); 400 million poultry have been culled since 2003 as a result of efforts to control highly pathogenic H5N1 avian influenza (http://www.fao.org/avianflu/en/index.html), and there are increasing biological and ecological consequences.
Examples of health threats to biodiversity include the “spillover” of human diseases to great ape populations (Köndgen et al., 2008), the near-extirpation of
_______________
31 USGS National Wildlife Health Center, Madison, Wisconsin, USA.
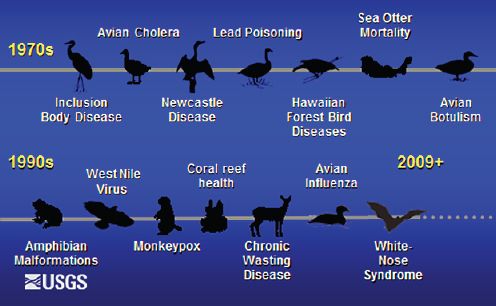
FIGURE A12-1 Emerging diseases investigated by the U.S. Geological Survey.
SOURCE: U.S. Geological Survey.
the black-footed ferret from canine distemper and sylvatic plague (for a review see Abbott et al., 2012), and threats to Hawaiian forest birds from introduced pathogens such as avian malaria and avian pox (van Riper et al., 1986, 2002). There are also newly discovered pathogens or diseases that have resulted in population declines, and global extinctions of several species. Examples include Batrachochytrium dendrobatidis, which causes a cutaneous fungal infection of amphibians and is linked to declines of amphibians globally (Kriger and Hero, 2009); and recently discovered Pseudogymnoascus (Geomyces) destructans, the etiologic agent of white-nose syndrome (WNS), which has caused precipitous declines of North American bat species (Blehert et al., 2009). Furthermore, there is increasing evidence of the subsequent impacts on human and ecosystem health; for example, increasing risk of exposure to Lyme disease as a consequence of decreased biodiversity (LoGiudice et al., 2003) as well as the economic cost of the loss of bats due to decreased insect control services (Boyles et al., 2011). Figure A12-1 is a timeline of important diseases investigated by the U.S. Geological Survey since the 1970s, which illustrates three factors:
- The unprecedented emergence of new pathogens and geographic spread of known pathogens since the 1990s;
- Diseases are increasingly causing large-scale, negative impacts on wildlife populations and spreading over larger geographic areas rather than remaining localized; and
- Diseases are increasingly of concern for multiple sectors, including public health, agriculture and wildlife management agencies.
Of increasing concern are these novel diseases such as WNS as they are hard to anticipate, particularly devastating to human health or wildlife populations, challenging to manage, spread over large geographic areas in short time periods, and may result in ecological ripple effects that are difficult to predict.
The following article provides examples of recently emerged viral diseases of wildlife origin. The examples have been selected to illustrate the drivers of emerging viral diseases, both novel pathogens and previously known diseases, the impacts of these diseases, as well as the role of wildlife both as “villains” or reservoirs as well as “victims” of these viral diseases. The article also discusses potential management strategies for emerging viral diseases in wildlife populations and future science directions in wildlife health to prevent, prepare, respond to, and recover from these disease events. Finally, the concept of One Health and its potential role in developing solutions to these issues of mutual concern is discussed.
Avian Influenza
Wild Birds as Victims?
Extensive phylogenetic analysis of avian influenza viruses has shown that wild birds do not normally harbor highly pathogenic lineages; instead waterfowl and shorebirds in particular are the reservoirs of a vast diversity of low pathogenic avian influenza (LPAI) viruses (Rohm et al., 1995; Sakoda et al., 2010). These LPAI viruses, following their introduction into domestic poultry, then evolve into highly pathogenic strains (Monne et al., 2014). The adaptation to respiratory transmission and ability to replicate in extra-intestinal organs in poultry can take as little as weeks to months following lapses in biosecurity in facilities (Berhane et al., 2009).
Highly pathogenic avian influenza (HPAI) outbreaks are rare in wild birds. Sparrows, starlings, and other peridomestic free-living species have occasionally died in association with HPAI outbreaks in poultry, but these are usually few in number and were exposed to the same virus as in nearby outbreaks in poultry (Alexander, 2007). Of the 31 HPAI outbreaks that have occurred since the discovery in 1955 that fowl plague, a devastating disease in poultry, was caused by avian influenza viruses, only two have involved significant number of wild birds (Table A12-1). The first large-scale wild bird HPAI outbreak killed more than 1,300 common terns (Sterna hirundo) in 1961 in South Africa (Becker, 1966).
