3
Opportunities and Challenges
for Big Data and Analytics
After the introductory remarks, six panelists discussed some of the ways in which researchers envision using big data and the associated analytic tools to track infectious diseases and also discussed some of the obstacles that need to be addressed before that promise becomes reality. Guillaume Chabot-Couture, an associate principal investigator at the Institute for Disease Modeling at Intellectual Ventures, described the defining characteristics of big data and showed how big data–informed mapping techniques have the potential to predict disease outbreaks. Jonna Mazet, the executive director of the One Health Institute at the University of California, Davis, discussed how the PREDICT team has been using big data to address the challenge of preempting disease outbreaks that result from the transmission of infectious agents from animals into humans. Simon Hay, a professor of global health and the director of geospatial science at the University of Washington, outlined a process for using big data to map the global distribution of infectious diseases. Michael Edelstein, a research fellow at the Chatham House Centre on Global Health Security, spoke about the potential for digital disease detection, also known as digital epidemiology, to augment traditional “shoe leather” epidemiology and increase the speed at which disease outbreaks are spotted. Catherine Ordun, a deputy project manager for data science and health surveillance at Booz Allen Hamilton, provided an overview of the typical information technology architecture used to compile, organize, and analyze big datasets. Colonel Emil Lesho, an activity duty physician with the U.S. Army and the director of the U.S. Department of Defense’s (DoD’s) Multidrug-Resistant Organism Repository and Surveillance Network and Antimicrobial Resistance Monitoring and Research Program, provided examples of how big data can be used to identify and combat microbial drug resistance.
DEFINING AND USING BIG DATA
As noted by each of the first four speakers, big data holds tremendous promise for infectious diseases research, surveillance, and prevention, but that begs the question, What exactly is big data? Chabot-Couture explained that the “four Vs”—volume, velocity, variety, and veracity—are often used to determine whether a dataset is big data or not. Volume refers to the amount of data, whether it is a terabyte, petabyte, or exabyte, while velocity reflects the speed at which live data is coming in for analysis. The two less commonly noticed attributes of big data are variety—the different data formats that are not necessarily easy to combine or the different types of data coming in for simultaneous analysis—and veracity, a reflection of the imperfectness, incompleteness, or unreliability of the data.
For Chabot-Couture, the goal of using big data is not just to find something interesting for the sake of discovery but to find something interesting that is actionable at scale. One approach to taking advantage of the variety of data available in big datasets, he said, is to explore the data using data mining and feature engineering techniques and analyze the data using methods outside of the typical operational analyses to look at questions outside of the normal day-to-day use of the data. Big data may also improve the predictive power of existing tools that may not be performing as well as desired, he added. The variety of data in multiple big datasets might, for example, enable the refinement of multiple predictors as a means of optimizing forecasting models.
As with any data, big datasets are only as useful as they are reliable, and there are a number of ways to test the veracity of a dataset. One approach, Chabot-Couture said, is to compare two different sets of data that use different ways of measuring the same underlying quantities. Another approach uses internal consistency metrics. “If you expect your dataset to behave a certain way, and it does not, that’s an indication that something is going wrong,” Chabot-Couture said. In some instances, there may an accepted gold standard that can be used as a benchmark against which to compare a big dataset.
Combining Big Data with Shoe Leather Epidemiology
While the reliability of big datasets is an obvious concern when it comes to managing global health risks, big datasets are not meant to be used in isolation. “That’s certainly something that I believe,” said Scott Dowell, the deputy director for surveillance in epidemiology at the Bill & Melinda Gates Foundation and the moderator of one of the two morning panel sessions that explored some of the opportunities and challenges for big data and analytics. Instead, Dowell said, big data can be most useful when combined with old-fashioned “shoe leather” epidemiology—researchers walking door-to-door collecting primary data to understand the spread of an outbreak. In the end, he said, big data and analytics should be complementary with shoe leather epidemiology, and these two sources of information can enhance each other when used thoughtfully.
SHIFTING TO A PREVENTION PARADIGM FOR EMERGING INFECTIOUS DISEASES
Some 7 years ago, the U.S. Agency for International Development (USAID) issued a challenge to preempt or combat at their source the first stage of emergence of zoonotic diseases—those originating in animals—that pose a significant threat to public and animal health and create and have the potential to produce pandemic infections. As Mazet, put it, “This was a pretty crazy challenge because it was attempting to stop everything we do not know might happen before it happens, and if we are successful, no one will know about it.” Nonetheless, Mazet and her colleagues at the One Health Institute, with support from USAID, put together a consortium, now known as PREDICT, of ministries of health, agriculture, and environment, multiple universities, and nongovernmental organizations in 31 countries to tackle this problem using big data as a critical tool.
The PREDICT Strategy
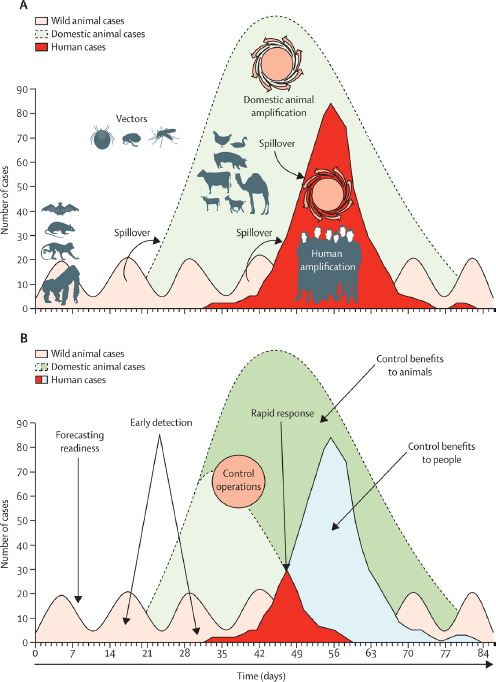
According to the USAID Emerging Pandemic Threats Program, nearly 75 percent of all new, emerging, or reemerging diseases affecting humans at the beginning of the 21st century are zoonotic (USAID, 2016), and Mazet noted that between 1990 and 2010, 91 percent of all zoonotic disease spillover events were from wildlife (Karesh et al., 2012) (see Figure 3-1A). The challenge, then, Mazet said, is to identify those potential human pathogens circulating in wildlife and enact control operations before they have a chance to cause widespread disease in domestic animals and humans (see Figure 3-1B).
