4
A Vision for Open Science by Design
SUMMARY POINTS
- In the previous chapters we defined the goals, benefits, and motivations of open science, as well as some barriers and concerns. In this chapter we take a closer look at how open science can be implemented “by design.” We define Open Science by Design as a set of principles and practices that fosters openness throughout the entire research lifecycle.
- Scientific progress is largely influenced by factors that promote—or constrain—the dissemination of knowledge. These factors may be social, economic, or both, and they contribute directly not only to the amount of time it takes a community to fully understand and embrace the implications of scientific discoveries, but also, in some cases, to the successful conduct of science itself.
- Over the past two decades, a related number of activities that include open publications, open data, and open source code have gradually been adopted and shared by increasing numbers of researchers in multiple fields. Many researchers are responding to evidence that openly sharing articles, code, and data in all phases of the research process is beneficial to the research community, to the broader scientific establishment, to policy makers, and to the public at large.
- The committee’s concept of open science by design is by necessity general and idealized. Some discipline-specific nuances cannot be captured in such a broad concept. Also, and importantly, open science by design is intended as a framework to empower the researcher.
PRINCIPLES OF OPEN SCIENCE BY DESIGN
The overarching principle of open science by design is that research conducted openly and transparently leads to better science. Claims are more likely to be credible—or found wanting—when they can be reviewed, critiqued, extended, and reproduced by others. All phases of the research process provide opportunities for assessing and improving the reliability and efficacy of scientific research. As an example, even early in the research process, research plans that are made openly available through a preregistration service, such as Registered Reports
(COS, 2018b), allow review and potential revision of the proposed methodology before data are collected and resources are needlessly expended.
A related principle is that integrating open practices at all points in the research process eases the task for the researcher who is committed to open science. Making research results openly available is not an afterthought when the project is over, but, rather, it is an effective way of doing the research itself. That is, in this way of doing science, making research results open is a by-product of the research process, and not a task that needs to be done when the researcher has already turned to the next project. Researchers can take advantage of robust infrastructure and tools to conduct their experiments, and they can use open data techniques to analyze, interpret, validate, and disseminate their findings. Indeed, many researchers have come to believe that open science practices help them succeed.
The enormous changes effected by the Internet and the wide range of digital technologies that are now available have had an immense impact in all areas of life, and scientific research is no exception. Digital computing technologies have enhanced scientific knowledge discovery in open networked environments, including information retrieval and extraction, artificial intelligence, data mining, and distributed computing, as a new paradigm in the conduct of research (NRC, 2012a). These technologies have the potential to accelerate the discovery and communication of knowledge, both within the scientific community and in the broader society. A principle that ensues is that open science by design is a natural consequence of the fact that “digital science” is rapidly supplanting other ways of doing science.
PRACTICING OPEN SCIENCE BY DESIGN
The researcher is at the center of the practice of open science by design. From the very beginning of the research process, the researcher both contributes to open science and takes advantage of the open science practices of other members of the research community. Practicing open science by design means that researchers develop new habits and customs to promote better science. Figure 4-1 illustrates that open science practices in all phases of the research life cycle are essential to the realization of open science by design.1
The research life cycle begins with the Provocation phase. During this phase, researchers have immediate access to the most recent publications and have the freedom to search archives of papers, including preprints, research software code, and other open publications, as well as databases of research results, including digital information related to physical specimens, all without charge or other
___________________
1 The committee acknowledges the work groups such as of the Center for Open Science and its Open Science Framework (https://osf.io) and FOSTER (https://www.fosteropenscience.eu/content/what-open-science-introduction) in developing innovative approaches to supporting and conceptualizing openness in the research life cycle. There is a long history of explaining and describing the life cycle of information regarding access to research resources to achieve open science.
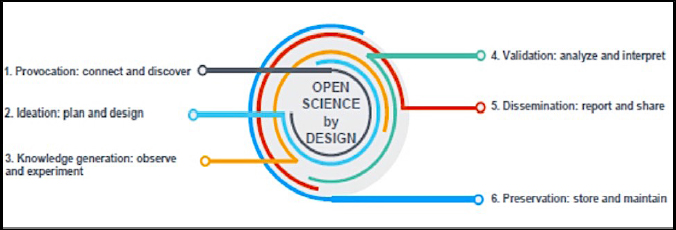
barriers. Researchers use the latest database and text mining tools to explore these resources, to identify new concepts embedded in the research, and to identify where novel contributions can be made. Robust collaborative tools are available to network with colleagues in preparation for the Ideation phase of the research.