TABLE A12-1 Highly Pathogenic Avian Influenza Outbreaks in Poultry and Wild Birds Since 1959
| Year | Strain | Location | Species | Number of Animals |
| 1959 | H5N1 | Scotland, UK | Chicken | Unknown |
| 1961 | H5N3 | South Africa | Gulls | 1,300 |
| 1963 | H7N3 | England, UK | Turkey | 29,000 |
| 1966 | H5N9 | Ontario, Canada | Turkey | 8,000 |
| 1975 | H7N7 | Victoria, Australia | Chicken | 58,000 |
| 1979 | H7N7 | Germany | Chicken | 600,000 |
| 1979 | H7N7 | England, UK | Turkey | 9,000 |
| 1983 | H5N2 | Pennsylvania, US | Chicken | >17 million |
| 1983 | H5N8 | Ireland | Ducks | 307,000 |
| 1985 | H7N7 | Victoria, Australia | Chicken | 240,000 |
| 1991 | H5N1 | England, UK | Turkey | 8,000 |
| 1992 | H7N3 | Victoria, Australia | Chicken | 18,000 |
| 1994 | H7N3 | Queensland, Australia | Chicken | 22,000 |
| 1994 | H5N2 | Mexico | Chicken | Unknown |
| 1995 | H7N3 | Pakistan | Chicken | >6 million |
| 1996 | H5N1 | Hong Kong | Chicken | 3 million |
| 1997 | H7N4 | New South Wales, Australia | Chicken | 160,000 |
| 1997 | H5N2 | Italy | Chicken | 8,000 |
| 1999 | H7N1 | Italy | Chicken | 14 million |
| 2003 | H5N1 | Hong Kong, China | Chicken | 1.5 million |
| 2002 | H7N3 | Chile | Chicken | 700,000 |
| 2003 | H7N7 | Netherlands | Chicken | >25 million |
| 2004 | H7N3 | British Columbia, Canada | Chicken | 17 million |
| 2004 | H5N2 | Texas, US | Chicken | 6,600 |
| 2004 | H5N2 | South Africa | Ostrich | 30,000 |
| 2004-Present | H5N1 | Asia, Europe, Africa | Chicken | >400 million |
| 2005 | H5N2 | British Columbia, | Chicken | 16 million |
| Canada | ||||
| 2007 | H7N3 | Saskatchewan, Canada | Chicken | 53,000 |
| 2008 | H7N7 | England, UK | Chicken | 15,000 |
| 2011 | H5N2 | South Africa | Ostrich | 26,000 |
| 2014 | H5N8 | South Korea | Chicken | 14 million |
It was speculated at the time that unspecified stress in the colony might have “converted a latent into an overt infection” (Becker, 1966). Most of the HPAI outbreaks listed in Table A12-1 were limited in geographical location and were eliminated through prompt management actions and resulted in limited wild bird
exposure. However, the HPAI H5N1 outbreak is exceptional as it had gone largely unchecked from its emergence in 1996 until the events of 2005 (the extraordinary efforts by the government of Hong Kong in 2001 and 2003 are obvious exceptions), resulting in significant spread by human action throughout Southeast Asia (FAO, 2011a). As might be expected from such a large-scale infection in poultry, spill back into wild birds was likely. The HPAI H5N1 outbreaks in wild birds that occurred in 2005–2006, particularly the event in Qinghai, China, where more than 6,000 birds including 3,282 bar-headed geese perished, was the second HPAI outbreak with significant wild bird involvement (Chen et al., 2005; Liu et al., 2005).
Wild Birds as Vectors?
The spectacular feats of long-distance migration that some species of birds undertake makes the concept of long distance disease spread easy to assume. For example, because the strain of West Nile virus (WNV) isolated in New York in 1999 is most closely related to WNV that caused an outbreak in geese and storks in Israel 1 year earlier, Rappole et al. (2000) proposed 31 species of birds that might act as vectors between the Old and New World (Rappole et al., 2000). Other mechanisms of introduction such as the importation of a WNV-infected bird, mosquitoes, or even an infected person are also possible and the exact mechanism of how WNV arrived in North America remains a subject of debate (Roehrig, 2013).
The spread of WNV following its introduction into the Americas in 1999 may serve as an informative example of the spread of a newly introduced disease agent across the landscape. Since WNV has a wide host range (infection has been documented in more than 300 species of birds) and it is a mosquito-transmitted arbovirus infecting at least 10 genera of mosquitoes, the virus was predicted to spread rapidly from its initial introduced location in New York. However, the virus was restricted to the Northeast United States and the Atlantic Flyway for 3 years despite multiple cycles of annual migration, expanding only to the Gulf states in 2001. That year, the first evidence of WNV infection in birds of the Mississippi Flyway was reported, and in the following year, the virus was reported in the rest of the continental United States. However, it was not until 2005, 6 years after its introduction, that WNV was detected in all 48 contiguous states (Gubler, 2007).
Large-scale surveillance programs have detected infections of HPAI H5N1 in very few healthy wild birds, but wild birds are likely to have introduced Clade 2.3.2 HPAI H5N1 viruses into Russia and Mongolia in 2009 and Romania in 2010 (FAO, 2011b). The outbreaks of HPAI H5N1 in Japan in 2010/2011 were preceded by the detection of the virus in wild bird fecal samples collected from Lake Onuma, Hokkaido, in October 2010 (Kajihara et al., 2011; Sakoda et al., 2012). A similar situation occurred in South Korea over the same period, with initial detection in healthy wild birds, followed by outbreaks in poultry farms (Kim et al., 2012). In both Korea and Japan, the virus was assumed to have been
introduced by migratory birds, and following introduction into poultry, the subsequent spread from farm to farm was due to agricultural practices. Together, Japan and South Korea culled 7.3 million birds during eradication efforts (Korea, 2011; Tsukamoto, 2012). Conversely, the persistence of viruses belonging to Clade 1 within Vietnam while having been extirpated elsewhere, and the continued circulation of Clade 2.1 viruses in Indonesia since 2004 and Clade 2.2.1 virus in Egypt since 2006 without any evidence of long-distance export of these clades to adjacent or distant countries, suggests that long-distance transportation of HPAI H5N1 by migratory birds is not usually a significant mechanism of transmission (Scotch et al., 2013; WHO, 2011).