The first way in which big data helped the PREDICT team was to inform its modeling efforts aimed at forming a surveillance program that would target species, locations, and interfaces—the contact points between humans and animals that allow pathogen spillovers to occur—and creating a diagnostic platform capable of spotting both known and novel pathogens in the least developed countries. The team used an iterative approach of “Get the data, process the data, get it back to the modeling and analytics team to improve the models so that we can continue to target where to work” Mazet explained. They also used a network analysis to evaluate the transmission interfaces of every zoonotic virus in the past 20 years (see Figure 3-2). Many of the resulting models produced what she called commonsense results in that they predicted that the viruses most likely to spill over would be those that have done so in the past or be closely related to those that have done so in the past. The models also predicted that animals serving as food sources, both domesticated and hunted, and species that have adapted well to living with humans, such as rodents and bats, would be the most likely hosts.
Modeling using data mined from the literature only went so far, though, because the goal of this effort is to identify not just known hazards but new ones, too, Mazet explained. For example, bat guano farming is growing as an
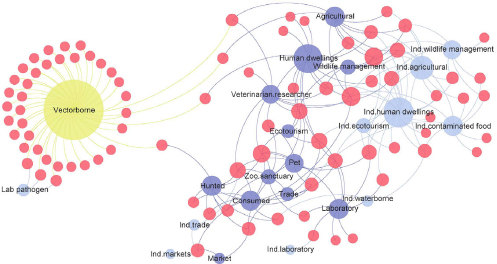
SOURCES: Mazet presentation; Kreuder Johnson et al., 2015.
industry in Southeast Asia, and while there is little information about this activity in the literature, bats are known transmission sources for coronaviruses, and so this becomes a new high-risk interface to be added to a model. In addition to boots-on-the-ground research to identify high-risk interfaces in the target nations, Mazet said, the PREDICT team also used big data to model large-scale ecological drivers, such as regional wildlife biodiversity, human population density, changes in land use, and agricultural industry changes.
5 Years of Achievements
In its first 5 years, the PREDICT team was able to train workers in 20 countries to safely collect samples from live animals, keeping in mind that these samples will be screened from some of the deadly viruses. The team also developed a consensus polymerase chain reaction (cPCR) approach as a diagnostic platform and deployed that platform in the 20 countries (PREDICT Consortium, 2014).
This cPCR platform is capable of detecting 20 different viral families, including all of the viruses that have caused epidemics and pandemics in the past, Mazet explained. Over the initial 5-year period, these trained workers have run some 400,000 diagnostic tests on samples collected from more than 56,000 animals and have identified 984 viruses, 182 of them known and the others newly discovered viruses (see Table 3-1), from 28 viral families (see Table 3-2).
This work remapped the geographic location where different viruses are found and their host species, Mazet said. It also identified novel viruses in certain families, such as the coronavirus and herpes virus families, with the potential to jump from animals to humans. In particular, Mazet said, the study found a number of coronaviruses closely related to the Middle East respiratory syndrome (MERS) virus in regions outside of the Middle East, where MERS was known, and as a result the PREDICT team was able to work with teams in countries such as Mexico and Thailand and to identify closely related viruses in real time. The PREDICT team has now used the data from the first 20 countries to develop viral discovery curves that show how many samples are needed to find every virus infecting each species. “This helped us determine that it is a realistic endeavor to take this to scale beyond these initial 20 countries,” Mazet said.
When asked how to clarify that comment, Mazet said that by her estimates the cost of discovering every virus that could spill over into humans would be more than $1 billion and less than $10 billion. This amount is on the same order of the $6.7 billion that the World Bank projected would be saved annually by preventing zoonotic disease outbreaks and much less than the tens of billions of dollars spent containing the most recent Ebola outbreak in West Africa. An added benefit, aside from the eventual cost savings, would be the opportunity for researchers to develop more effective vaccines and countermeasures for families of viruses that would enable the world to be ahead of these outbreaks rather than always catching up.
TABLE 3-1 Viral Detection Results from PREDICT’s First 5 Years of Operations
| Bats | Nonhuman Primates | Rodents and Shrews | Humans | Other Taxa | Totala | |
|---|---|---|---|---|---|---|
| Novel | 431 | 234 | 143 | 3 | 9 | 820 |
| Known | 80 | 55 | 15 | 31 | 1 | 182 |
| Total | 1,002 |
a Note numbers of viruses do not total to 984 as cited in text because viruses have been found in more than one wildlife host taxa.
SOURCES: Re-created from Mazet presentation; PREDICT Consortium, 2014.
TABLE 3-2 PREDICT Virus Detection Results by Viral Family
| Viral Family | Novel Bat | Known Bat | Novel Primate | Known Primate | Novel Rodent/Shrew | Known Rodent/Shrew | Novel Human | Known Human |
|---|---|---|---|---|---|---|---|---|
| Adenovirus | 53 | 3 | 6 | 4 | 32 | 1 | 1 | 3 |
| Astrovirus | 153 | 33 | 19 | 3 | 31 | 1 | 0 | 1 |
| Coronavirus | 61 | 30 | 3 | 0 | 6 | 0 | 0 | 2 |
| Dependovirus | 0 | 0 | 11 | 0 | 0 | 0 | 0 | 0 |
| Flavivirus | 3 | 0 | 0 | 1 | 0 | 0 | 0 | 2 |
| Hantavirus | 3 | 1 | 0 | 0 | 0 | 2 | 0 | 1 |
| Herpesvirus | 46 | 0 | 48 | 25 | 43 | 6 | 0 | 5 |
| Orbivirus | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 |
| Paramyxovirus | 63 | 7 | 0 | 2 | 11 | 2 | 0 | 3 |
| Polyomavirus | 27 | 1 | 4 | 3 | 8 | 0 | 0 | 1 |
| Arenavirus | 0 | 0 | 0 | 0 | 2 | 2 | 0 | 0 |
| Rhabdovirus | 19 | 0 | 2 | 0 | 7 | 0 | 1 | 0 |
| Seadornavirus | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Bocavirus | 0 | 2 | 1 | 3 | 0 | 0 | 0 | 0 |
| Enterovirus | 0 | 0 | 5 | 4 | 2 | 0 | 0 | 5 |
| Retrovirus | 0 | 0 | 4 | 7 | 0 | 0 | 0 | 1 |
| Alphavirus | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 |
| Poxvirus | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 |
| Influenza | 0 | 2 | 0 | 0 | 0 | 1 | 0 | 5 |
| Mononegavirales | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 |
| Papillomavirus | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 |
| Picobirnavirus | 0 | 0 | 120 | 0 | 0 | 0 | 0 | 0 |
| Picornavirus | 0 | 0 | 4 | 0 | 0 | 0 | 0 | 0 |
| Picornavirales | 0 | 0 | 4 | 0 | 0 | 0 | 0 | 0 |
| Phlebovirus | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Rotavirus | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 |
| Anellovirus | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 |
| Hepadnavirus | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 |
NOTE: Numbers of viruses do not total to 984 as cited in text because viruses have been found in more than one wildlife host taxa.