During the Ideation phase, researchers and their collaborators develop and revise their research plans. During this phase they may collect preliminary data from publicly available data repositories and conduct a pilot study to test their new methods on the existing data. When applying for research funding, they develop the required data management plans, stating where data, workflow, and software code will be archived for use by other researchers. In addition, in some cases, they may decide to preregister their research plans and protocols in an open repository, as has, for example, become common practice in clinical research. Publicly preregistering the experimental design and analysis plan in advance of data collection is an effective means of minimizing bias and enhancing credibility in a number of fields. Throughout this phase, they pay close attention to the methods and tools they will use during the Knowledge Generation phase, in order to ensure that their final research results will be available in accordance with open principles.
During the Knowledge Generation phase, researchers collect data, using tools that guarantee that the dataset will be stored in an interoperable format and includes appropriate documentation and metadata for easy reuse by other interested researchers at some time in the future. Some data are artifacts, physical samples, and specimens, such as rocks, ice core samples, or tissue samples, and researchers develop concrete plans to archive these data according to disciplinary best practices. With the availability of open software, the researcher can document approaches to cleaning and preparing data for analysis in a research notebook. Electronic research notebooks are both human readable and executable documents that can be run to perform data analyses and are useful in the Validation phase.
During the Validation phase, researchers use open data techniques to analyze, interpret, and validate findings. They present their preliminary findings at conferences and other venues and refine their methods based on relevant comments and critiques. They may deposit their initial working paper in a preprint server of their choice and revise the paper based on the open peer review afforded by the service. They prepare their data in standard formats according to disciplinary norms, and they document and describe both their data and the code that generated their results in optimal ways for reuse and replication. Analysis and interpretation of data are key elements of the scientific process, and the algorithms and workflows for data analysis and interpretation are important research objects in their own right. As they prepare for the Dissemination phase, they review and amend, if necessary, their data management plans to ensure that they have met all of the criteria for making their data and code available for broad and open sharing in an appropriate FAIR repository. As discussed in Chapter 3 and Chapter 5, efforts are ongoing to develop new models of scientific communication that rely on open community review, and where the validation stage follows publication. Implementing such models at a large scale will be an important step forward.
During the Dissemination phase, researchers select the best venue for open publication of their work, including articles, data, code, and other research products. They revise and, in some cases, substantially improve their work based on the comments of the peer reviewers. Journal articles are currently the primary method for summarizing and sharing scientific results, and the journal’s impact factor plays a large role in the assessment of academic achievement. In the digital age, compiling articles in journals for distribution is no longer a requirement for broad distribution. New models are appearing, in which authors publish their work, which then goes through open quality review and certification. The article might then be included in a mega-journal, which are essentially online collections of published articles. Proposals for non-article formats for scholarly communication are also appearing. For example, several neuroscientists have proposed the single figure paper as a form of “nano-publication” that would communicate key findings in a manner optimized for machine-readability (Do and Mobley, 2015). Upon acceptance and before final submission of their work, they select a public copyright license, such as the GNU General Public License for software or a Creative Commons license for other works, including scholarly articles. In preparation for the Preservation phase, they make final adjustments to the metadata that describe their research data and code, making sure that these will be reusable by other interested researchers and specific physical samples are preserved and curated for use by other researchers.
During the Preservation phase, researchers deposit the final peer-reviewed articles in an openly accessible university archive, or they deposit the articles in another publicly accessible archive as required by their research funders. They deposit their research data and software in one or more FAIR data archives, with clear and persistent links that interlink the article, data, and software. Publicly accessible data may then be used by others in the Provocation phase to generate new ideas, marking the beginning of a new research life cycle. Note that data are
often most effectively stewarded and preserved if planned for from the outset, and not at publication time.
Also, and importantly, open science by design is intended as a framework to empower the researcher. As expressed in other NASEM work, the principle for openness of data and other information underlying reported results is that they should ordinarily be available no later than the time of publication, or when the researcher is seeking to gain credit for the work (NAS-NAE-IOM, 2009; NRC, 2003). For journal publication, any sharing prior to the point of final publication is up to the researcher, who is in full control of the decision of when to share.