Phylogenetics
The genetic sequence of a virus can be used in phylogenetic studies and when appropriately applied can be used to infer the evolutionary relationship and possible routes of introduction and transmission. For example, HPAI H5N1 outbreaks have occurred in South Korea and Japan in 2004, 2007, and 2010. During each outbreak, genetic information of the viruses provided information independent of the traditional epidemiological investigations to support determinations of the mechanism and pattern of spread. Moreover, the comparison of nucleotide sequence differences showed that the viruses belonged to different clades in different years (2004, Clade 2.5; 2007, Clade 2.2; 2010, Clade 2.3.2.1) and supported the countries’ assertion that the stamping-out policies were effective in the eradication of the virus each time (Kim et al., 2012; Sakoda et al., 2012).
Genetic information can also be used to infer the possible origin of an outbreak strain. The persistence of HPAI H5N1 in China since 1996 has allowed it to re-assort with other avian influenza viruses circulating in poultry (Chen et al., 2004). The HPAI H5N8 virus that was first detected in South Korea in January 2014 is characterized by a hemagglutinin from the HPAI H5N1 Clade 2.3.4.6 lineage, but the other genes are from a variety of viruses including other H5N8, H5N2, and H11N9 viruses (Lee et al., 2014). These types of re-assortments might provide the progeny virus with improved fitness such as increased ability to replicate in domestic poultry or amantadine resistance. Similarly, the nucleotide sequence of the strains of Bluetongue virus found in the European Union (EU) has been used in phylogenetic comparisons with those found elsewhere to infer possible sources of introductions (Purse et al., 2005).
A Direct Pipeline?
Do existing viruses always have to mutate or undergo adaptation before they are able to infect new hosts? Recent examples of avian influenza A viruses suggest that some contemporary viruses already possess the ability to infect mammals efficiently. An avian influenza H3N8 virus was shown to be the causative
agent in a large-scale mortality event of harbor seals (Phoca vitulina) in the northeastern United States in 2011 (Anthony et al., 2012). The virus was most closely related to contemporary wild bird H3N8 viruses in North America and did not contain RNA segments from mammalian influenza viruses, including canine and equine H3N8 lineages. In a second example, an H7N9 virus was found in an 87-year-old man from Shanghai, China, who had died in March 2013 of pneumonia. Since then, more than 450 cases of additional H7N9 infection have been reported with an estimated 32 percent fatality rate. In spite of the serious disease in people, these H7N9 viruses do not cause mortality in experimentally infected chickens and have been found in apparently healthy poultry in live bird markets (Morens et al., 2013). In both the seal H3N8 and poultry H7N9 viruses, mutations that confer increased ability to replicate in mammalian hosts were already present in the avian viruses, suggesting these viruses have a preexisting ability to cross species barriers.
In summary, the literature reveals a complex picture of the role of migratory birds in avian influenza epidemiology, and the view that migratory birds are primarily responsible for highly pathogenic avian influenza outbreaks or drive viral diversity is too simplistic. However, knowing that influenza viruses will move from wild to domestic birds and to people, focusing on interventions such as farm biosecurity will remain key to reducing risks.
There is also a need for ongoing surveillance of genetic diversity. A recent paper showed that gene segments from the 1918 Spanish flu virus circulate today in wild birds and that an artificial construct that brings together modern descendants of the 1918 virus is capable of elevated pathogenicity in mice as well as being able to be spread by respiratory droplets (Watanabe et al., 2014). As the authors of the paper note, a better understanding of the genetic diversity and the molecular mechanisms of pathogenicity will aid in improved risk assessment and outbreak preparedness.
Global Trade: Schmallenberg Virus
Schmallenberg virus (SBV) is a bunyavirus and a member of the Simbu serogroup (OIE, 2013). Its introduction in Europe serves as an example of the expansion of a virus into an area where susceptible hosts and suitable vectors already exist. Schmallenberg virus was first detected in Europe in November 2011 in Schmallenberg, Germany, and 1 month later in The Netherlands (Tarlinton et al., 2012). Since then, 16 countries in Europe have been affected. Infection with SBV is associated with deformities in lambs and calves, abortions, and decreased milk production in cattle, goats, and sheep. The virus is suspected to have been first introduced into Europe in early 2011 because there is no evidence of SBV in archived samples prior to 2011 and its effects only became apparent during the 2011 fall lambing season (Beer et al., 2013). While the actual route of introduction has not been established, the virus may have been introduced by midges that
arrived in shipments of cut flowers, or produce arriving daily from Africa into the clusters of international airports including Amsterdam, Brussels, and Cologne as well as seaports such as Rotterdam (Beer et al., 2013). These are also areas with a high density of susceptible hosts including cattle and sheep as well as established populations of Culicoides species that are competent vectors of SBV.