SOURCES: Re-created from Mazet presentation; PREDICT Consortium, 2014.
Modeling Outbreaks and Saving Lives
Combining the viral scan data with data on human population, wildlife biodiversity, and changes in agricultural practices and land use that together can capture the routes by which viruses can enter the human population, the PREDICT team also developed a model that predicts the relative risk of zoonotic viral outbreaks. The team has also modeled outbreaks in real time and assisted health officials in 10 countries deal with 23 outbreaks over the project’s first 5 years, Mazet said. At the same time as the recent Ebola outbreak in West Africa, the PREDICT team detected the Ebola virus in the Democratic Republic of the Congo, enabling the prime minister’s office to restrict the movement of people, which, Mazet said, resulted in that outbreak ending quickly and minimized mortality there to under 100 people. In contrast, this outbreak lasted much longer and resulted in many more deaths in nearby countries that were not part of the PREDICT program.
Disseminating information is a critical piece of the PREDICT program, Mazet said. Once viruses are identified, that information goes to the ministries of health, agriculture, and environment in the partner countries for coordinated review and interpretation. The information is then released to the public as a HealthMap application on the PREDICT website.1 As an aside, Mazet said, getting the ministries to release these data to the public was much less of a challenge than the team had expected. The HealthMap application enables any user to click on a hot spot and obtain data on every sample collected and every virus identified, including whether a specific virus is considered a threat to human health. Eventually, she added, every virus will be sequenced and the data will be deposited into GenBank.2
Potential Next Steps for the PREDICT Strategy
Having identified so many novel viruses, the PREDICT team is now addressing the challenge of determining which ones are important to human health. Mazet said that one of her frustrations has been that the wealth of existing data has not helped her team rank which viruses are most important for further study. PREDICT’s modeling, epidemiology, and virus teams are now working to develop a first-of-its-kind ranking system, which she acknowledged will be far from perfect but will at least stimulate conversation about how to rank viral threats. The team is also expanding its efforts to detect viruses in human populations and to understand how human behavior, particularly with regard to interactions with livestock and local wildlife, relates to the viruses found in people. “The idea is to target behaviors and find palatable interventions ahead of the next big epidemic or pandemic,” she explained.
___________________
1 HealthMap is an organization that scans and tracks a variety of health sources online. More information is available at http://www.healthmap.org/site/about (accessed October 31, 2016).
2 See http://www.ncbi.nlm.nih.gov/genbank (accessed October 31, 2016).
In closing, Mazet said the world’s increasing population and expansion into previously uninhabited areas will increase potential spillover and the risk of disease. However, what PREDICT has been able to show so far is that it is possible to identify the viruses that exist in specific regions before spillover occurs, rather than chasing them down as an epidemic or pandemic is occurring. As a result, she said, “we can give people the information they need to make good choices and make good response plans.”
POTENTIAL OPPORTUNITIES FOR BIG DATA TO HELP MAP DISEASES
A number of inputs are needed to use geospatial techniques to generate occurrence maps, Hay explained. These inputs include
- occurrence points—the latitude, longitude, and time for every diagnosis of a particular disease;
- environmental covariates—the readily available data on the rainfall, temperature, vegetation, population density, and other associated demographic variables that have been collected continuously across the planet;
- control data—information reflecting areas in which the particular disease is absent; and
- a constraint parameter called the definitive extent—the probability that a disease will be in a particular location.
Using a technique for species-distribution modeling based on boosted regression trees, it is possible to take all of these data and make a continuous map of disease occurrence (Bhatt et al., 2013).
While this approach works, a number of challenges make it difficult. The first challenge Hay discussed is compiling occurrence data from the literature by identifying every paper documenting the occurrence of a particular disease and manually mapping each occurrence. For dengue fever, which did not involve big data, this exercise took Hay and his colleagues 1 year and generated some 8,000 points from more than 100 countries (Messina et al., 2014). He said that, as is the case with the PREDICT project, his group collaborates with HealthMap. Using an automated data retrieval system, the HealthMap team was able to add another 1,500 occurrence points from its digital archives. For dengue, this exercise showed that there is good coverage in the Americas and Asia, but poor coverage in Africa with respect to data collection (see Figure 3-3). Hay said that even in regions that appear well-covered, drilling into the data reveals there are many areas that are under-sampled.
Geographically detailed information of this sort is not readily available, Hay said, and it is available at a crude national-scale resolution for only 350 of the 1,400 known infectious diseases. This raises the question of whether it is possible
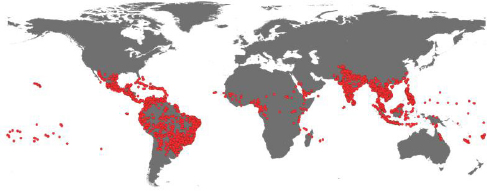
SOURCES: Hay presentation; Messina et al., 2014.
to map all known diseases at a scale that would be useful for prevention and response efforts. For the 350 diseases that haven been mapped at a national level, Hay and his team conducted a systematic review and determined that it would be possible to map 176 of them at a more detailed level, but that only 4 percent of these clinically important infectious diseases have been completely mapped (Hay et al., 2013a).
