1
INTRODUCTION
Advances in data sciences and applied mathematics that facilitate deep learning and machine learning for computational biology, along with advances in engineering that have enabled automation and high-throughput experimentation, are accelerating discovery within the life sciences.1 The collective progress in these fields and the application of engineering principles to biology have in turn made possible the creation of new products based on biological processes, materials, and information. These products, and the research and development (R&D) that has created them, are changing the face of many industries and stimulating economic activity.2 The term “bioeconomy,” which has emerged over the past two decades as a way to conceptualize this economic activity, has differing attributions, and its meaning is continually evolving. Given that the term links biology and economic activity, moreover, its meaning differs across contexts and countries, reflecting the vast range of natural resources and technological strengths around the world. Despite these variations, more than 40 countries have recognized the potential of a bioeconomy to address a number of societal needs, and have articulated their intent to boost their own bioeconomies by incorporating the concept
___________________
1 For the purposes of this report, the term “life sciences” is intended to encompass the biological, biomedical, environmental biology, and agricultural sciences.
2 Disclaimer: Mention of examples of commercial companies or products made in the report are for illustrative purposes only and are not meant to imply endorsement by the committee; the National Academies of Sciences, Engineering, and Medicine; or any organizations providing funding for this study.
into their policy strategies (El-Chichakli et al., 2016), with the aim of leveraging the power of biology to enable new paths of creation and product development.
The United States has a long history of supporting and growing a vibrant life science research enterprise that is increasingly contributing to the growth of many economic sectors and has provided the nation with many benefits, such as improved health and environment and new and innovative products, generally leading to a better quality of life. The nation currently leads in many biotechnology3 arenas, and also has tremendous natural and agricultural resources and sources of bio-derived feedstocks, both actual and potential, as well as technological capabilities.
The future of the U.S. bioeconomy offers promise of growth and prosperity, and improved quality of life through health and environmental benefits. For example, the bioeconomy offers potential new biobased pathways for creating chemicals, energy sources, and materials, enabling the replacement of traditional inputs such as petroleum feedstocks. Therefore, the bioeconomy can also contribute to climate change mitigation. However, this promise does not come without vulnerabilities and concerns. The many aspects of the bioeconomy rely heavily on a healthy and strong agricultural sector as both a consumer of and a contributing producer of bioeconomy goods and services. Moreover, the nation’s clear leadership in biotechnology will be challenged as other countries make biotechnology investments at increasing rates (enabling them to advance their research and innovation base), reflecting a normal aspect of rivalry in the global economy, and recognizing that U.S. citizens will benefit from bioeconomy advances elsewhere even as the world benefits from U.S. advances. In light of these crucial benefits and the challenges they bring, the Office of the Director of National Intelligence (ODNI) requested that the National Academies of Sciences, Engineering, and Medicine (the National Academies) convene an ad hoc committee to consider how the U.S. bioeconomy can be safeguarded and sustained. This report presents the results of that study.
HISTORY OF THE U.S. BIOECONOMY
Several events led the United States to define and consider areas of structural importance to its own bioeconomy. The Great Recession, a period of significant general economic decline in world markets in the late 2000s and early 2010s, nucleated a series of efforts in the United States to stimulate economic recovery. During this time, the 2009 National Research
___________________
3 Any technological application that uses biological systems, living organisms, biological processes, or derivatives thereof, to make or modify products or processes for specific use.
Council (NRC) report A New Biology for the 21st Century (NRC, 2009) was issued. This report describes the growing power of biology, and explains how biotechnology advances and has critical intersections with a number of scientific disciplines, including computing and engineering, addressing a broad range of human needs in such diverse areas as human health, food and nutrition, energy, and the environment (NRC, 2009). While that report is focused on social benefits, it also points to the deep ties between research innovation and economic benefits.
The following year, the U.S. government first acknowledged the need for strategic planning for the nation’s bioeconomy. In their joint guidance memorandum on science and technology priorities for the fiscal year 2012 budget, the White House Office of Management and Budget and the Office of Science and Technology Policy (OSTP)4 directed agencies to prioritize efforts to promote sustainable economic growth and job creation. Specifically, agencies were advised to “support research to establish the foundations for a 21st century bio-economy” in areas in which “advances in biotechnology and improvements in our ability to design biological systems have the potential to address critical national needs in agriculture, energy, health and the environment.” This specific reference to biotechnology as a key feature of the future U.S. bioeconomy was aligned with the strengths of the nation’s public and private research sectors in cutting-edge engineering biology and big data approaches to harness the potential of biological research for addressing national-scale challenges.
In addition to developing specific guidance for science and technology priorities in federal research to drive the U.S. bioeconomy, considerable effort was focused on reforming the patent system, stimulating economic growth, and enabling entrepreneurs to create new companies and new jobs. The resulting America Invents Act of 2011 (P.L. 112-29) addressed barriers that hindered the key industries of biotechnology, medical devices, and advanced manufacturing. The act was intended to accelerate innovation by providing a fast-track patent application process that would allow applicants to obtain a decision within 12 months, reducing the then-current patent backlog and, importantly, moving the U.S. patent system from a “first-to-invent” to a “first-inventor-to-file” system, thereby aligning U.S. patent policies with those of other patent systems around the world.
In 2012, the National Bioeconomy Blueprint5 laid out strategic objectives that included strengthening relevant R&D efforts, advancing
___________________
4 See https://www.whitehouse.gov/sites/whitehouse.gov/files/omb/memoranda/2010/m10-30.pdf.
5 See https://obamawhitehouse.archives.gov/sites/default/files/microsites/ostp/national_bioeconomy_blueprint_april_2012.pdf.
discoveries from laboratory to market, reducing regulatory barriers, developing a 21st-century bioeconomy workforce, and fostering key public–private partnerships. It also highlighted the need to include biotechnology as a key driver of the U.S. bioeconomy strategy. Since its release, a number of major advances have accelerated the growth of the U.S. bioeconomy:
- The U.S. Department of Agriculture (USDA) expanded efforts to enable the procurement of biobased products through the BioPreferred Program,6 and the BioRefinery Assistance Program (rebranded as the Biorefinery, Renewable Chemical, and Biobased Product Manufacturing Assistance Program7), and the Biomass Crop Assistance Program.8
- Major advances have occurred in engineering biology, including gene-editing approaches involving meganucleases, zinc fingers, transcription activator-like nucleases (TALENs), and clustered regularly interspaced short palindromic repeats (CRISPR).