Climate Change: Bluetongue Virus
Bluetongue virus (BTV) is an orbivirus belonging to the family Reoviridae and is transmitted by Culicoides spp. midges. Bluetongue virus had historically been restricted to a zone between approximately 40°N and 35°S, and its expansion into Europe is an example of spread as a result of climate change leading to conditions permissive to sustained transmission. Since 1998, the geographical range of BTV has expanded, sometimes as far as 700 km northward. Between 1998 and 2005, BTV belonging to five serogroups (BTV-1, BTV-2, BTV-4, BTV-9, and BTV-16) was present continuously in the Mediterranean basin (Saegerman et al., 2008). In 2006, BTV-8 was detected for the first time in Europe, initially in The Netherlands and by 2007, more than 60,000 farms in six countries were affected (Wilson and Mellor, 2009). At least six countries (Bulgaria, France, Italy, Macedonia, Tunisia, and Yugoslavia) had never had BTV previously. Prior to 2004, BTV was associated with periodic expansion and contraction from endemic into adjacent areas as conditions favorable for transmission such as vector availability alternated (Walton, 2004), but the widespread outbreak of BTV-8 in 2006 suggests that factors, including climate change, allowing for sustained transmission, including successful overwintering, may now be present in the EU (Purse et al., 2005; Saegerman et al., 2008).
Diseases at the Human–Primate Interface
The hunting, butchering, and consumption of primates is recognized as a major source of viral disease emergence. This has resulted in cross-species transmission of several retroviruses to humans including simian immunodeficiency virus (SIV), simian T-lymphotropic virus (STLV), and simian foamy virus, the former of which resulted in a human disease of pandemic proportions in the form of human immunodeficiency virus and AIDS (Gao et al., 1999). The extent of the wildlife trade is difficult to measure due to the clandestine nature of the business; however, bushmeat continues to present threats to public health. In a recent study, samples were collected at several international airports from illegally imported nonhuman primate and rodent species, including baboon, chimpanzee, mangabey, guenon, green monkey, cane rat, and rat (Smith et al., 2012). Pathogen screening identified retroviruses (simian foamy virus) and/or herpes viruses (cytomegalovirus and lymphocryptovirus) in the primate samples. These results demonstrated that illegal bushmeat importation into the United States could act as a conduit
for pathogen spread, and the authors suggested that implementation of disease surveillance of the wildlife trade would help facilitate prevention of disease emergence. The uncontrolled extraction and trade of wildlife is also a threat to the persistence of fish and wildlife species, and changing attitudes toward this trade would not only benefit public health, but would assist in the conservation of threatened and endangered species.
Recent outbreaks of zoonotic diseases in African great apes also illustrate the potential role of infectious diseases in jeopardizing the persistence of great ape populations. Controlled contact as well as unavoidable contact between humans and great apes is increasing due to ecotourism, expanding human populations, as well as other ecologic factors such as deforestation and the bushmeat trade (Figure A12-2) (Adams et al., 2001; Guerrera et al., 2003; Walsh et al., 2003). Great apes are especially vulnerable to human diseases due to the close taxonomic relationship, and there are increasing reports of human-associated diseases in great ape populations including outbreaks of sarcoptic mange in mountain gorillas (Gorilla beringei beringei) (Graczyk et al., 2001; Kalema-Zikusoka et al., 2002) suspected to be of human origin. More recently, Köndgen et al. (2008) presented evidence of human paramyxovirus transmission from humans to wild chimpanzees (Pan troglodytes) that resulted in respiratory disease, mortality, and decline of the chimpanzee population. In these areas where humans and primates coexist, improvements in public health infrastructure and measures to reduce disease transmission (for example, the creation of open defecation free zones) would benefit the human populations that have high burdens of disease and unmet health needs as well as these endangered species. This, again, illustrates

the connectivity between human and wildlife health, and provides an additional conservation-related argument for the improvement of public health in these developing countries.
Bats and Emerging Viral Diseases
Bats are being increasingly recognized as an important reservoir of zoonotic viruses of different families, including SARS coronavirus, Nipah virus, Hendra virus, and Ebola virus (Smith and Wang, 2013). The question of whether bats have unique biological features making them ideal reservoir hosts has been the subject of recent discussion (Kupferschmidt, 2013). However, these unique features may also increase their susceptibility to infectious diseases. Pseudogymnoascus (Geomyces) destructans, the etiologic agent of white-nose syndrome (WNS), which has caused precipitous declines of North American bat species, infects bats during hibernation. It has been hypothesized that bats are particularly vulnerable to infection during that period of their life cycle due to natural immunosuppression (Blehert et al., 2009).
The triptych of bats as reservoirs of zoonotic viruses, their ecological and economic importance (Boyles et al., 2009), and threats to their persistence creates the question on how humans and bats can coexist. A recent study of 2007–2008 outbreaks of Marburg virus associated with caves in Uganda may provide some answers (Amman et al., 2012). These caves were used by local population for mining, were also tourist attractions, and contained large population of Rousettus aegyptiacus fruit bats, which were implicated as the reservoir of Marburg virus. Between August 2008 and November 2009, 1,622 bats were captured and tested for Marburg virus, and the Q-RT-PCR data showed distinct pulses of virus infection in older juvenile bats that temporarily coincided with the peak twice yearly birthing seasons. Retrospective analysis of historical human infections suspected to have been the result of discrete spillover events directly from nature found 83 percent (54/65) of events occurred during these seasonal pulses in virus circulation, perhaps demonstrating periods of increased risk for human infection. These results provide a basis for risk-reduction strategies through temporal separation of human caving activities and bats during the high-risk birthing seasons.