Prioritization and Automation
Given that it took a year to construct a geospatially detailed map of dengue, the idea of mapping 176 infectious diseases at that resolution seemed implausible to Hay and his colleagues. To prioritize their efforts, they plotted disability-adjusted life-years (DALYs) for each disease against a policy priority ranking based on a review of global health stakeholders, existing literature, and national health priorities (Pigott et al., 2015) (see Figure 3-4). This analysis identified five disease clusters—diarrhea, pneumonia, malaria, human immunodeficiency virus (HIV), and tuberculosis—that have big policy and research interests and huge morbidity and mortality burdens, providing a clear rationale for prioritizing those diseases and the organisms that cause them. Hay cautioned that neither Zika nor Ebola would have appeared in the initial list of 350 potentially mapable diseases. “If you are going to conduct a prioritization exercise such as this you are not going to get all diseases,” he said. Some other tool will be needed for emergency pandemic situations, he added.
The Atlas of Baseline Risk Assessment for Infectious Disease (ABRAID) represents Hay’s attempt to automate the mapping of disease risk using available big datasets (Hay et al., 2013b). ABRAID uses automated data retrieval from sources such as PubMed, HealthMap, and even Facebook, Twitter, and
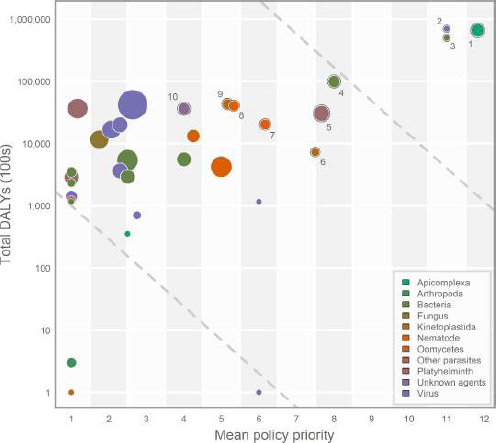
SOURCES: Hay presentation; Pigott et al., 2015.
other lower-provenance data sources. Hay explained that crowdsourcing and machining-learning systems trained by experts are used to validate the accumulated occurrence data, which is then fed into an automated mapping system. His hope is that automating this process will enable real-time updating of the occurrence maps. He noted, for example, that the dengue distribution map generated in 2013 is already out of date and does not reflect the ongoing shifts in dengue distribution that are occurring today.
The ABRAID prototype is complete, and a mechanism for crowdsourcing the validation of the occurrence data is in place, though it had yet to be launched publicly at the time of this workshop. Hay’s team has also finished programming software based on species-distribution modeling that will update the maps in real time for 10 different diseases, including dengue. “Once we have a disease in the system, it will always be up to date and be kept contemporary,” Hay said.
Big Data Inputs
Where Hay sees promise going forward is in using big geopositioned data sources, such as Twitter, Google, and Facebook, to improve real-time disease mapping, assuming that there are useful occurrence data in the hundreds of million tweets and other social media exchanges taking place daily. These data are not meant to replace high-provenance surveillance data from public health agencies and researchers, but Hay said he believes that they could prove useful with appropriate data fusion and a range of tools to help extract signal from noise. His hope is that such data sources can generate usable occurrence data for the many infectious diseases for which there is little or no information about geographical distribution.
As an example of potential applications he hopes to see in the future, Hay concluded his talk by describing his group’s most recent work mapping the environmental suitability for the Zika virus (see Figure 3-5). The group completed the map in a little over a month thanks to the many similarities Zika shares with dengue (Messina et al., 2016). In the event of a public health emergency this type of map could be published and used to estimate where travelers would be at risk of exposure, in this case to Zika virus. “I think big data is the way to speed this process up, both for pandemic preparedness and to filling in the landscape of our geographical knowledge across all diseases,” Hay said.
PREDICTING DISEASE OUTBREAKS
Since 1988, when a global campaign to eradicate poliomyelitis (polio) was started, the number of polio cases has plummeted from as many as 600,000 cases per year to 74 in 2015: 20 in Afghanistan and 54 in Pakistan (Global Polio Eradication Initiative, 2016). Chabot-Couture described the key challenges to eradicate polio as identifying those areas with poor vaccination, which allow the polio virus to continue circulating, and preventing virus reintroduction in those regions cleared of the virus. Traditional datasets may not contain the information needed to discover areas of incomplete vaccination coverage, Chabot-Couture said, so he and his colleagues at the Institute for Disease Modeling have turned to big data to address this shortcoming. Their goal is to develop a model for predicting where polio cases in these two countries are likely to be detected next in order to direct monitoring, vaccinations, and political support activities to high-risk areas.
The data that he and his colleagues used in their model included the location and time of past polio cases, surveys of vaccination coverage, operational data from previous vaccination campaigns, multiple indicator cluster surveys,3 and population distribution, among many others. Even though these data are imperfect and incomplete, Chabot-Couture said that, taken together, these datasets contain patterns and the resulting models are predictive. The Pakistan Polio
___________________
3 More information about multiple indicator cluster surveys is available at http://www.unicef.org/statistics/index_24302.html (accessed October 31, 2016).
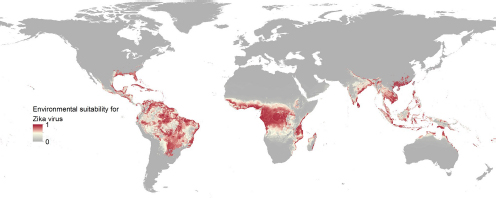
SOURCES: Hay presentation; Messina et al., 2016.
Eradication Program is now using this model to rank each district in the country into four risk categories and to prioritize its resources to those in the highest risk tier. The model is run every 6 to 12 months, depending on epidemiologic findings, Chabot-Couture said. His team has used the same approach to model polio susceptibility in Africa as a tool for understanding where susceptibility to reintroduction exists and to prevent outbreaks in areas that are currently free of polio (Upfill-Brown et al., 2014). They have also successfully modeled measles outbreaks in Nigeria (see Figure 3-6).
Lessons Learned
It is not enough to simply input big datasets into models, Chabot-Couture said. Rather, it is important to extract specific features of the data to include in the model. “You need insights about how to engineer features of your dataset,” he explained. For example, a key feature in the polio datasets is the historical sum of all polio cases in an area. This feature proved to be an excellent predictor of future cases, probably because that feature is measuring chronic underperformance in a way that other datasets are missing. The team found that the same thing was true for measles, he added. Other specific features from the available datasets that informed their models included regional natural immunity and proximity in location and time to reported cases. Chabot-Couture noted, too, that the specific relationship between predictors and outcomes can change from one country to another, but the overall approach for engineering dataset features will remain the same.