- The launch of the Precision Medicine Initiative9 occurred in 2016. It aims to use biological data and new analytics tools to derive inferences that can be applied to understand disease and develop diagnostics and treatments.
- In 2016, The Billion Ton Biomass report (USDA and the U.S. Department of Energy [DOE]) provided evidence and data on the potential for 1 billion tons of renewable biomass in the United States to give rise to 50 billion gallons of biofuels/25 percent of liquid transportation fuels, 50 billion pounds of biobased chemicals/products, reductions of 450 million tons of carbon dioxide (CO2) emissions, and 1.1 million direct jobs/$250 million kept in the United States by 2030 (DOE, 2016; Rogers et al., 2017).
- In 2016, DOE established the first open, public biofoundry, the Agile BioFoundry,10 to address precompetitive research challenges identified by industry.
- In 2016, the U.S. National Science Foundation launched its Big Idea initiative, including the Rules of Life Program.11
___________________
6 See https://www.biopreferred.gov/BioPreferred/faces/pages/AboutBioPreferred.xhtml.
7 See https://www.rd.usda.gov/programs-services/biorefinery-renewable-chemical-and-biobased-product-manufacturing-assistance.
8 See https://www.fsa.usda.gov/programs-and-services/energy-programs/BCAP/index.
9 See https://obamawhitehouse.archives.gov/precision-medicine.
10 See https://agilebiofoundry.org/how-we-got-here.
11 See https://www.nsf.gov/news/special_reports/big_ideas/life.jsp.
- The release of the “2017 Update to the Coordinated Framework for the Regulation of Biotechnology” was aimed at increasing transparency, ensuring safety, streamlining regulatory processes, and accelerating the translation of bioinventions to market (EOP, 2017).
- In 2017, USDA released an interagency task force report outlining the need to increase public acceptance of biotechnology products, modernize and streamline the federal regulatory system for such products, and expedite their commercialization, all of which would improve the bioeconomy through biotechnology (USDA, 2017).
- In 2018, LanzaTech partnered with Pacific Northwest National Laboratory and Virgin Atlantic to develop and test new biojet fuel (Bauer and Burton, 2018).
- In 2019, the Biomass Research and Development Board of DOE and USDA issued The Bioeconomy Initiative: Implementation Framework (BRDB, 2019).
- In 2019, the Engineering Biology Research Consortium (EBRC) released its technical research roadmap, Engineering Biology: A Research Roadmap for the Next-Generation Bioeconomy,12 which outlines technical themes and application sectors for engineering biology.
In addition to the previously mentioned 2009 NRC report A New Biology for the 21st Century, a number of more recent National Academies reports have elaborated specific sectors of biotechnology. Among them are the following:
- Industrialization of Biology: A Roadmap to Accelerate the Advanced Manufacturing of Chemicals (NRC, 2015) also speaks to specific aspects of chemical and fuel production via microbial biotechnology. It provides a roadmap for expanding the application of engineering biology in the production of chemicals.
- Genetically Engineered Crops: Experiences and Prospects (NASEM, 2016) showcases progress in the development and use of genetically engineered crops.
- Preparing for the Future Products of Biotechnology (NASEM, 2017) imagines possible developments on a 5- to 10-year horizon and considers regulatory frameworks needed to support them.
___________________
- Biodefense in the Age of Synthetic Biology (NASEM, 2018) considers possible misuse of the powerful tools of synthetic biology.
- Gaseous Carbon Waste Streams (NASEM, 2019) identifies a number of feedstocks (CO2, carbon monoxide [CO], methane [CH4]) with the potential to drive the U.S. bioeconomy.
Beyond these publications, the NRC and the National Academies have worked with the science academies in the United Kingdom and China since 2013 to conduct a series of symposia titled Positioning Synthetic Biology to Meet the Challenges of the 21st Century. Additionally, the NRC and the National Academies convened three workshops in 2014, 2015, and 2016 exploring the bioeconomy, emerging technologies, and security concerns related to life sciences data.
The United States is not alone in seeing the economic advantages that can be derived from having a bioeconomy or from focusing investments in biotechnology. The Organisation for Economic Co-operation and Development, the European Commission, and several European countries individually have written their own related position papers and roadmaps. In 2012, the United Kingdom launched its Synthetic Biology Leadership Council, co-chaired by government and private-sector representatives. And China sees synthetic biology as having potential to accelerate economic growth, having developed its own long-term (20-year) plans and objectives. A detailed discussion of other nations’ approaches to defining their bioeconomies and organizing their bioeconomy strategies can be found in Chapter 2.
The global bioeconomy, then, involves economic rivalry and cooperation among nations, in addition to significant scientific collaboration. Leadership in biotechnology has the potential to lead to economic advantage, whereas falling behind in biotechnology could have a cost, or at a minimum, the cost of lost opportunity.
ADVANCES IN BIOTECHNOLOGY AND THE LIFE SCIENCES
For the U.S. bioeconomy, the innovation process often begins with fundamental research. Fundamental discoveries in basic biology are crosscutting and often agnostic to potential application areas. A revolution in life sciences is accelerating, powered by technologies for reading and writing genomes, facile gene and genome editing, and the leveraging of natural diversity through genome-wide association studies (GWASs) to identify the genes underlying desirable traits. Breakthroughs in systems biology and synthetic biology then provide an unprecedented capacity for engineering plants, animals, and microbes. This cycle of discovery
leading to technology that then amplifies discovery can be illustrated by the four examples described below. These are but a few examples of how the basic research enterprise of discovery science is now fueled by enabling technologies to such an extent that the rate of production of new knowledge continues to accelerate. Although the timescale for translation and advancement is different for different application areas, an important question is how the benefits of these knowledge gains can be translated most effectively into the bioeconomy and into positive impacts on society. The components of the innovation ecosystem that need to be in place to realize the potential of these scientific breakthroughs are discussed later in this chapter.