Future Directions in Wildlife Health
Early detection of emerging viral diseases in wildlife is an important component of an overall strategy to prevent, prepare for, and respond to emerging infectious diseases. A primary component is the field epidemiological capacity and network to detect and respond to unusual wildlife mortality events. This includes natural resource agency field biologists and wildlife health professionals as well as epidemiologists trained in wildlife disease outbreak investigation and surveillance. State-of-the-art wildlife diagnostic laboratory capacity, including
virology, microbiology, chemistry, and pathology to detect and identify novel emerging pathogens is essential, including a network of laboratories specializing in wildlife pathogen detection and characterization. The application of new molecular diagnostic technologies such as next-generation sequencing has opened up previously unknown avenues for pathogen discovery (Relman, 1998; Wang et al., 2003) and should become mainstream in this context. Such systems will also contribute to upstream surveillance for hazards, and strengthen the capacity of nations to detect infectious diseases that may represent potential public health emergencies (Baker et al., 2010).
We have established Earth (Landsat: landsat.usgs.gov) and climate monitoring systems (NOAA: www.nesdis.noaa.gov) that provide continuous imagery, atmospheric measurements, and climatic data, and we have global public health surveillance systems for human diseases (WHO: http://www.who.int/research/en), yet we lack the same ongoing, systematic collection of data for fish and wildlife health. Collection and integration of data from such a long-term data system with data from a variety of sources, including human and animal health data, climatic, ecologic, hydrologic, geologic, and socioeconomic data, among other sources, will allow a deeper understanding of the environmental drivers and the generation of predictive models of “hot spots” of disease emergence (Jones et al., 2008). This will ultimately allow for the targeting of resources to geographic areas and populations at greater risk and the prevention of disease emergence and spread. The development of new analytical models will also provide us with the mathematical tools to identify and anticipate threats to wildlife, understand the distribution, dynamics, and impacts of disease, and ultimately provide better information for guiding management decisions. Recognizing that not all diseases will be predicted and prevented, the biggest deficiency is a suite of tools that can be mobilized to manage diseases in wildlife populations. The current methods such as culling are crude, unpopular, and generally ineffective. Vaccines are probably the primary, cost-effective public health and veterinary intervention available and have been used widely to save millions of lives and reduce economic losses. Very few vaccines are available for use in free-ranging wildlife populations due to the challenges of delivery; however, the oral rabies vaccine has reduced the prevalence of rabies infection in wildlife, and was used to successfully eradicate fox rabies from Western Europe (Brochier et al., 1991). Further research in the development of safe and effective vaccines that can be mass delivered to wildlife populations, as is being done to develop a sylvatic plague vaccine for prairie dogs (Abbott et al., 2012), would allow for this technique to be increasingly applied to vaccinate upstream and prevent pathogen spillover. Increased focus on other disease management tools should include biocontrol strategies and research on social attitudes and behaviors related to natural resources and disease management.
Finally, we need robust partnerships to address these pressing issues of mutual concern. While the One Health concept recognizes the interconnectedness of human, animal, wildlife, and ecosystem health, the infrastructure to respond
TABLE A12-2 Factors That Contributed to the Success of One Health Projects
| • | Sense of urgency and common purpose |
| • | Delegated authority or mandate to conduct the work |
| • | Good governance: An interagency steering committee or working group is formed to oversee the work |
| • | Foundation of trust exists among key individuals in different agencies, built on a willingness to acknowledge the other agencies’ concerns |
| • | Mutually agreed-upon outcomes are science based |
| • | Clearly defined roles and responsibilities |
| • | Leadership rotates rather than being monopolized by one sector |
SOURCE: Rubin et al., 2014.
to wildlife emerging diseases and wildlife health emergencies is lacking. Until we have the operational framework (the network of partners, with appropriate governance, policies, procedures, etc.) by which agencies and institutions with a stake in wildlife diseases cooperate and collaborate to achieve optimal outcomes for human, animal, and ecosystem health, the third leg of the One Health stool (the three legs being human health, domestic animal health, and wildlife health) will always be missing. In one sense this is a leadership challenge. Interdisciplinary teams are more likely to be successful when there is a unified task and shared goals and values, and when personal relationships are developed from a foundation of trust and respect (Anholt et al., 2012). A recent review of successful One Health projects revealed common factors that contributed to their success (Rubin et al. 2014; Table A12-2). Consequently, what are the common core values of One Health, and do we have the individual leadership skills, such as an ability to think beyond the boundaries of one’s own agency or institution to make One Health successful? Addressing emerging viral diseases is a shared leadership responsibility we all must willingly accept, and doing so will help us make significant progress.
References
Abbott, R. C., J. E. Osorio, C. M. Bunck, and T. E. Rocke. 2012. Sylvatic plague vaccine: A new tool for conservation of threatened and endangered species? EcoHealth 9(3):243-250.
Adams, H., J. Sleeman, I. Rwego, I., and J. New. 2001. Self-reported medical history survey of humans as a measure of health risk to the chimpanzees (Pan troglodytes schweinfurthii) of Kibale National Park, Uganda. Oryx 35:308-312.