Chabot-Couture said that 5 years of experience working with these models has shown that they can identify counterintuitive relationships in the data that can be ignored because they are in fact not linked. For example, the raw data for polio in Nigeria seemed to show that better sanitation was related to an increased risk of developing polio. However, the model showed that this was mostly because
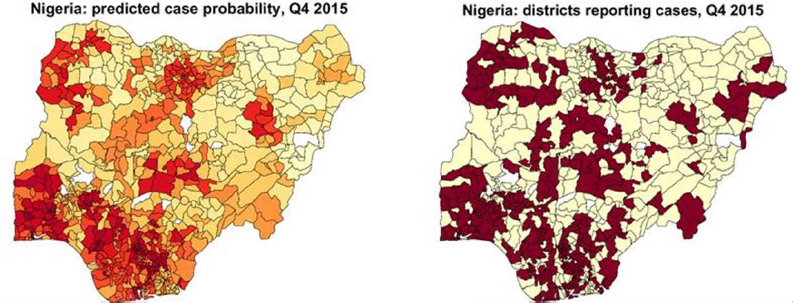
SOURCE: Chabot-Couture presentation.
one particular district that had experienced many cases of polio had subsequently improved sanitation. Chabot-Couture also noted the importance of being careful about noise in the predictors. “If you are having sparse datasets, if you have infrequent data points,” he said, “you want to be careful about the relationship between those noisy data and your outcome.”
Testing Veracity
One problem that Chabot-Couture and his colleagues encountered was that administrative vaccination coverage—a measure of whether a region is supplied with enough vaccines—was often estimated simply by dividing the number of doses distributed in a region by the size of the target population. In at least some cases, the size of the target population, as estimated, was less than the number of doses distributed, which could mean, Chabot-Couture explained, that the program achieved 100 percent coverage, but it could also mean that, for example, only 80 percent coverage was achieved. Coverage surveys would provide a definitive result, but they are expensive and are only conducted every 5 years or so, Chabot-Couture said. The opportunity for big data in this case is to use disease surveillance data—reported cases as well as asking patients how many doses of vaccine they received—as a benchmark and then triangulate toward a more accurate estimate of actual vaccination coverage.
For example, Chabot-Couture said, when the administrative coverage and disease surveillance data are not in agreement, that is an indication that adjustments need to be made. Doing so requires having an understanding of the uncertainty in each dataset. The administrative dataset in this case has high precision because the number of doses is a well-known quantity, but it has a lower accuracy because complete coverage cannot be ensured; the disease surveillance dataset, on the other hand, is sparse and therefore has intrinsic uncertainties associated with it, making it more accurate but less precise. The lesson, Chabot-Couture said, is that comparing multiple imperfect datasets is a useful approach to testing the veracity of data. Concordance will reveal relationships to a common underlying quantity, while outliers will help identify anomalies in the datasets.
When a gold standard dataset is available, it can be used to measure bias and variance in a less accurate but richer dataset, Chabot-Couture noted. He also said that testing for internal consistency can be useful for driving changes in data collection strategy when such a test reveals nonsensical aspects of the data.
In closing, Chabot-Couture said that the opportunities for innovation in the area of big data and analytics do not necessarily involve tackling very large sets of data, but rather derive from bringing multiple pieces of data together to make comparisons and draw actionable insights. He said that data for diseases other than the one of interest can serve as an important ancillary dataset and that higher resolution surveillance and population data are becoming available. The growing amount of genetic sequence data, for example, can provide insights into disease
provenance or movement. He also said that there are mathematical techniques—such as Benford’s law of digit frequency—that can test whether data have been manipulated or falsified.
Concerning the challenges faced when doing this type of work, Chabot-Couture said that the primary one is the limited accessibility to health data related to privacy concerns. “You have to establish collaborations to get access and gain the trust of the people that collect these very large datasets,” he said. Other challenges are that the data platforms that collaborators use may make data sharing difficult, in which case it will be necessary to build an infrastructure to access the data, and that the data may be incomplete or of questionable quality.
DISCUSSION
Dealing with Uncertainty
In response to a question from Jeffrey Duchin from the University of Washington School of Medicine about the wider utility of predictive maps, Hay spoke of the difficulty of conveying the uncertainty that goes along with these predictions. For example, his team spends half of its time developing the correct uncertainty envelopes for the maps, and he does not have a good idea on how to communicate this uncertainty to the many constituencies that would find the maps useful. One aspect of these maps that he finds particularly vexing is the tendency for people to just look at the map and ignore all of the richer detail about uncertainty that his team provides with the maps. Chabot-Couture echoed the importance of uncertainty, noting that understanding uncertainty can prevent overreaction to the changes in a model’s output that result from a change in the underlying data provided to the model. “If you are trying to identify differences over time, seeing if those differences are greater than your uncertainty is a key part to determine whether you need to change the actions or recommendations that you are making,” Chabot-Couture said.
Looking Forward
When asked by Lonnie King, a professor and dean emeritus of the College of Veterinary Medicine at The Ohio State University, to look a decade ahead and talk about innovative strategies for using big data, Chabot-Couture commented that even in an era where a growing percentage of the world’s population uses a mobile phone, the reality is that a great deal of potentially useful data is still being collected on paper and, as Mazet added, lack any detail on where the data were collected geographically. As data collection moves to mobile devices, every data point will have global positioning system coordinates and time stamps. It will also be transmitted directly from the field to data analysis centers, rather than having to make its way from the field to district office and provincial offices and
finally to the one person in federal office that keeps a spreadsheet. As an example, the more than 10,000 vaccination teams in Nigeria now have mobile phones and their activity was tracked every day over the course of a year-long vaccination campaign, which enabled program administrators to ensure that a team visited even the tiniest village. Chabot-Couture predicted that the ways in which Amazon and Google now analyze big data in real time will become the standard in epidemiology and disease surveillance.
Hay is optimistic that as these modeling efforts being conducted in academic institutions continue to prove their worth, organizations with better programming and design capabilities will take over some of this work and develop software that will be more user-friendly, more efficient, and more widely distributed. He also said there is “massive potential” to use advanced geospatial technologies to identify inequities and populations that various disease-prevention efforts are missing and to use that information with boots-on-the-ground approaches to help those underserved groups. Martin Sepúlveda, a recently retired senior physician at IBM’s Watson Research Laboratory, added that the advancement of natural language processing will enable new sources of data to be mined efficiently in response to highly nuanced inquiries.