Example 1: Next-Generation Sequencing
In the last decade of the 20th century, conventional Sanger sequencing and then shotgun sequencing were used to generate the publicly funded sequences of the human genome and those of genetic model species, a painstaking labor involving international research consortia (Shendure et al., 2017). Over the first two decades of this century, next-generation deep-sequencing technologies have built on this foundation, using the previous era’s sequences as definitive libraries against which to match short sequences produced by more modern instruments. Next-generation sequencing reduces the scale (size) of the sample, enabling massively parallel sequencing reactions—the simultaneous sequencing and analysis of millions of oligonucleotides (short strings of DNA bases). Thus, miniaturization of sequencing reactions allowed multiplexing of the number of reactions that can be run in a single experiment, dramatically increasing the speed of data acquisition (Shendure et al., 2017). It became possible to obtain complete genome sequences of prokaryotic organisms and the protein-coding regions of complex eukaryotic organisms on a routine basis and at a cost within the reach of a single investigator in a university. Gene and genome sequences and RNA transcripts can be compared between species, within a species, and within selected populations. Miniaturized equipment for DNA and RNA sequencing is available to further advance field work. And on the near horizon is instrumentation that can read the sequences of single molecules of DNA at the speed of DNA polymerization and in devices about the size of a thumb drive (Jupe et al., 2019).
Geneticists have used association mapping for more than a century to identify causative genes underlying a mutant phenotype. For such diseases as Huntington’s disease and cystic fibrosis for which mutations in a single gene are causative, this is a reasonable approach. However, many human diseases, or desired traits in crops and animals, have a polygenic basis, meaning that more than one gene is likely responsible for the
disease or trait. The availability of thousands of genome sequences from a diseased population versus a healthy population of humans, plants, or animals has allowed identification of the suites of mutations that contribute to the risk of a particular disease. The more sequences are available, the greater is the statistical power in the association of genetic differences between the diseased and healthy populations with disease risk. These GWASs have identified tens or hundreds of new candidate genes13 and biochemical pathways involved in autism, schizophrenia, obesity, and heart disease (Hall et al., 2016). Thus, the research community is beginning to understand the complex molecular bases of these diseases and to provide new therapeutic targets for drug development and diagnosis. Similarly, the application of GWASs to plant and animal populations is contributing to fundamental understanding of growth and development, resistance to stresses, and desired traits such as increased yield (Rai et al., 2019; Sun and Guan, 2018). The first GWAS study was published in 2005, and there are now about 4,000 curated studies with more than 130,000 associations (GWAS catalog14). Companies such as 23andMe and Ancestry DNA utilize GWAS predictions to provide reports to their individual consumers on genetic disease risks or other characteristics with a genetic basis. They can also use the data contributed by consumers in the aggregate to provide pharmaceutical companies with a rich dataset for their own GWAS analyses.
Example 2: Analytical Chemistry
New analytical methods have been developed that can identify the structures and concentrations of chemical species in complex mixtures in plant and microbial cells and in fermentation media, and those generated during the processing of lignocellulosic biomass. These complex mixtures can contain previously unknown compounds of a variety of characteristics (e.g., size, volatility, solubility, polarity, acidity, basicity, ionization energy, reactivity, and concentration). Both high-throughput and highly specialized analyses are now available, including methods based on high-resolution separations, novel ionization and dissociation methods, high-resolution mass spectrometry, and multistage tandem mass spectrometry (Aksenov et al., 2017).
The capability to inventory amounts and types of molecules beyond nucleic acids in living cells has had two major impacts. First, in basic
___________________
13 GWASs have grown in the complexity of the gene networks they can connect. For example, a study examining the genetic basis for height recognizes the potential contributions of roughly 700 genes (Yengo et al., 2018).
research, the accurate characterization of the protein and metabolite contents of living cells revealed the lack of correlation between transcript levels and their translated products, and then the primary and secondary metabolites synthesized by enzymes and enzyme complexes. Using proteomics and metabolomics data, computational modeling of biochemical pathways and their metabolic fluxes provides a systems-level view whereby hypotheses about the effects of perturbation of a component within the system can be tested in silico and then validated experimentally (Ideker et al., 2001). Second, as understanding of living systems increases, it becomes possible to move beyond a description of the system and its parts to the design of new parts and pathways and their genetic control. This mechanistic understanding is used to control native or synthetic pathways at the cell, tissue, and organismal levels. For example, the production of the antimalarial drug artemisinin by an engineered pathway in yeast rather than its native pathway in the plant Artemisia annua15 was one of the first proofs of this concept (Paddon and Keasling, 2014), and was extended to the production of jet fuel precursors in E. coli (Liu et al., 2018).
Metabolic engineering and engineering biology, enabled by new analytical capabilities, are poised to enable use of a national resource of more than a billion tons of lignocellulosic biomass (DOE, 2011, 2016). Beyond ethanol produced from fermentation of biomass-derived sugars, early-stage research is mapping chemical, biochemical, and fast-pyrolytic conversion pathways to liquid hydrocarbon fuels similar to jet fuel, gasoline, or their components (McCann and Carpita, 2015). Lignin in intact woody biomass can be converted efficiently by chemical catalysts to methoxyphenols and then deoxygenated to propylcyclohexane (Parsell et al., 2015), and cellulose can be converted to 5-hydroxymethylfurfural (Yang et al., 2012). Plant species accumulate large amounts of carbon in the form of soluble phenylalanine-derived products and polyketides. Many of these compounds are conjugates of highly reduced aromatic molecules, and together with sugars and aromatics derived from plant cell walls, have the potential to be converted to next-generation fuels or co-products. In one example, Gevo Inc. has blended its renewable jet fuel, derived from wood waste, in test flights, but the current excitement about green (sustainable) chemistry has yet to translate to commercial application.