ENABLING TECHNOLOGIES FOR OPEN SCIENCE BY DESIGN
The practice of open science by design means that the researcher plans for openness right from the start of the research project. Researchers can choose from a growing number of tools, technologies, and platforms as they design and conduct their research. These choices include, for example, the most appropriate data mining algorithms for exploring an unfamiliar dataset, the best workflow tools for capturing and sharing computational workflows, and the established standards for preparing their data for optimal use in FAIR archives. A wide variety of organizations are developing these tools, technologies, and platforms, including community-based nonprofits supported by philanthropy or membership dues, nonprofit coalitions that bring together multiple stakeholders, for-profit startups, and large corporations. A first step might be to register for an author identifier through a service such as ORCID, which provides unique, persistent identifiers for researchers (Meadows, 2016; Wilson and Fenner, 2012). Individuals with an ORCID unique identifier can associate their identifier with their research outputs, whether those outputs are articles, datasets, or other scholarly works. ORCID identifiers can also be used to unambiguously identify researchers in manuscript submission systems, grant application systems, and thesis deposit systems. Because the identifier is unique and persistent, it is not affected by changes in an individual’s location, name, or affiliation.
ORCID’s User Facilities and Publications Working Group brings together “publishers and facilities to better understand research, publication, and reporting workflows” (ORCID, 2018a). ORCID’s Reducing Burden and Improving Transparency (ORBIT) project encourages “funders to use persistent identifiers to automate and streamline the flow of research information between systems” (ORCID, 2018b). Since 2015, Crossref has enabled ORCID records to be automatically updated (Hendricks, 2015). Also, FREYA, a 3-year project funded by the European Commission under the Horizon 2020, builds the infrastructure for persistent identifiers as a core component of open science in the EU and globally.
With eventual publication in mind, researchers can choose an openly available system, such as Docear or Zotero (Beel et al., 2011; Docear, 2018; Vanhecke, 2008; Zotero, 2018) to collect, manage, organize, and format their references. These systems serve as traditional reference managers, but with additional features, including searching and downloading from public databases, tagging and
annotation capabilities, and sharing and exchanging data with collaborators or other software applications. These systems also interoperate with BibTeX, another open source reference manager for those working in the TeX public domain document preparation environment (BibTex, 2018; Hefferon and Barry, 2009; Patashnik, 2003).
Researchers can choose among a variety of open tools for exploring and mining existing datasets. Two popular tools are the R programming language and the WEKA machine learning workbench. The R programming language and software environment is designed for statistical analysis and data mining and integrates with the RStudio user interface (Ihaka, 2010; Tippman, 2015; Verzani, 2011). The WEKA workbench is a toolkit implemented in Java, and like the R software suite, it is also designed to support the entire workflow for experimental data mining, including multiple preprocessing tools, machine learning algorithms, and visualization techniques (Holmes et al., 1994; Eibe et al., 2016).
Automated documentation and sharing of workflows is a key aspect of open science by design. Because many, if not most, areas of science now involve computational analysis of often very large datasets, a variety of tools, both general and domain-specific, have been developed to manage computational data processing and workflows. Importantly, these tools allow researchers to publish their methods and algorithms not only in textual form, but also to publish the code itself, enhancing the reproducibility of the results. In the “Science Code Manifesto,” Barnes et al. (2018) emphasized, “the code is the only definitive expression of the data-processing methods used: without the code, readers cannot fully consider, criticize, or improve upon the methods.” Stodden et al. stated “Access to the computational steps taken to process data and generate findings is as important as access to the data themselves (Stodden et al., 2016; see Box 4-1). Some recently established journals are dedicated to publishing software, including the Journal of Open Source Software and Journal of Open Research Software, allowing authors to receive credit equivalent to “traditional” journal publication for the code they publish (Shamir et al., 2018).