Alexander, D. J. 2007. An overview of the epidemiology of avian influenza. Vaccine 25(30):5637-5644. Amman, B. R., S. A. Carroll, Z. D. Reed, T. K. Sealy, S. Balinandi, R. Swanepoel, A. Kemp, B. R. Erickson, J. A. Comer, S. Campbell, D. L. Cannon, M. L. Khristova, P. Atimnedi, C. D. Paddock, R. J. Crockett, T. D. Flietstra, K. L. Warfield, R. Unfer, E. Katongole-Mbidde, R. Downing, J. W. Tappero, S. R. Zaki, P. E. Rollin, T. G. Ksiazek, S. T. Nichol, and J. S. Towner. 2012. Seasonal pulses of Marburg virus circulation in juvenile Rousettus aegyptiacus bats coincide with periods of increased risk of human infection. PLoS Pathogens 8(10):e1002877.
Anholt, R., C. Stephen, and R. Copes. 2012. Strategies for collaboration in the interdisciplinary field of emerging zoonotic diseases. Zoonoses and Public Health 59(4):229-240.
Anthony, S. J., J. A. St Leger, K. Pugliares, H. S. Ip, J. M. Chan, Z. W. Carpenter, I. Navarrete-Macias, M. Sanchez-Leon, J. T. Saliki, J. Pedersen, W. Karesh, P. Daszak, R. Rabadan, T. Rowles, and W. I. Lipkin. 2012. Emergence of fatal avian influenza in New England harbor seals. mBio 3(4):e00166-12.
Baker, M. G., S. Easther, and N. Wilson. 2010. A surveillance sector review applied to infectious diseases at a country level. BMC Public Health 10:332.
Becker, W. B. 1966. The isolation and classification of Tern virus: Influenza A—Tern South Africa—1961. Journal of Hygiene 64(3):309-320.
Beer, M., F. J. Conraths, and W. H. van der Poel. 2013. “Schmallenberg virus”—a novel orthobunyavirus emerging in Europe. Epidemiology and Infection 141(1):1-8.
Berhane, Y., T. Hisanaga, H. Kehler, J. Neufeld, L. Manning, C. Argue, K. Handel, K. HooperMcGrevy, M. Jonas, J. Robinson, R. G. Webster, and J. Pasick. 2009. Highly pathogenic avian influenza virus A (H7N3) in domestic poultry, Saskatchewan, Canada, 2007. Emerging Infectious Diseases 15(9):1492-1495.
Blehert, D. S., A. C. Hicks, M. Behr, C. U. Meteyer, B. M. Berlowski-Zier, E. L. Buckles, J. T. Coleman, S. R. Darling, A. Gargas, and R. Niver. 2009. Bat white-nose syndrome: An emerging fungal pathogen? Science 323(5911):227.
Boyles, J. G., P. M. Cryan, G. F. McCracken, and T. H. Kunz. 2011. Conservation. Economic importance of bats in agriculture. Science 332(6025):41-42.
Brochier, B., M. P. Kieny, F. Costy, P. Coppens, B. Bauduin, J. P. Lecocq, B. Languet, G. Chappuis, P. Desmettre, K. Afiademanyo, et al. 1991. Large-scale eradication of rabies using recombinant vaccinia-rabies vaccine. Nature 354(6354):520-522.
Chen, H., G. Deng, Z. Li, G. Tian, Y. Li, P. Jiao, L. Zhang, Z. Liu, R. G. Webster, and K. Yu. 2004. The evolution of H5N1 influenza viruses in ducks in southern China. Proceedings of the National Academy of Sciences of the United States of America 101(28):10452-10457.
Chen, H., G. J. Smith, S. Y. Zhang, K. Qin, J. Wang, K. S. Li, R. G. Webster, J. S. Peiris, and Y. Guan. 2005. H5N1 virus outbreak in migratory waterfowl. Nature 436(7048):191-192.
Daszak, P., A. Cunningham, and A. Hyatt. 2001. Anthropogenic environmental change and the emergence of infectious diseases in wildlife. Acta Tropica 78(2):103-116.
FAO (Food and Agriculture Organization of the United Nations). 2011a. Approaches to controlling, preventing and eliminating H5N1 highly pathogenic avian influenza in endemic countries, Animal Production and Health Paper. No. 171. Rome, Italy: Food and Agriculture Organization of the United Nations.
FAO. 2011b. Approaches to controlling, preventing and eliminating H5N1 highly pathogenic avian influenza in endemic countries, FAO animal production and health paper. Rome, Italy: Food and Agriculture Organization of the United Nations.
Gao, F., E. Bailes, D. L. Robertson, Y. Chen, C. M. Rodenburg, S. F. Michael, L. B. Cummins, L. O. Arthur, M. Peeters, and G. M. Shaw. 1999. Origin of HIV-1 in the chimpanzee Pan troglodytes troglodytes. Nature 397(6718):436-441.
Graczyk, T. K., A. B. Mudakikwa, M. R. Cranfield, and U. Eilenberger. 2001. Hyperkeratotic mange caused by Sarcoptes scabiei (Acariformes: Sarcoptidae) in juvenile human-habituated mountain gorillas (Gorilla gorilla beringei). Parasitology Research 87:1024-1028.
Gubler, D. J. 2007. The continuing spread of West Nile virus in the Western Hemisphere. Clinical Infectious Disease 45(8):1039-1046.
Guerrera, W., J. M. Sleeman, S. B. Jasper, L. B. Pace, T. Y. Ichinose, and J. S. Reif. 2003. Medical survey of the local human population to determine possible health risks to the mountain gorillas of Bwindi Impenetrable Forest National Park, Uganda. International Journal of Primatology 24:197-207.