The bio-derived monomers, although abundant, in these examples represent a tiny proportion of the more than 400,000 kinds of molecules synthesized by living plant cells (Hur et al., 2013). Some natural plant
___________________
15 The penultimate molecule in the reaction, artemisinic acid, can be produced using an engineered pathway in yeast. This molecule undergoes one final chemical reaction to produce the drug.
products have nutritional or pharmaceutical value and form the basis of foods, nutritional supplements (e.g., vitamins), and drugs, while others govern interactions of the plant with its environment (Farré et al., 2014; Fitzpatrick et al., 2012; Martin, 2013). The diversity of plant metabolism thus provides a foundation for metabolic engineering and engineering biology to meet societal goals in biofuels and bioproducts, as well as in food and feed production, biomedicine, and sustainability. Efficient production of target compounds requires systems-level understanding of metabolism and constraints, including trade-offs between carbon fluxes and cellular energy balances. Distributed network control, genetic redundancy, compartmentation of metabolic activities, and multicellularity together increase metabolic complexity in plants, making the design–build–test–improve engineering cycle more challenging than is the case for microbial systems. However, the ability to generate haploids and induce genome duplication such that plants are homozygous for all genomic loci is a breakthrough technology that significantly shortens the timeline for crop breeding (Kalinowska et al., 2019). Future technologies will facilitate both plant metabolic engineering itself and implementation strategies for engineering crops or plant cell cultures as bioproduction systems.
Example 3: Epigenetics
The cloning of Dolly the sheep by reproductive cloning was a technology landmark because it demonstrated that the nucleus of a differentiated cell could be reset to an undifferentiated state from which all cell lineages could be derived (Campbell et al., 1996). Since that landmark was achieved, it has become clear that development and disease in eukaryotic organisms are a function of both mutations in DNA and the modifications to the structure of chromatin that are made during a cell’s or organism’s lifetime, affecting the expression of the gene or genes in that area (epigenetics). The three pillars of epigenetics are methylation of cytosine in DNA; methylation, acetylation, and phosphorylation of the histone proteins around which the DNA is wound on nucleosomes; and RNA-mediated gene-silencing mechanisms that promote heterochromatin formation (Allis and Jenuwein, 2016). These DNA structures and modifications modulate gene expression to maintain the differentiated state of somatic cells. Some epigenetic marks on genomes are now known to occur as a result of chemical exposure, and some chemicals, including morphine, alcohol, and nicotine, show transgenerational effects (Boškovi and Rando, 2018).
In fundamental studies leading to a Nobel Prize, the transcription factors that maintain the pluripotent state of embryonic stem cells were
identified, and shown to be necessary and sufficient to reset fully differentiated somatic cells to a pluripotent state (Takahashi and Yamanaka, 2006). The resulting cells are referred to as induced pluripotent stem cells (iPSCs). These iPSCs can then be induced to differentiate and form organoids, three-dimensional tissue cultures that recapitulate some of the complexity of animal or human organs (Franchini and Pollard, 2015). A patient’s own skin cells, for example, can be reset to iPSCs and then triggered with a specific cocktail of transcription factors to form liver cells. This technology could eventually give rise to organ replacements fully compatible with a patient’s own immune system (Kimbrel and Lanza, 2016). In combination with gene-editing technology, iPSCs and derived organoids have the potential to become patient-specific testbeds for drug responses.
Example 4: Gene and Genome Editing
Basic research investigating the mechanisms by which bacteria protect themselves from viral infections has led to a gene- and genome-editing technology for routine laboratory use (Sander and Joung, 2014). The CRISPR/Cas system uses noncoding RNAs to guide the Cas9 nuclease to induce site-specific double-stranded DNA cleavage. This DNA damage is repaired by cellular DNA repair mechanisms. A single guide RNA is generated to direct the Cas9 nuclease to the specific genomic location. Homologous recombination at the target site allows replacement of endogenous gene sequences with sequence variants encoded in DNA vectors (Lander et al., 2016). Careful genotyping is still required to identify the desired transformants and eliminate transformants resulting from off-target genetic modifications.
The ubiquity of gene editing using CRISPR/Cas9 in public and private research has arisen as the result of a noteworthy conjunction of circumstances. Other methods for creating changes to DNA, such as meganucleases, zinc-finger nucleases, and TALENs, were laborious because the protein recognition domain for each target sequence had to be designed and correctly expressed. Shifting from a system that depended on protein recognition of target DNA sequences to a system that depends on complementary DNA recognition of target DNA sequences simplified and resolved many of the underlying issues of molecular engineering. The rapidity of adoption of CRISPR/Cas9 by the research community is a function of the ease with which the technology can be used for the design of genetic modifications, the affordability of oligonucleotide synthesis, and the low cost of sequencing modified organisms.
One of the first human clinical trials using CRISPR/Cas9 gene editing is now ongoing in the United States for sickle cell disease, led by CRISPR
Therapeutics/Vertex Pharmaceuticals and Sangamo Therapeutics/Sanofi (Collins, 2019). The advantage of blood as an organ for gene editing is that it can be removed from a patient and reintroduced after the treatment. Red blood cells are short-lived and are continuously replaced by hematopoietic stem cells. Sickle cell disease is caused by a single base pair mutation in the beta-hemoglobin protein that reversibly binds oxygen in red blood cells. Unlike normal hemoglobin, the mutated hemoglobin polymerizes inside cells when deoxygenated, injuring the membrane of the cell and causing its rupture, and also distorting the shape of cells in a manner that leads to vaso-occlusion. Two strategies are being explored for efficient editing of induced hematopoietic stem cells (Sugimura et al., 2017) initially derived from sickle cell patients: the single nucleotide polymorphism in the beta-hemoglobin gene itself can be edited to the wild-type sequence, or a repressor of fetal hemoglobin can itself be mutated, leading to expression of normal fetal hemoglobin in adult patients (Bourzac, 2017). The U.S. Food and Drug Administration granted fast-track designation for the CRISPR-based treatment called CTX001 for this latter strategy.16
The convergence of the technologies described in the previous four examples for animal and human disease studies is now easy to imagine: the availability of DNA sequences permits GWAS analyses of healthy and diseased populations, from which candidate genes are inferred by genetic association. Expression of each of the tens or hundreds of candidate genes can be modulated using CRISPR/Cas9 technology in iPSCs and their derived organoids to test hypotheses of development and disease or to provide a testbed for evaluating therapeutic drugs.