A growing number of scientific journals, including Nature, PNAS, and Science, require authors to make materials, data, code, and associated protocols available to readers (Nature, 2018; PNAS, 2018; Science, 2018). The American Economic Review, the flagship journal of the American Economic Association, has hired a data editor to assist authors with the proper approach to archiving data and code associated with published articles. To incentivize open practices, the Transparency and Openness Promotion (TOP) Guidelines provide a set of recommended standards for scholarly journals to increase reproducibility of research (COS, 2015; Nosek et al., 2015). The TOP Guidelines consist of eight modular standards, with each guideline including three levels of increasing transparency. For example, for the analytic methods (code) transparency standard, a journal that only encourages code sharing or says nothing about it would be at level 0; a journal that requires authors to state whether and where code is available would qualify for level 1; a journal that requires code to be posted on a trusted repository would qualify for level 2; and to qualify for the most transparent level, level 3, the
journal would require that code not only be posted to a trusted repository, but also that reported analyses be reproduced independently before publication.
Mitchum noted that “many are looking to the culture of software programming as a potential model for a more open world of science” (Mitchum, 2015). GitHub, a startup launched in 2008 and originally intended for and used heavily by the open source software development community, is, in fact, increasingly used by researchers as a public platform for sharing their scientific data and code openly (GitHub, 2018; Perkel, 2016). GitHub agreed to be acquired by Microsoft in June 2018 (Ford, 2018). Project Jupyter is an open source framework for scientific software, standards, and services (Project Jupyter, 2018). Jupyter Notebooks, the project’s flagship resource, is a domain-independent, web-based platform for supporting reproducible scientific workflows, from “interactive exploration to publishing a detailed record of computation” (Kluyver et al., 2016). Jupyter works with code in several different programming languages and enables notebook sharing when integrated with the recently developed Binder service that provides a computational environment for users to inspect and execute code, and to publish it seamlessly on GitHub (Forde et al., 2018).
The Center for Open Science’s Open Science Framework (OSF) provides a platform for users to design and create projects, engage with collaborators, manage their research using a suite of tools, prepare their research reports, and preserve their research outcomes. (Foster and Deardorff, 2017; see Box 4-2). The OSF can also be integrated with other open tools, including, notably, support for storage of OSF facilitated research outputs in open repositories. Stodden and colleagues have identified many additional research environments, workflow systems, and dissemination platforms that are now available for researchers’ use across a broad spectrum of academic disciplines (Stodden, 2017; Stodden et al., 2014). In addition, other groups, such as the Research Data Alliance, and Earth Science Information Partners, work with communities to create and disseminate open science and open data tools.
Curated data are the cornerstone of interoperable systems, allowing others to access, understand, compare, and reuse the data stored within those systems in optimal ways. Curation applies throughout the research lifecycle, from the point of data collection to its eventual deposit in an open repository. Many academic disciplines have established and made available community-driven metadata specifications. The Digital Curation Centre maintains an extensive list of domain-specific metadata standards and, in many cases, includes pointers to tools to help implement those standards (Digital Curation Centre, 2018). For example, the ISA initiative has developed a framework and an open source software suite for creating metadata for -omics-based experiments (Sansone et al., 2012), and the Data Documentation Initiative has developed an international standard and a set of tools for describing data in the social and behavioral sciences (Vardigan, 2013).
There are a variety of efforts underway to ensure that datasets are reliably cited, such that they can be found and appropriately attributed (Altman and Crosas, 2013). Although these practices are not yet as mature as article citation practices, their importance is beginning to be acknowledged. In 2012, the National Academies of Sciences, Engineering, and Medicine (NASEM) hosted a workshop that addressed various dimensions of this topic, including technical requirements, legal and socio-cultural aspects, and disciplinary considerations (NRC, 2012b). Recent community-driven efforts have resulted in a set of principles for data citation (Data Citation Synthesis Group, 2014), the Data Citation Implementation Pilot (DCIP) project (Cousijn et al., 2017), as well as implementation guidelines focused more specifically on scholarly data repositories (Fenner et al., 2016). In
addition, the Center for Expanded Data Annotation and Retrieval (CEDAR), a standards-based metadata authoring system developed under the NIH Big Data to Knowledge Program, is a good example of a general-purpose tool for creating standards-based metadata in a domain-independent manner and that fits into the data submission pipeline for open repositories. There is strong agreement that datasets should have a persistent, globally unique method for identification that is both human understandable and machine-actionable.