Jones, K. E., N. G. Patel, M. A. Levy, A. Storeygard, D. Balk, J. L. Gittleman, and P. Daszak. 2008. Global trends in emerging infectious diseases. Nature 451(7181):990-993.
Kajihara, M., K. Matsuno, E. Simulundu, M. Muramatsu, O. Noyori, R. Manzoor, E. Nakayama, M. Igarashi, D. Tomabechi, R. Yoshida, M. Okamatsu, Y. Sakoda, K. Ito, H. Kida, and A. Takada. 2011. An H5N1 highly pathogenic avian influenza virus that invaded Japan through waterfowl migration. Japanese Journal of Veterinary Research 59(2-3):89-100.
Kalema-Zikusoka, G., R. Kock, and E. Macfie. 2002. Scabies in free-ranging mountain gorillas (Gorilla beringei beringei) in Bwindi Impenetrable National Park, Uganda. Veterinary Record 150:12-15.
Kim, H. R., Y. J. Lee, C. K. Park, J. K. Oem, O. S. Lee, H. M. Kang, J. G. Choi, and Y. C. Bae. 2012. Highly pathogenic avian influenza (H5N1) outbreaks in wild birds and poultry, South Korea. Emerging Infectious Diseases 18(3):480-483.
Köndgen, S., H. Kühl, P. K. N’Goran, P. D. Walsh, S. Schenk, N. Ernst, R. Biek, P. Formenty, K. Mätz-Rensing, and B. Schweiger. 2008. Pandemic human viruses cause decline of endangered great apes. Current Biology 18(4):260-264.
Korea, S. 2011. Highly pathogenic avian influenza, Korea (Rep. of) (Follow-up Report 15: 08/09/2011). http://www.oie.int/wahis_2/temp/reports/en_fup_0000010982_20110908_181911.pdf (accessed June 7, 2014).
Kriger, K. M., and J.-M. Hero. 2009. Chytridiomycosis, amphibian extinctions, and lessons for the prevention of future panzootics. EcoHealth 6(1):6-10.
Kupferschmidt, K. 2013. Link to MERS virus underscores bats’ puzzling threat. Science 341:948-949. Lee, Y. J., H. M. Kang, E. K. Lee, B. M. Song, J. Jeong, Y. K. Kwon, H. R. Kim, K. J. Lee, M. S. Hong, I. Jang, K. S. Choi, J. Y. Kim, H. J. Lee, M. S. Kang, O. M. Jeong, J. H. Baek, Y. S. Joo, Y. H. Park, and H. S. Lee. 2014. Novel reassortant influenza A(H5N8) viruses, South Korea, 2014. Emerging Infectious Diseases 20(6):1086-1089.
Liu, J., H. Xiao, F. Lei, Q. Zhu, K. Qin, X. W. Zhang, X. L. Zhang, D. Zhao, G. Wang, Y. Feng, J. Ma, W. Liu, J. Wang, and G. F. Gao. 2005. Highly pathogenic H5N1 influenza virus infection in migratory birds. Science 309(5738):1206.
LoGiudice, K., R. S. Ostfeld, K. A. Schmidt, and F. Keesing. 2003. The ecology of infectious disease: Effects of host diversity and community composition on Lyme disease risk. Proceedings of the National Academy of Sciences of the United States of America 100(2):567-571.
Monne, I., A. Fusaro, M. I. Nelson, L. Bonfanti, P. Mulatti, J. Hughes, P. R. Murcia, A. Schivo, V. Valastro, A. Moreno, E. C. Holmes, and G. Cattoli. 2014. Emergence of a highly pathogenic avian influenza virus from a low-pathogenic progenitor. Journal of Virology 88(8):4375-4388.
Morens, D. M., J. K. Taubenberger, and A. S. Fauci. 2013. H7N9 avian influenza A virus and the perpetual challenge of potential human pandemicity. mBio 4(4):e00445-13.
OIE (World Organization for Animal Health). 2013. OIE technical factsheet: Schmallenberg virus. http://www.oie.int/fileadmin/Home/eng/Our_scientific_expertise/docs/pdf/A_Schmallenberg_virus.pdf (accessed July 7, 2014).
Purse, B. V., P. S. Mellor, D. J. Rogers, A. R. Samuel, P. P. Mertens, and M. Baylis. 2005. Climate change and the recent emergence of bluetongue in Europe. Nature Reviews: Microbiology 3(2):171-181.
Rappole, J. H., S. R. Derrickson, and Z. Hubalek. 2000. Migratory birds and spread of West Nile virus in the Western Hemisphere. Emerging Infectious Diseases 6(4):319-328.
Relman, D. A. 1998. Detection and identification of previously unrecognized microbial pathogens. Emerging Infectious Diseases 4(3):382.
Roehrig, J. T. 2013. West Nile virus in the United States—a historical perspective. Viruses 5(12):3088-3108.
Rohm, C., T. Horimoto, Y. Kawaoka, J. Suss, and R. G. Webster. 1995. Do hemagglutinin genes of highly pathogenic avian influenza viruses constitute unique phylogenetic lineages? Virology 209(2):664-670.