DIGITAL DISEASE DETECTION4
As Mazet, Hay, and Chabot-Couture noted, public health surveillance—the continuous, systematic collection, analysis, and interpretation of health-related data—can serve as an early warning system for impending health emergencies. Public health surveillance can also be a monitoring and evaluation tool for documenting the impacts of an intervention or for tracking progress toward specified goals, and it can be used to describe the epidemiology of a problem, allowing priorities to be set and informing public health policy and strategies.
The reason public health surveillance is so important in the field of infectious diseases and why it is important to realize the promise of big data to augment public health surveillance, Edelstein said, is that infectious diseases have had such an tremendous impact on human populations over the ages. As recently as the 20th century, influenza outbreaks killed as many as 100 million people worldwide, and the 2014 Ebola outbreak killed at least 11,000 people. “In the 1960s there was a wave of optimism because of the advent of antibiotics, because of the widespread use of vaccines, and because of generally improving health care conditions,” Edelstein said. “There was this idea that infectious diseases were going to be thing of the past, and there is this famous quote from the U.S. Surgeon General in 1967 that it was time to close the chapter on infectious diseases.” Instead, there has been an increase in the number of infectious diseases
___________________
4 Digital disease detection, also known as digital epidemiology, uses electronic data sources, many of which are generated directly by the population through their use of online services, without the sources necessarily having engaged with the health care system (Vayena et al., 2015).
spilling over into humans, including severe acute respiratory syndrome (SARS), HIV, MERS, and most recently Zika. In addition to the human impact, the economic impact of these outbreaks has been in the hundreds of billions of dollars (Newcomb et al., 2011). The H1N1 influenza outbreak in 2009 cost the global economy some $50 billion, Edelstein said, and the SARS outbreak in 2003 cost upward of $60 billion. At an individual country level, the human and economic costs can be devastating.
Quantitative But Slow
Traditionally, public health surveillance has relied on health care workers and laboratories notifying national public health institutes. This system, which is in place in much of the world, provides validated and quantitative data and is clearly tied into national and even global response operations. The drawback is that there can be a significant lag, measured in weeks and months, between the time an outbreak starts and the time the alarm sounds and a response is mounted. In addition, as Mazet noted, this system is only capable of reporting known diseases.
In response, many organizations have begun harnessing the potential of the Internet and online data to supplement the traditional surveillance landscape, Edelstein said. Today, big data analytics, including search engine query analytics, social media analysis, rumor tracking, and mobile phone geotracking, are being tested as tools for disease surveillance. For example, Google Flu Trends used search engine analytics to spot emerging outbreaks of influenza, and between 2009 and 2012 its predictions closely matched data from the Centers for Disease Control and Prevention (CDC), Edelstein said.5 Another source of digital disease detection data is participatory surveillance, which enrolls volunteers to regularly report their health status online.
Using digital data as a tool for disease detection is not easy, Edelstein said, but it is possible to produce valid public health data on a timely basis. One study, for example, found that a combination of six different big data analytic tools was able to detect H5N1 influenza outbreaks an average of 10 days sooner than official public health surveillance (Barboza et al., 2013). HealthMap spotted reports of a “strange new disease of unknown origin with symptoms resembling Lassa fever” on March 14, 2014, some 2 weeks before the Ebola outbreak was detected through formal channels.
New Prospects and New Challenges
In addition to detecting events that would be detected by traditional surveillance, online technologies have the potential to look at health and disease in ways
___________________
5 As Edelstein noted later, Google Flu Trends failed to accurately report influenza cases in 2013. See also Lazer et al. (2014).
that have been previously impossible using traditional surveillance, Edelstein said. For example, geolocating mobile phones calling an Ebola hotline to report cases during the 2014 outbreak provided national public health agencies with leads on new areas of transmission, enabling them to send investigative teams at the initial signs of new outbreaks. Sweden’s public health agency is using a digital disease detection tool—a tool that Edelstein and his colleagues validated using 8 years of retrospective data—to spot the onset of the norovirus season before hospitals report outbreaks. Another example that Edelstein described, which is unrelated to infectious disease detection, involves the use of the so-called dark Web to track trends in transactions of illegal substances or prescription drugs used for recreational purposes, creating a picture of a public health problem that could not be captured using traditional surveillance techniques.
Potential aside, there are technical pitfalls that need to be overcome. The widely reported failure of Google Flu Trends (Lazer et al., 2014) to accurately report influenza cases in 2013 serves as a cautionary note to those developing online predictive tools using non-health-related big datasets, Edelstein said. Given the amount of information available and the increasing odds that for every true positive signal there may be dozens of false positive signals that require investigation, the signal-to-noise issue is particularly challenging. In fact, Edelstein said that he worries that in a landscape of ever-expanding information, the need for human moderation to distinguish signal from noise may not be sustainable.
The representativeness of online data can also skew findings, something Edelstein said that he and his colleagues discovered when they recruited a cohort of Swedes to regularly report health events online. It turned out that those reporting their health events were younger, more educated, and more likely to be female than the population they were supposed to represent. This type of misrepresentation is a particular issue when trying to investigate health events in populations that are disadvantaged and less likely to be regular Internet users, he said.
The Current and Future Landscape
Today, digital disease detection activity largely sits outside of the traditional surveillance landscape. Academia, private technology companies, and other organizations are producing informal, unstructured information that may be fast, flexible, and capable of detecting novel events but that also might be inaccurate and biased, Edelstein said. However, he said he believes that the accuracy and utility of digital data is improving constantly, and he expressed confidence that within a decade, digital online data will be as reliable as the data produced from traditional surveillance sources. Nonetheless, he added, it is clear that digital disease detection will not replace traditional shoe leather epidemiology but will rather complement it as a new tool.
For that to happen, several nontechnical issues will need to be addressed. The most important one, Edelstein said, is the actionability of data generated by these
new sources of information. Digital disease detection, he said, generates valuable public health information, but that information is not tied to a response mechanism. “Until there is a formal link between making those detections and having a response mechanism, the utility of such information is limited,” Edelstein said.
