A12
DISRUPTION OF INSECT TRANSMISSION OF PLANT VIRUSES1
Anna E. Whitfield2and Dorith Rotenberg
Summary
Plant-infecting viruses are transmitted by a diverse array of organisms including insects, mites, nematodes, fungi, and plasmodiophorids. Virus interactions with these vectors are diverse, but there are some commonalities. Generally the infection cycle begins with the vector encountering the virus in the plant and the virus is acquired by the vector. The virus must then persist in or on the vector long enough for the virus to be transported to a new host and delivered into the plant cell. Plant viruses rely on their vectors for breaching the plant cell wall to be delivered directly into the cytosol. In most cases, viral capsid or membrane glycoproteins are the specific viral proteins that are required for transmission and determinants of vector specificity. Specific molecules in vectors also interact with the virus and while there are few-identified to no-identified receptors, candidate recognition molecules are being further explored in these systems. Due to the specificity of virus transmission by vectors, there are defined steps that represent good targets for interdiction strategies to disrupt the disease cycle. This review focuses on new technologies that aim to disrupt the virus–vector interaction and focuses on a few of the well-characterized virus–vector interactions in the field. In closing, we discuss the importance of integration of these technologies with current methods for plant virus disease control.
Introduction
The virus transmission cycle involves host-finding, feeding and acquisition of virus, transport and delivery of virus to a new host plant (see Figure A12-1). Each step in the transmission process provides an opportunity for interdiction. Strategies for disrupting transmission are the focus of this review and we highlight recent biotech-based approaches to reduce vectorial capacity and populationreduction approaches that utilize the specificity of the virus–vector interaction to target insects.
___________________
1 Reprinted from Current Opinion in Insect Science, Vol. 8, Pages 79-87, Copyright 2015, with permission from Elsevier.
2 Department of Plant Pathology, Kansas State University, Manhattan, KS 66502, USA.
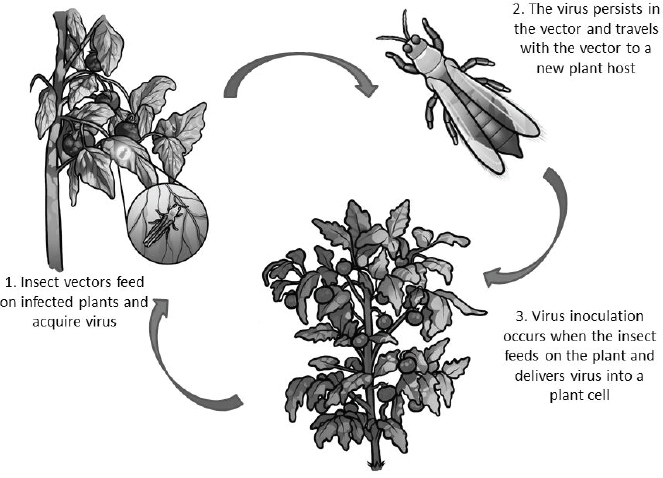
Overview of the Mechanisms and Methods of Plant Virus Transmission
Plant virus transmission by insects is classified into two major categories: non-circulative and circulative transmission. The non-circulative-externally borne viruses associate with specific cuticular structures of the insect stylet or foregut (see Figure A12-2) and the attached virus particles are lost during the insect molt (reviewed in Ng and Falk, 2006; Blank et al., 2014).