Rubin, C., B. Dunham, and J. Sleeman. 2014. Making One Health a reality: Crossing bureaucratic boundaries. Microbiology Spectrum 2(1):OH-0016-2012.
Saegerman, C., D. Berkvens, and P. S. Mellor. 2008. Bluetongue epidemiology in the European Union. Emerging Infectious Diseases 14(4):539-544.
Sakoda, Y., S. Sugar, D. Batchluun, T. O. Erdene-Ochir, M. Okamatsu, N. Isoda, K. Soda, H. Takakuwa, Y. Tsuda, N. Yamamoto, N. Kishida, K. Matsuno, E. Nakayama, M. Kajihara, A. Yokoyama, A. Takada, R. Sodnomdarjaa, and H. Kida. 2010. Characterization of H5N1 highly pathogenic avian influenza virus strains isolated from migratory waterfowl in Mongolia on the way back from the southern Asia to their northern territory. Virology 406(1):88-94.
Sakoda, Y., H. Ito, Y. Uchida, M. Okamatsu, N. Yamamoto, K. Soda, N. Nomura, S. Kuribayashi, S. Shichinohe, Y. Sunden, T. Umemura, T. Usui, H. Ozaki, T. Yamaguchi, T. Murase, T. Ito, T. Saito, A. Takada, and H. Kida. 2012. Reintroduction of H5N1 highly pathogenic avian influenza virus by migratory water birds, causing poultry outbreaks in the 2010-2011 winter season in Japan. Journal of General Virology 93(Pt 3):541-550.
Scotch, M., C. Mei, Y. J. Makonnen, J. Pinto, A. Ali, S. Vegso, M. Kane, I. N. Sarkar, and P. Rabinowitz. 2013. Phylogeography of influenza A H5N1 clade 2.2.1.1 in Egypt. BMC Genomics 14:871.
Smith, I., and L. F. Wang. 2013. Bats and their virome: An important source of emerging viruses capable of infecting humans. Current Opinion in Virology 3(1):84-91.
Smith, K. M., S. J. Anthony, W. M. Switzer, J. H. Epstein, T. Seimon, et al. 2012. Zoonotic viruses associated with illegally imported wildlife products. PLoS One 7(1):e29505. doi:10.1371/journal.pone.0029505.
Spellberg, B. 2008. Dr. William H. Stewart: Mistaken or maligned? Clinical Infectious Diseases 47:294.
Tarlinton, R., J. Daly, S. Dunham, and J. Kydd. 2012. The challenge of Schmallenberg virus emergence in Europe. Veterinary Journal 194(1):10-18.
Taylor, L. H., S. M. Latham, and E. Mark. 2001. Risk factors for human disease emergence. Philosophical Transactions of the Royal Society of London. Series B: Biological Sciences 356(1411): 983-989.
Tsukamoto, K. 2012. Highly pathogenic avian influenza in Japan: Outbreaks, control measures, and roles of wild birds. Journal of Disaster Research 7(3):324-331.
van Riper III, C., S. G. van Riper, M. L. Goff, and M. Laird. 1986. The epizootiology and ecological significance of malaria in Hawaiian land birds. Ecological Monographs 56(4):327-344.
van Riper III, C., S. G. van Riper, W. R. Hansen, and S. Hackett. 2002. Epizootiology and effect of avian pox on Hawaiian forest birds. The Auk 119(4):929-942.
Walsh, P. D., K. A. Abernethy, M. Bermejo, R. Beyers, P. De Wachter, M. E. Akou, B. Huijbregts, D. I. Mambounga, A. K. Toham, A. M. Kilbourn, S. A. Lahm, S. Latour, F. Maisels, C. Mbina, Y. Mihindou, S. Ndong Obiang, E. N. Effa, M. P. Starkey, P. Telfer, M. Thibault, C. E. G. Tutin, L. J. T. White, and D. S. Wilkie. 2003. Catastrophic ape decline in western equatorial Africa. Nature 422(6932):611-614.
Walton, T. E. 2004. The history of bluetongue and a current global overview. Veterinaria Italiana 40(3):31-38.
Wang, D., A. Urisman, Y.-T. Liu, M. Springer, T. G. Ksiazek, D. D. Erdman, E. R. Mardis, M. Hickenbotham, V. Magrini, and J. Eldred. 2003. Viral discovery and sequence recovery using DNA microarrays. PLoS Biology 1(2):e2.
Watanabe, T., G. Zhong, C. A. Russell, N. Nakajima, M. Hatta, A. Hanson, R. McBride, D. F. Burke, K. Takahashi, S. Fukuyama, Y. Tomita, E. A. Maher, S. Watanabe, M. Imai, G. Neumann, H. Hasegawa, J. C. Paulson, D. J. Smith, and Y. Kawaoka. 2014. Circulating avian influenza viruses closely related to the 1918 virus have pandemic potential. Cell Host Microbe 15(6):692-705.
WHO (World Health Organization). 2011. FAO-OIE-WHO Technical Update: Current evolution of avian influenza H5N1 viruses. http://www.who.int/entity/influenza/human_animal_interface/tripartite_notes_H5N1.pdf?ua=1 (accessed June 6, 2014).
Wilson, A. J., and P. S. Mellor. 2009. Bluetongue in Europe: Past, present and future. Philosophical Transactions of the Royal Society of London Series B: Biological Sciences 364(1530):2669-2681.