The Digital Object Identifier (DOI) is a system for identification of content on digital networks. DOI identifiers are persistent, unique, resolvable, and interoperable for management of content on digital networks (Paskin, 2010). The system is implemented through a federation of registration agencies under agreed upon policies and common infrastructure and is now overseen by the Swiss DONA foundation (DONA). The DataCite organization was founded in 2009 to support data archiving through data citation (Neumann and Brace, 2014; DataCite, 2018). The organization provides persistent DOIs for research data, providing data citation support and services to researchers, data centers, journal publishers, and funding agencies. Many general open data repositories, including Dryad, Figshare, and Zenodo, assign DOIs to the data they store.
As the calls for FAIR research archives have increased, a number of European efforts, including the OpenAIRE project and the European Open Science Cloud, have initiated large-scale infrastructure projects. OpenAIRE’s focus is on developing a technical infrastructure for an interoperable network of research repositories from throughout Europe through the establishment of common guidelines and shared metadata standards (Schirrwagen et al., 2013; OpenAIRE, 2018). The project involves the collaboration of researchers from several scientific disciplines as well as librarians, data and information technology experts, and legal specialists. OpenAIRE is collaborating with a number of other groups internationally, including the US-based SHARE (SHared Access Research Ecosystem) project. The SHARE project was launched in 2013 by the Association of Research Libraries, the Association of American Universities, and the Association of Public and Land-grant Universities to strengthen efforts to identify, discover, and track research outputs (COAR, 2015b; Hudson-Vitale et al., 2017; SHARE, 2018).
The European Open Science Cloud (EOSC) project grounds its work in the EOSC Declaration, which includes recommendations and implementation suggestions in the areas of data culture and FAIR data, research data services and architecture, and governance and funding. (EC, 2017a). EOSC has expanded its scope beyond Europe, and they, together with many others, acknowledge that because scientific knowledge is not confined within national boundaries, a globally interoperable open research infrastructure is needed (NASEM, 2018c; Wittenburg and Strawn, 2018).
STRENGTHENING TRAINING FOR OPEN SCIENCE BY DESIGN
As researchers adopt the habits and practices of open science by design, they may need help in identifying and making use of the most effective tools and approaches to use at various stages of their work.2
Several recent studies have noted that training of researchers early in their careers is critical, suggesting that open science training can be integrated into existing graduate curricula (OECD, 2015; McKiernan et al., 2016). Others have addressed the practical guidance and training that is needed to help researchers learn how to open up their research processes and results (Carvalho, 2017; EC, 2017f; FOSTER, 2018). The FOSTER (Facilitate Open Science Training for European Research) project, for example, has developed a suite of open science training materials, including courses on open science policies, practices, and resources, research workflow and design, text and data mining methods, data management, legal issues, and responsible conduct of research (See Box 4-3). As described in Chapter 3, the Committee on Data for Science and Technology (CODATA) also supports educational opportunities for early career researchers.
Librarians and other information professionals who have extensive experience in the preservation, publication, and dissemination of digital scientific materials, may, nonetheless, need additional training to address the challenges of curating large-scale open data resources. Several years ago, NASEM undertook a consensus study that examined the career paths for individuals working in the field of digital curation, which they defined broadly as the “active management and enhancement of digital information assets for current and future use” (NRC, 2015). They concluded that although the number of educational opportunities has grown, the available opportunities are well below what is needed to meet the demands of the current data-rich era. Individuals who have discipline-specific knowledge as well as skills in computer and information science are best suited to meet these demands.
Similarly, a recent report from the EC noted that “there is an alarming shortage of data experts both globally and in the European Union” and that there is a “chasm” that needs to be addressed between those who work on digital infrastructure and scientific domain specialists (EC, 2016). An analogous point is made in the current strategic plan for the National Library of Medicine. In discussing the intersection between biomedical data science and open science, the report suggests that more training is needed that involves “enhancing the computational and statistical skills of researchers with biomedical knowledge, and training computer scientists to apply their work to biomedical problems.” (NLM, 2018a). The National Science Foundation’s Research Traineeship Program trains graduate students in high priority interdisciplinary research areas, including “Harnessing the Data Revolution” (NSF, 2018b).
___________________
2 Indeed, the OSTP 2013 memo explicitly highlights the importance of supporting “training, education, and workforce development related to scientific data management, analysis, storage, preservation, and stewardship” (OSTP, 2013).