FOUR DRIVERS OF THE U.S. BIOECONOMY
As noted earlier, definitions of the term “bioeconomy” vary across different contexts and countries. The focus of this study was on the U.S. bioeconomy, and so it is important before reporting the study’s results to define what the term means in the U.S. context and for the purposes of this report. To this end, the committee identified four defining drivers of the U.S. bioeconomy (see Figure 1-1).
The first is the life sciences—the subdisciplines of biology that yield understanding of all forms of life on Earth. These subdisciplines include botany and agronomy, which focus on plants and agriculture, respectively; microbiology, which studies single-cell organisms; and
___________________
16 See http://ir.crisprtx.com/news-releases/news-release-details/crispr-therapeutics-andvertex-announce-fda-fast-track.
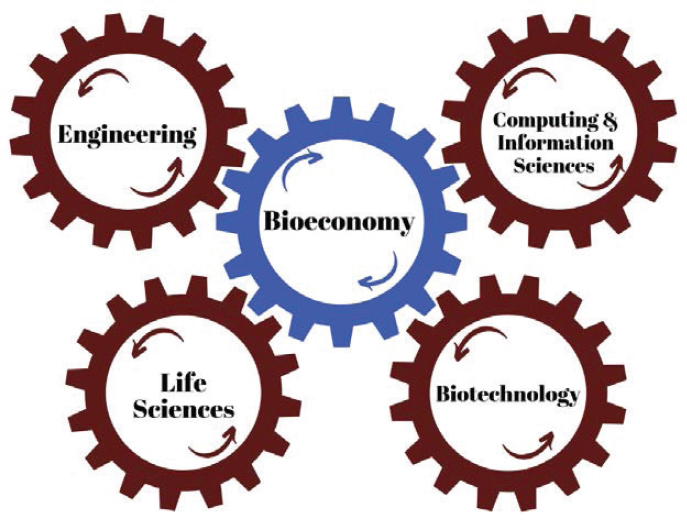
environmental biology, the study of how plants and animals interact with their environment.
Second is biotechnology, which enables understanding biology at the level of genetics, the code for all living organisms. Advances in biotechnology have now made it possible not just to read the genetic code but to write it, and to engineer it to such purposes as curing a disease, improving a crop yield, or addressing an environmental need. Biotechnology advances have also enabled new methods for growing and analyzing cells and tissues, as well as for purifying enzymes for use in driving chemical reactions outside of their native cellular context. The four examples presented in the preceding section—next-generation sequencing, analytical chemistry, epigenetics, and gene and genome editing—are all powerful biotechnology tools that have accelerated the development of applications for the bioeconomy.
Advances in biotechnology require experimentation: to bring a biotechnology drug to market requires millions of experiments, and the same is true for a biotechnology crop or a new detergent enzyme. Engineering has made it possible to automate and miniaturize the experimental process, which enables high-throughput experimentation. Engineering advances in robotics and microfluidics support high-throughput techniques for product development, while advances in analytical techniques
enable use of smaller samples to derive results. In addition to robotics and microfluidics, examples of the use of engineering in the development and production of bioeconomy products include tissue engineering and cell culture, and advanced fermentation. Moreover, the application of engineering principles, such as design–build–test, to biology has greatly accelerated the field of synthetic biology.
Finally, computing and information sciences have made it possible to model experiments mathematically before they are run, as well as to predict outcomes. Experimentation results in large datasets—the “omics” (genomics, proteomics, metabolomics) data from humans, animals, plants, and microorganisms—along with the massive datasets associated with digital imaging. Today, advanced computing techniques such as machine learning are dramatically accelerating the ability to observe nonobvious patterns in large, complex datasets; to make “wise guesses,” eliminating improbable experiments; and to continue to pursue the most promising leads. Biological datasets can also be paired with data from disparate sources, such as medical clinical observations, plant-breeding records, workplace exposure data, family histories, and lifestyle information from social media. Applications of artificial intelligence to these datasets will deepen and accelerate understanding of the interrelation between cause and effect, and between genotype and phenotype. It is this dimension that holds particular promise for the future of the U.S. bioeconomy, and it is also an area in which U.S. leadership in the increasingly global bioeconomy could be disrupted.
The committee’s definition of the U.S. bioeconomy derives from its identification of the above four drivers:
The U.S. bioeconomy is economic activity that is driven by research and innovation in the life sciences and biotechnology, and that is enabled by technological advances in engineering and in computing and information sciences.
The U.S. bioeconomy thus defined rests on both the nation’s natural resources and American ingenuity. It encompasses the products of biological processes and those based on biological feedstocks. It also includes the value chains that have formed to support these research and production activities, such as DNA sequencing services; “foundries” that produce domesticated “host” production organisms and DNA constructs; and consumables that are specific to biotechnology research, such as the ubiquitous 96-well plates and polymerase chain reaction kits. Perhaps most significant, this definition (and thus the bioeconomy) fully embraces the convergence of many different scientific and engineering principles and domains with the life sciences. The transdisciplinary nature of the
bioeconomy is key to its success and growth. It is this aspect that has enabled the bioeconomy to spread to sectors that have traditionally been completely independent of the life sciences.