Non-circulative viruses are transmitted in a non-persistent or semi-persistent manner which means that they are acquired within seconds to minutes of feeding and transmitted rapidly as well. Semi-persistent viruses require longer periods to be acquired and transmitted (minutes to hours). By contrast, the circulative or internally-borne viruses require a greater time for acquisition and transmission (hours to days) and must traverse the gut and reach the salivary glands for transmission to occur. These viruses are not lost during insect molts and have a latent period between initial acquisition and transmission. The latent period is synonymous with extrinsic incubation period in animal vector biology. For all types of insect transmission, viral determinants of transmissibility have been defined. For the non-circulative viruses, some viruses bind directly to insect stylets or foreguts and other viruses need the assistance of another viral protein(s) that serves as a bridge between the insect structures and the virion (Chen et al., 2011;
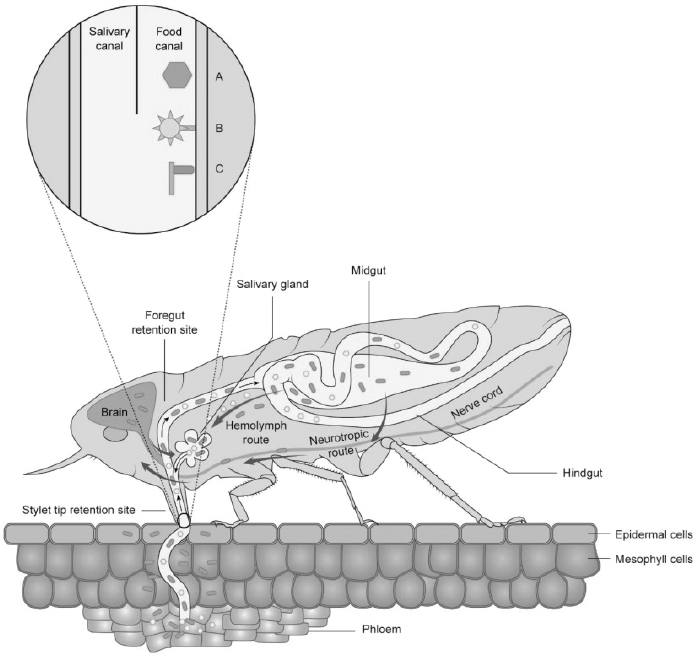
SOURCES: Modified from Blanc et al., 2014, and Ammar et al., 2009.
Liu et al., 2006; Lung and Pirone, 1974; Govier and Kassani, 1974). For the circulative viruses, the viral capsid proteins and glycoproteins have been identified as viral determinants of insect transmission (reviewed in Hogenhout et al., 2008). Similarly, for the viruses transmitted by soil-dwelling plant–virus vectors (nematodes, fungi, and plasmodiophorids) the viral coat protein(s) is responsible for binding and retention in the vector (Ohki et al., 2010; Andret-Link et al., 2004; Adams et al., 2001). Despite being transmitted by different mechanisms, the requirement of a viral protein–insect molecular interaction is a consistent theme in transmission by insects and provides a common target for interrupting the transmission process.
Blocking Virus Transmission with Viral Capsid Proteins and Glycoproteins
Viral proteins are required for attachment and/or entry into the insect vector. Therefore, exploiting these proteins for their specific binding affinities to vector tissues is an obvious approach for blocking virus acquisition and transmission. For all the vector-borne plant viruses, a specific viral protein(s) is required for virus transmission. Genomes of plant viruses are quite small, and defining the viral attachment protein(s) (VAP) has been completed for diverse and seemingly intractable virus–vector systems. Using this knowledge, recombinant VAP can be used to (1) reduce transmission of viruses by blocking virus binding and subsequent dissemination in the vector and (2) reducing the vector population using the viral protein to deliver toxic cargo to the insect (see Table A12-1).
Exploiting Viral Proteins to Control Vectors of Circulative Viruses
For circulative viruses, the structural proteins of the viral capsid are the determinants of insect vector specificity (reviewed in Grey et al., 2014). The route of virus dissemination has been well-characterized for members of the family Luteoviridae and the coat protein (CP) and the readthrough extension of the coat protein are required for transmission. Luteovirids are small icosahedral virions (25–30 nm) that are composed of a major coat protein and a minor protein that has a carboxy-terminal extension termed the readthrough domain (RTD). Initial virus entry occurs in the insect gut and the specific region for entry varies with virus species, occurring in the midgut or hindgut. Several studies have documented that the coat protein is sufficient for delivery of virus into the hemocoel and the RTD is crucial for transmission. It is thought that the salivary glands are the barrier to transmission of particles with mutations in the RTD (Brault et al., 2000; Peter et al., 2008; Liu et al., 2009). Knowledge of Pea enation mosaic virus (PEMV) CP binding and movement through the insect gut was used to target a hemocoel-active toxin to aphids (Bonning et al., 2014). The authors found that a recombinant CP fused to non-viral toxin peptides could be delivered via transcytosis from the aphid gut to the hemocoel to be aphicidal.