The legal and ethical issues of using data from sources such as Facebook and Twitter for public health purposes without the express consent of users are only beginning to be explored, Edelstein said, as is the challenge of moving data from these new sources into the formal surveillance landscape. In addition, the public health infrastructure, which has the responsibility of responding to public health threats, is not yet equipped to analyze the broad range of different types of data that these unconventional sources generate. “We need to think about how we can standardize these different data sources so that we can transform these data into information that is directly actionable,” Edelstein said.
Confident that these issues will be addressed successfully, Edelstein said he believes that the country-level public health surveillance system will remain at the center of the surveillance system because the mandate to protect populations still rests with governments. However, digital disease detection organizations will become formal partners in this system, feeding actionable data into the surveillance system in a systematic manner. Toward that end, he said, he and his colleagues have begun a project to develop guidance for creating the proper environment to achieve good practices for sharing public health data.
BIG DATA ARCHITECTURE AND ANALYTICS
A typical big data architecture, often called a tech stack, comprises five components, Ordun said: (1) the data sources; (2) the data system, which houses and processes the data; (3) the consumption layer, which queries the data on request of the users; (4) the application program interface (API), which communicates the data between the different components; and (5) the specific client applications, which create reports and visualization (see Figure 3-7). While this is a general picture of a typical data architecture, Ordun stressed that one size does not fit all and that building the right architecture for a specific project or analytic requires starting with a good “use case,” or research question. It also requires determining how much data a specific project needs, given the growing number of available data sources. Often, she said, it is possible to build an architecture strategy using a small subset of the available data. Typically, it takes 9 to 12 weeks to develop a project, the first month of which is devoted to developing a use case and understanding what an organization wants from its data architecture.
The data system layer comprises technologies for importing a wide range of data sources and formats into the system, a means of storing the data—typically in the cloud—and big-data analytics tools to extract, map, batch process, transform, and dynamically change data for analysis. Ordun specifically referenced applications like Hadoop and Spark as technologies to increase computational speed

NOTE: The products and brands shown are exemplary and not exhaustive. API = application program interface.
SOURCE: Ordun presentation.
and data, but she noted that it is important to realize that the pace of technologies change rapidly and thus analytic projects should be built behind strong use cases as opposed to technologies of the moment. The consumption layer is typically customized for a particular use and can include different views of the most common variables being queried. The design of the consumption layer, Ordun explained, depends on the expertise, capabilities, and preferences of a project’s staff; on what databases are currently available; on the preference of administrators; and on the nature of the data. Additionally, within a consumption layer there are many types of databases that a technologist could deploy to package pre-staged views of data. The API layer, which is essentially a set of instructions that send data between the different programs, is designed to support the user’s analytics needs. So, too, are the specific programs in the client application layers.
Once an architecture is in place, the next challenge is to incorporate the capacity to collect, curate, extract, transform, and load the data for analysis—a process that might have taken many hours 5 years ago but which can be accomplished in minutes today using some of the advanced computational data system technologies, Ordun said. While these janitorial tasks are unexciting, doing them
effectively and systematically is what enables the users to explore the data to answer questions and develop analytics efficiently and effectively. Exploring data, Ordun said, is about moving beyond summary statistics and developing an understanding of the key features of a dataset, mapping the data, and identifying relationships in the data that enable modeling activities.
Concerning modeling, Ordun cautioned that it is important to consider the limitations of a model when it comes to looking at the subsets of a dataset. Often, she said, datasets are too large to analyze and thus require sampling, which can produce artifacts and miss rare events. “When dealing with this type of sampling of a big dataset,” Ordun said, “it is important to figure out what your assumptions and your limitations are because in the end, the confidence that your end user has in the results emanating from your analytics will be based on these assumptions. There are limitations. There are outliers. There are clusters. There are rare events. And there are oftentimes missing values. How will you deal with that?” She stressed that understanding the statistical metrics that correspond with your model, and consulting with a statistician is always helpful in any data model development.
Lessons for Building Analytics Applications
Ordun has managed several projects: one that analyzed 2 years of tweets, approximately 1 million tweets in total, using natural language processing to extract the number of people who were hospitalized or sick from food-borne illnesses; one that served as a digital disease detection dashboard for hypothesis testing and forecasting using multiple data sources, including CDC data, weather or climate data, as well as Census data; and a third that is a more complex geospatial analytics application that can superimpose hundreds of geospatial feeds like terrain, land use, or transportation for situational awareness in complex emergencies. Most recently, she helped lead a project with the Food and Drug Administration to create analytics to provide rapid signal detection of adverse events and medication errors using public mobile app reporting, as well as leading the team that developed the mobile application and the cloud architecture.
From these activities, Ordun said, she has learned a few lessons. One is that a data architecture can often start with a “minimal viable product” (MVP) which takes full advantage of the many different available technologies to create an application at the lowest cost and with the highest return possible. An MVP has enough value that people are willing to use it and demonstrates enough future benefit to retain early adopters. It also creates a feedback loop to guide the future development of a more advanced application capable of handling bigger datasets. Another lesson is that data standards vary across data types and formats, but they need to be considered in the analytics pipeline for data processing and preparation. The third lesson is that security remains critical—not an afterthought—and Ordun recommended including data encryption in every layer of the analytics architecture. There
are a variety of ways to encrypt data at rest and data in transit and, for her clients in government, security is a key component to the process from the beginning.
In summarizing, Ordun reiterated the importance of starting with a good question and use case. She also stressed that developing a useful architecture is a team activity that must consider the full team’s capabilities and include statisticians and subject-matter experts during the design process. “Even if your subject-matter expert is not a data scientist,” Ordun said, “it is incredibly important to build this team capability as you frame out your architecture and your analytics,” using the subject-matter experts’ insights, theories, and hypotheses to generate use cases to get to the key result the analytics are intended to deliver. She also recommended against over-engineering an architecture and going for the biggest datasets possible.
COMBATING MICROBIAL RESISTANCE WITH BIG DATA
As an example of how a large, geographically dispersed health care organization is using big data to combat microbial resistance to antibiotics, Lesho described efforts by the DoD to use big data to conduct epidemiologic surveillance of and applied research on multidrug resistant microorganisms. For the DoD, as for other large health care organizations, big data is a reality. For example, each year the DoD generates as many as 18 billion bacterial culture results, Lesho said, a total that does not include the whole-genome sequence data now being generated for many of those cultures or the cultures taken from the DoD’s many military working dogs.