CONSIDERATIONS IN SAFEGUARDING THE U.S. BIOECONOMY
In studying the overarching question of how to safeguard the U.S. bioeconomy, the committee identified a number of issues that need to be considered. For example, countries around the world rely on goods produced in the United States. Would the United States be comfortable relying on non-U.S. sources of therapies for treatment of U.S. citizens? Or non-U.S. sources for agricultural inputs needed to grow the nation’s food supply? Or foreign biotechnology solutions to critical U.S. environmental concerns? The answers, of course, depend. What are the circumstances in each instance? What is the need? What are the potential consequences in human, environmental, economic, and security terms? On whom would the United States be dependent? What alternatives exist? While addressing all of these questions was beyond the scope of this study, many of the topics and concerns explored in this report feed into those discussions.
Even the process through which the U.S. bioeconomy develops is worthy of examination. The scientific process is collaborative by nature. Scientific processes in the United States are open by intent and by design; openness in science is always preferred. The science and technology enterprise of the U.S. bioeconomy advances through the sharing of data and information and through collaboration among scientists around the world. Sharing works to build scientific expertise while also saving resources, enabling many researchers (in academia or industry, within or outside of the United States) to benefit from initial investments and to validate discoveries made by others. For example, through the use and continued growth of public datasets, researchers can access information without needing to fund the re-creation of those datasets.
While sharing of data and information is desired, certain types of data associated with the U.S. bioeconomy pose privacy and security concerns. In medical research, for example, the privacy of patients’ data, whether their electronic health records or their genomic sequence data, must be assured. This requirement limits what data can be shared and the manner in which it is shared. For instance, genetic data on the U.S. population and subpopulations may reveal vulnerabilities to specific diseases. Similarly, in the agricultural arena, genetic information on vital food crops could reveal vulnerability to disease or heightened susceptibility to genetically enhanced pathogens. Thus, the central issue arises of how to balance the intent to share openly with the legitimate privacy and security concerns involved.
Moreover, openness in science is extended with an expectation of reciprocity. A growing number of countries are restricting the sharing of genetic data or samples that can yield genetic information (conversely, others are sharing even more of these data and samples than is the United States). What is the appropriate response to this growing asymmetry and imbalance in openness in science?
These considerations represent the central impetus for this study.
STUDY CHARGE, SCOPE, AND APPROACH
As mentioned previously, in 2012 OSTP released the National Bioeconomy Blueprint, which laid the groundwork for characterizing and stimulating the U.S. bioeconomy. While the activity that followed focused on the scientific capabilities and potential for societal benefit, and there was some effort to characterize economic contributions in particular domains, little was done to holistically examine the value of the U.S. bioeconomy or assess the risks that relate to the bioeconomy. As a result, questions around the scope and scale of the bioeconomy persisted, a process by which to measure its value was never created, and concerns about the national strategic thinking and the ability to secure and protect the U.S. bioeconomy remained. The committee convened to conduct this study was tasked with delineating the scope of the U.S. bioeconomy, determining how to assess its economic value, identifying potential economic and national security risks related to the bioeconomy and associated policy gaps, considering cybersecurity solutions for protecting bioeconomy data and other outputs of the bioeconomy, and determining a mechanism for tracking future advances and developments within the bioeconomy. The committee’s full Statement of Task is presented in Box 1-1. Importantly, the committee was not asked to determine the value of the bioeconomy; however, in the course of its information gathering, the committee did collect enough data to present a pilot experiment for bioeconomy valuation. The committee was also not tasked to conduct a horizon scan of future innovations in the bioeconomy; rather, this report describes methodologies that could be used to conduct and establish a process for horizon scanning and foresight to enable policy makers to stay abreast of developments in the bioeconomy.
To address its Statement of Task, the committee held three information-gathering workshops in Washington, DC, and three online webinars. Speakers at the workshops and webinars were selected to complement the broad expertise of the committee members and to represent various stakeholder groups within the U.S. bioeconomy. The speaker list for the workshops and webinars can be found in Appendix B. The discussions covered the breadth of the bioeconomy; various perspectives on how to
define the bioeconomy, as well as measure the bioeconomy and assess the value of its various components; and the risks and benefits of the bioeconomy’s various facets. These discussions served as the initial basis for the committee’s deliberations, which were further informed by a review of the relevant literature.
ORGANIZATION OF THE REPORT
This report is organized into four parts addressing the key elements of the committee’s Statement of Task: “Defining and Measuring the U.S. Bioeconomy” (Part I), “Understanding the Ecosystem and Identifying New Trends in the U.S. Bioeconomy” (Part II), “Understanding the Risks Associated with the U.S. Bioeconomy” (Part III), and “Strategies for Safeguarding the U.S. Bioeconomy” (Part IV).
In Part I, the committee presents its perspectives on how to define and measure the bioeconomy. Chapter 2, on defining the U.S. bioeconomy, details the various approaches used by countries around the world to define their bioeconomies and organize their bioeconomy strategies. That chapter also explores the committee’s definition, presented earlier in this chapter, and its interpretation of how that definition sets the parameters of what is included in the U.S. bioeconomy. Chapter 3, on frameworks to measure the value of the U.S. bioeconomy, reviews the various approaches that can be used to assess the value of an economic sector and how those approaches can be applied to the U.S. bioeconomy. In light of its definition of the U.S. bioeconomy, the committee analyzes the data available for conducting such an assessment, undertaking a pilot experiment and examining the robustness of currently available data. In the process, steps were taken to identify data that are missing, not well characterized, or collected in such a way that it is difficult to incorporate them into an assessment of the value of the bioeconomy. The chapter presents a simplified framework for the process the committee undertook in this pilot experiment. Chapter 3 ends with a data-rich discussion of the current direction and status of the U.S. bioeconomy, examining national and private investments and indicators of innovation outcomes (e.g., patents, product approvals, sales). These U.S.-based data are examined in this chapter to prepare the reader for the global comparisons made in the subsequent chapter.