There are a growing number of collaborative activities among universities, nonprofit organizations, and the philanthropic community related to open science training. The University of California at Riverside and the Center for Open Science (COS) have initiated an NSF-supported randomized trial to evaluate the impact of receiving training on the use of the Open Science Framework for managing, archiving, and sharing lab research materials and data (McKiernan et al., 2016; Nosek, 2017; COS, 2018c). The COS also works with the University of Virginia on an NIH supplemental grant for the Biotechnology Training Program that develops and presents reproducible and open practice curriculum content (COS, 2018d).
Since 2012, the Berkeley Initiative for Transparency in the Social Sciences at the University of California, Berkeley has developed coursework to promote open science in social sciences research (BITSS, 2018). The Data Curation Network of nine major academic institutions, supported by the Alfred P. Sloan Foundation, provides a cross-institutional staffing model for training data curators to build an innovative community to promote data curation practices (Johnston et al., 2017; Data Curation Network, 2018). The Gordon and Betty Moore Foundation and the Alfred P. Sloan Foundation, in partnership with several universities, have recently created the Data Science Environments project to “advance data-intensive scientific discovery.” Among their efforts is the development of educational materials for researchers at all levels of their academic careers (MSDSE, 2018).
Several related areas of education and training might provide opportunities to incorporate content related to open science practices. First, the field of data science is rapidly growing and evolving. According to NASEM (2018a, p. 1), “Data science is a hybrid of multiple disciplines and skill sets, draws on diverse fields (including computer science, statistics, and mathematics), encompasses topics in ethics and privacy, and depends on specifics of the domains to which it is applied.” Those trained in data science at the undergraduate and graduate levels will go on to a wide range of careers, many outside of research. Nevertheless, the core skills and capabilities of data science are clearly relevant to the practice of open science.
In addition, several federal research agencies mandate that some subset of students or trainees that they support receive responsible conduct of research (RCR) training (NASEM, 2017b). Progress toward open science by design will certainly affect the treatment of some traditional RCR topics such as data handling and responsible authorship. RCR training and education programs might benefit from new approaches that incorporate more open science content as a way of ensuring that students and other researchers possess the knowledge and skills needed to practice open science by design.
OTHER CONSIDERATIONS
The committee’s concept of open science by design is by necessity general and idealized. It is important to note that research methods and processes vary by
field, and that some discipline-specific nuances cannot be captured in such a broad concept. As discussed in Chapter 3, some fields rely on large data resources that are shared by a defined community. In other fields, experimental data are generated by individuals or research groups, and there may or may not be incentives or rewards for making data openly available. Many disciplines are focused on the study of nondigital data and materials, and face the challenge of ensuring long-term availability. In fields that are focused on the study of one-time phenomena such as earthquakes, specific practices such as preregistration may not make sense or add value.
Likewise, publication cultures vary widely. Some fields such as physics and economics have a long history of utilizing preprints, while this practice is just beginning to gain popularity in the life sciences. In computer science, conferences are a central mechanism for the dissemination of results, and conference proceedings are more important publication venues than are journals (Vardi, 2010). Different fields can and should adapt the open science by design concept to fit their practices and circumstances.
In addition, open science by design raises some questions about roles and responsibilities that have not been fully resolved. For example, should the burden of deposition in publicly accessible archives be shouldered by the organization (publisher or otherwise) that provides services for Dissemination and Preservation, as opposed to the researcher? In order to secure researchers’ rights and assuage concerns about reuse and credit, should the choice of license be determined as soon as the research is disseminated, even if it allows for downstream changes (with a separate license for preservation)? Should the researcher be solely responsible for semantically linking research products or should this responsibility be shared?
Given that this is a U.S.-based project undertaken by a U.S. committee, much of the report’s discussion and analysis reflects the U.S. experience. However, international perspectives and examples are utilized frequently. At first, wide availability of the tools and infrastructure needed to practice open science by design may be limited to researchers working in well-resourced institutions, primarily in developed countries. Over time, as open science by design demonstrates its value, tools and resources will become more widely available. For example, the African Academy of Sciences recently launched AAS Open Research as “a platform for rapid publication and open peer review for researchers supported by AAS…” (AAS, 2018).
Finally, in order to take hold as a core concept for the future of research, open science by design needs to serve the needs of early career researchers. Chapter 2 discusses how a lack of incentives and supportive culture around open practices is a particular problem facing early career researchers. Open principles and practices must demonstrate their value by enabling early career researchers to become more effective, productive scientists than they would be in a closed environment.