Creating and Storing Big Data
The DoD creates and analyzes big data in three steps, Lesho said. First, it leverages existing public health and infection control surveillance mandates. Second, it creates specialized datamarts or data warehouses containing structured and unstructured data that can be queried and mined in real time to provide alerts and other products. Third, the DoD has established a high-throughput pipeline for organism identification, susceptibility testing, and genome sequencing (Lesho et al., 2014). A unique challenge the DoD faces that other health care systems might not is its high staff turnover rate. “Library preparations for sequencing are demanding, and sometimes, by the time you train an individual, that individual is then rotated out,” Lesho said. To cope with staff turnover, the DoD’s central antimicrobial resistance laboratory uses two high-capacity, liquid-handling robots to prepare samples for five sequencing platforms.
A key feature of the military’s antimicrobial resistance monitoring and research program database is that it links clinical demographic information—including personally identifiable information maintained behind a secure firewall—to a freezer location and the results of the microbial classification
assays, including whole-genome sequencing. The DoD’s data architecture makes heavy use of commercial off-the-shelf software and open-source software as well as some software developed in house. Lesho said that while whole-genome sequencing places a heavy demand on data storage and processing capabilities, it also replaces many conventional genetic fingerprinting tests that can require thousands of different polymerase chain reaction (PCR) runs to characterize all of an organism’s resistance and virulence genes and the mobile genetic elements responsible for transmitting resistance mechanisms between organisms. Concerning the velocity attribute of big data, Lesho said that the DoD’s pipeline consistently delivers a high-resolution outbreak report within 48 hours from the time the laboratory is contacted for outbreak support and receives an isolate. The DoD’s goal is to make its database accessible via the Web so that end users with the right clearance will have access to the data and the analytic tools.
Sample Results
One study that Lesho and his colleagues completed examined the relationship between antibiotic use and the development of carbapenem-resistant Enterobacteriaceae across the entire U.S. military health system (Lesho et al., 2015). The dataset for this study comprised 75 million person-years of surveillance data and 1.97 million cultures from 266 fixed-facility hospitals located around the globe. Lesho’s team examined the data both globally and by region, facility, and antibiotic. The analysis revealed that inpatient quinolone use at the two major referring facilities—one in the Northeastern United States, the other in Texas—correlated with carbapenem resistance, but not fluoroquinolone resistance, in Escherichia coli (E. coli), one of the most common human pathogens and the cause of urinary tract infections.
Another study looked at multidrug resistance in Acinetobacter, a bacterial pathogen that was relatively unknown before the conflicts in Afghanistan and Iraq but that almost led to the closure of Walter Reed Medical Center at one point. The dataset for this study included results from 14.7 million cultures used to identify 360,000 potentially carbapenem-resistant strains (Lesho et al., 2016), and the analysis revealed that isolation of bacteria overseas or isolation from the bloodstream was associated with a higher relative risk of carbapenem resistance. Enterobacteriaceae were isolated 11 times more frequently than either Pseudomonas aeruginosa, an important source of hospital-acquired infections, or all strains of Acinetobacter. However, compared to Enterobacteriaceae, carbapenem resistance was 73-fold higher in Pseudomonas aeruginosa and 210-fold higher in Acinetobacter. As a result of this study, Lesho said, health care providers are now alerted to the fact that if they are overseas and the preliminary lab report says a patient has a non-lactose fermenting gram-negative bacterial infection, that patient is at elevated risk of having a carbapenem-resistant infection and that therapy decisions need to be adjusted accordingly.
Another way in which the military is leveraging its warehoused data is that it is looking for ways of improving stewardship. For example, one of Lesho’s colleagues examined how antibiotics were being prescribed for patients with acute respiratory infections and found that antibiotics were more likely to be prescribed to females, retirees and their dependents, and persons 45 years and older. As a result, the military has been able to target its programs to reduce antibiotic misuse in the treatment of respiratory infections. In addition, this analysis found that 311 patients had received potentially problematic prescriptions or formulations of an antimicrobial agent.
Big Data Is Not Always Better Data
As a cautionary tale illustrating the challenge of data veracity, Lesho discussed a study using big data to identify specific strains of the gram-positive organism Staphylococcus aureus that elude identification by automated vancomycin-susceptibility platforms. Vancomycin-resistant Staphylococcus aureus are rare, and vancomycin-susceptible Staphylococcus aureus are common. However, Lesho explained that there are strains that are technically susceptible but that are clinically problematic and have been shown to be associated with poorer clinical outcomes. Identifying those strains requires a laborious, highly specialized assay that is not feasible for any large health care organization to run on hundreds of thousands of isolates.
The dataset for this study included some 230 million patient encounters, 6.5 million bacterial cultures, and 81,000 unique cultures tested on one of three different assay platforms. Using multiple platforms is a common situation in large, geographically dispersed health care systems (Sparks et al., 2015). The analysis produced a model that determined the point prevalence of the problematic strains and predicted trends in their occurrence and drug usage, Lesho said. Just before issuing the findings, the team’s bioinformatics specialists spotted a potential problem with the data from one particular platform on which about 20 percent of the isolates in the database were tested. In fact, further analysis showed that model results using data from one platform predicted that 54 percent of the isolates would be problematic, while results using data from either of the other platforms predicted a 2 percent incidence. In this case, big data did not prove to be better data, though it did show that one platform was not suitable for trend analysis.
Big Data Challenges for Large Health Care Organizations
Lesho sees four challenges for large health care organizations related to big data: generating the data, storing it, analyzing it, and sharing it. Next-generation sequencing is evolving at a pace that will make it difficult for the lengthy, burdensome, and convoluted procurement processes at many health care systems
and government-funded laboratories to keep up with technological change. The growth of data is placing huge demands on both storage systems and the supply of information technology professionals with the skills to manage the influx of data from a large health care organization. Analyzing and sharing such large datasets also requires access to high-bandwidth local area networks. In response to a question from Rima Khabbaz at the CDC, Lesho said that the growing use of whole-genome sequencing is creating a new problem—laboratories are not culturing organisms anymore, which is a potentially troubling trend, given that it does not currently allow for typing the organisms.