Chapter 4, on areas of leadership in the global economy, presents a detailed examination of the metrics commonly used to determine scientific and economic leadership within a domain. The metrics compared here include government investment in R&D, scientific output (captured in publications and patents), training indicators for students (degrees
granted), investments by private entities (corporations and venture capital firms), and the number of bioeconomy-relevant firms.
In Part II, the committee examines the innovation occurring within the bioeconomy and how new trends and developments can be tracked. Chapter 5, on the ecosystem of the U.S. bioeconomy, explores the nature of the life sciences research enterprise and the associated processes and structures that support and sustain it. This chapter includes examples of how advances in engineering and in computing and information sciences have created new opportunities for growth and development in life sciences research. Chapter 6, on horizon-scanning and foresight methods, assesses the various methodologies for bioeconomy-related horizon scanning and forecasting, providing examples of approaches relevant to the life sciences. The chapter also offers the committee’s assessment of desirable elements for a future-thinking and horizon-scanning mechanism for the bioeconomy.
In Part III of the report, the committee explores the potential risks associated with the bioeconomy and provides its conclusions and recommendations for safeguarding the bioeconomy. Chapter 7, on economic and national security risks pertaining to the bioeconomy, outlines the various risks related to the U.S. bioeconomy, although the committee notes that much of this discussion does not differentiate economic from national security risks, which often cannot be decoupled. Within this chapter, the committee also examines policy mechanisms that can be used to address these risks, pointing out how these policies can be used to mitigate some risks but also may raise additional concerns through the potential for unintended consequences of particular actions.
In Part IV, Chapter 8, the committee presents its overall conclusions and recommendations, explaining their underlying logic and intent, and in some cases discussing different approaches for fulfilling the respective goals. The committee avoided being prescriptive and identified the relevant players when necessary. The committee’s conclusions and recommendations encompass many of the subjects covered in this report, as the committee attempted to take a holistic approach when considering what elements to elevate to the top of its priority list. However, the recommendations are not presented in an order indicating priority, but rather in a manner designed to present a logical and holistic view of the bioeconomy.
REFERENCES
Aksenov, A. A., R. da Silva, R. Knight, N. P. Lopes, and P. C. Dorrestein. 2017. Global chemical analysis of biology by mass spectrometry. Nature Reviews Chemistry 1:0054.
Allis, C. D., and T. Jenuwein. 2016. The molecular hallmarks of epigenetic control. Nature Reviews Genetics 17(8):487–500.
Bošković, A., and O. J. Rando. 2018. Transgenerational epigenetic inheritance. Annual Review of Genetics 52:21–41.
Bourzac, K. 2017. Gene therapy: Erasing sickle-cell disease. Nature 549(7673):S28–S30.
BRDB (Biomass Research and Development Board). 2019. The bioeconomy initiative: Implementation framework. https://biomassboard.gov/pdfs/Bioeconomy_Initiative_Implementation_Framework_FINAL.pdf (accessed October 21, 2019).
Campbell, K. H. S., J. McWhir, W. A. Ritchie, and I. Wilmut. 1996. Sheep cloned by nuclear transfer from a cultured cell line. Nature 380:64–66.
Collins, F. 2019. A CRISPR approach to treating sickle cell. NIH Director’s Blog, April 2. https://directorsblog.nih.gov/2019/04/02/a-crispr-approach-to-treating-sickle-cell (accessed August 30, 2019).
DOE (U.S. Department of Energy). 2011. U.S. Billion-Ton Update: Biomass Supply for a Bioenergy and Bioproducts Industry. R. D. Perlack and B. J. Stokes (Leads). ORNL/TM-2011/224. Oak Ridge, TN: Oak Ridge National Laboratory. https://www.energy.gov/sites/prod/files/2015/01/f19/billion_ton_update_0.pdf (accessed October 21, 2019).
DOE. 2016. 2016 billion-ton report: Advancing domestic resources for a thriving bioeconomy, Vol. 1: Economic availability of feedstocks. http://energy.gov/eere/bioenergy/2016-billion-ton-report (accessed October 21, 2019).
El-Chichakli, B., J. von Braun, C. Lang, D. Barben, and J. Philp. 2016. Policy: Five cornerstones of a global bioeconomy. Nature 535(7611):221–223. doi: 10.1038/535221a.
EOP (Executive Office of the President). 2017. Modernizing the regulatory system for biotechnology products: An update to the coordinated framework for the regulation of biotechnology. https://obamawhitehouse.archives.gov/sites/default/files/microsites/ostp/2017_coordinated_framework_update.pdf (accessed August 30, 2019).
Farré, G., D. Blancquaert, T. Capell, D. Van Der Straeten, P. Christou, and C. Zhu. 2014. Engineering complex metabolic pathways in plants. Annual Review of Plant Biology 65:187–223.
Fitzpatrick, T. B., G. J. Basset, P. Borel, F. Carrari, D. DellaPenna, P. D. Fraser, H. Hellmann, S. Osorio, C. Rothan, V. Valpuesta, C. Caris-Veyrat, and A. R. Fernie. 2012. Vitamin deficiencies in humans: Can plant science help? Plant Cell 24:395–414.
Franchini, L. F., and K. S. Pollard. 2015. Genomic approaches to studying human-specific developmental traits. Development 142(18):3100–3112.
Hall, M. A., J. H. Moore, and M. D. Ritchie. 2016. Embracing complex associations in common traits: Critical considerations for precision medicine. Trends in Genetics 32(8):470–484. doi: 10.1016/j.tig.2016.06.001.
Hur, S. J., S. Yuan Lee, Y.-C. Kim, I. Choi, and G.-B. Kim. 2013. Effect of fermentation on the antioxidant activity in plant-based foods. Food Chemistry 160:346–356.
Ideker T., V. Thorsson, J. A. Ranish, R. Christmas, J. Buhler, J. K. Eng, R. Bumgarner, D. R. Goodlett, R. Aebersold, and L. Hood. 2001. Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 292(5518):929–934.
Jupe, F., A. C. Rivkin, T. P. Michael, M. Zander, S. T. Motley, J. P. Sandoval, R. K. Slotkin, H. Chen, R. Castanon, J. R. Nery, and J. R. Ecker. 2019. The complex architecture and epigenomic impact of plant T-DNA insertions. PLoS Genetics 15(1):e1007819. doi: 10.1371/journal.pgen.1007819.
Kalinowska, K., P. Lenartowicz, J. Namieśnik, and M. Marć. 2019. Analytical procedures for short chain chlorinated paraffins determination—How to make them greener? Science of the Total Environment 671:309–323.
Kimbrel, E. A., and R. Lanza. 2016. Pluripotent stem cells: The last 10 years. Regenerative Medicine 11(8):831–847.
Lander, N., M. A. Chiurillo, and R. Docampo. 2016. Genome editing by CRISPR/Cas9: A game change in the genetic manipulation of protists. Journal of Eukaryotic Microbiology 63(5):679–690. doi: 10.1111/jeu.12338.
Liu, C.-L., T. Tian, J. Alonso-Gutierrez, B. Garabedian, S. Wang, E. E. K. Baidoo, V. Benites, Y. Chen, C. J. Petzold, P. D. Adams, J. D. Keasling, T. Tan, and T. S. Lee. 2018. Renewable production of high density jet fuel precursor sesquiterpenes from Escherichia coli. Biotechnology for Biofuels 11(1):285.
Martin, C. 2013. The interface between plant metabolic engineering and human health. Current Opinion in Biotechnology 24:344–353.
McCann, M. C., and N. C. Carpita. 2015. Biomass recalcitrance: A multi-scale, multi-factor, and conversion-specific property. Journal of Experimental Botany 66(14):4109–4118.
NASEM (National Academies of Sciences, Engineering, and Medicine). 2016. Genetically engineered crops: Experiences and prospects. Washington, DC: The National Academies Press. https://doi.org/10.17226/23395.
NASEM. 2017. Preparing for future products of biotechnology. Washington, DC: The National Academies Press. https://doi.org/10.17226/24605.
NASEM. 2018. Biodefense in the age of synthetic biology. Washington, DC: The National Academies Press. https://doi.org/10.17226/24890.
NASEM. 2019. Gaseous carbon waste streams utilization: Status and research needs. Washington, DC: The National Academies Press. https://doi.org/10.17226/25232.
NRC (National Research Council). 2009. A new biology for the 21st century. Washington, DC: The National Academies Press. https://doi.org/10.17226/12764.
NRC. 2015. Industrialization of biology: A roadmap to accelerate the advanced manufacturing of chemicals. Washington, DC: The National Academies Press. https://doi.org/10.17226/19001.
Paddon, C. J., and J. D. Keasling. 2014. Semi-synthetic artemisinin: A model for the use of synthetic biology in pharmaceutical development. Nature Reviews Microbiology 12(5):355–367.
Parsell, T., S. Yohe, J. Degenstein, T. Jarrell, I. Klein, E. Gencer, B. Hewetson, M. Hurt, J. I. Kim, H. Choudhari, B. Saha, R. Meilan, N. Mosier, F. Ribeiro, W. N. Delgass, C. Chapple, H. I. Kenttämaa, R. Agrawal, and M. M. Abu-Omar. 2015. A synergistic biorefinery based on catalytic conversion of lignin prior to cellulose starting from lignocellulosic biomass. Green Chemistry 17(3):1492–1499.
Rai, A., M. Yamazaki, and K. Saito. 2019. A new era in plant functional genomics. Current Opinion in Systems Biology 15:58–67.
Rogers, J. N., B. Stokes, J. Dunn, H. Cai, M. Wu, Z. Haq, and H. Baumes. 2017. An assessment of the potential products and economic and environmental impacts resulting from a billion ton bioeconomy. Biofuels, Bioproducts and Biorefining 11:110–128. doi: 10.1002/bbb.1728.
Sander, J. D., and J. K. Joung. 2014. CRISPR-Cas systems for editing, regulating and targeting genomes. Nature Biotechnology 32(4):347–355.
Shendure, J., S. Balasubramanian, G. M. Church, W. Gilbert, J. Rogers, J. A. Schloss, and R. H. Waterston. 2017. DNA sequencing at 40: Past, present and future. Nature 550(7676):345–353.
Sugimura, R., D. K. Jha, A. Han, C. Soria-Valles, E. L. da Rocha, Y. F. Lu, J. A. Goettel, E. Serrao, R. G. Rowe, M. Malleshaiah, I. Wong, P. Sousa, T. N. Zhu, A. Ditadi, G. Keller, A. N. Engelman, S. B. Snapper, S. Doulatov, and C. Q. Daley. 2017. Haematopoietic stem and progenitor cells from human pluripotent stem cells. Nature 545(7655):432–438.
Sun, H.-Z., and L. L. Guan. 2018. Feedomics: Promises for food security with sustainable food animal production. TrAC Trends in Analytical Chemistry 107:130–141.
Takahashi, K., and S. Yamanaka. 2006. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126(4):663–676.
USDA (U.S. Department of Agriculture). 2017. Report to the President of the United States from the Task Force on Agriculture and Rural Prosperity. https://www.usda.gov/sites/default/files/documents/rural-prosperity-report.pdf (accessed October 21, 2019).
Yang, Y., C. W. Hu, and M. M. Abu-Omar. 2012. Conversion of carbohydrates and lignocellulosic biomass into 5-hydroxymethylfurfural using AlCl3·6H2O catalyst in a biphasic solvent system. Green Chemistry 14(2):509–513.
Yengo, L., J. Sidorenko, K. E. Kemper, Z. Zheng, A. R. Wood, M. N. Weedon, T. M. Frayling, J. Hirschhorn, J. Yang, and P. M. Visscher. 2018. Meta-analysis of genome-wide association studies for height and body mass index in ∼700000 individuals of European ancestry. Human Molecular Genetics 27(20):3641–3649. doi: 10.1093/hmg/ddy271.






