TABLE A12-1 A Summary of Strategies Used to Disrupt Plant Virus–Vector Interactions
| Disruption strategy | Virus | Vector | Mode of transmission | In planta experiments conducted | References |
|---|---|---|---|---|---|
| Blocking virus entry into vector using a viral protein | Rice ragged stunt virus (RRSV, Oryzavirus, Reoviridae) | Planthopper, Nilaparvata lugens | Circulative, propagative | Yes | Guoying et al., 1999; Shao et al., 2003 |
| Tomato yellow leaf curl virus (TYLCV, Begomovirus, Geminiviridae) | Whitefly, Bemisia tabaci | Circulative | No | Wang et al., 2014 | |
| Tomato spotted wilt virus (TSWV, Tospovirus, Bunyaviridae) | Thrips, Frankliniella occidentalis | Circulative, propagative | Yes | Whitfield et al., 2004, 2008; Montero-Astua et al., 2014 | |
| Viral coat protein/toxin fusions | Pea enation mosaic virus (PEMV, Enamovirus, Luteoviridae) | Aphids, Acyrthosiphon pisum, Myzus persicae, Aphis glycines and Rhopalosiphum padi a | Circulative | Yes | Bonning et al., 2014 |
| Insect-gut binding peptide/toxin fusions | PEMVb | Aphids, A. pisum, M. persicae | Circulative | No | Liu et al., 2010; Chogule et al., 2013 |
| Targeting vector proteins that interact with virus | Rice stripe virus (RSV, Tenuivirusc) | Planthopper, Laodelphax striatellus d | Circulative, propagative | No | Huo et al., 2014 |
a A. glycines and R. padi do not transmit PEMV but were susceptible to the PEMV CP-toxin fusion protein.
b Peptide identified to bind aphid vector guts and reduced PEMV access to insect hemocoel.
c Member of a floating virus genus; tenuiviruses are closely related to the family Bunyaviridae however they lack an envelope.
d Disrupted transovarial transmission by RNA-interference of vitellogenin.
The benefit of using this system is that luteovirids are transmitted specifically by aphids. Additionally, the insect gut is not the major barrier to luteovirids entry into the insect and the salivary gland appears to be a more significant barrier to aphid transmission of these viruses. Additionally, the CP-toxin fusion killed non-vector aphids but had no apparent effect on an off-target lepidopteran species, Heliothis virescens. Begomoviruses are transmitted in a similar circulative manner by whitefly vectors and the viral CP was shown to bind to whitefly midguts and reduce the amount of virus in whiteflies in feeding experiments (Wang et al., 2014). The ability of viral CPs to bind to insect guts and block virus entry indicates that preventing virus entry and delivering toxic peptides may prove to be transmission inhibition-based approaches for other viruses that circulate through the insect body.
An alternative strategy to CP-mediated transport of toxins to aphid vectors has been documented with the use of aphid gut-binding peptides. A bio-panning approach identified a 12 amino acid peptide that bound to pea aphid guts (Liu et al., 2010). Interestingly, this peptide, GBP3.1, reduced PEMV abundance in the vector for up to 70 minutes after acquisition of the peptide. Although the primary amino acid sequence of GBP3.1 was dissimilar to the PEMV CP sequence, structural similarity was identified between the peptide and a specific surface loop of the viral protein, suggesting that reduced virus abundance may have resulted in competitive binding for gut molecules between the peptide and the virus.
The utility of this aphid binding peptide has been exploited to expand the target range of a Bacillus thuringiensis cytolytic toxin, Cyt2Aa (Chougule et al., 2013). The GBP3.1 peptide was incorporated into the surface loops of the toxin and the modified toxin bound aphid membranes. The modified toxin retained activity and was found to be toxic to Acyrthosiphon pisum and Myzus persicae. Modification of insect-specific toxins with the addition of aphid-binding peptides and/or virus CP is a promising new control strategy for vector and non-vector aphids.
Disruption of Transmission of Circulative, Propagative Viruses Using Viral Proteins
Tomato spotted wilt virus (TSWV) is an enveloped negative strand RNA virus and the type member of the genus Tospovirus within the family Bunyaviridae. Tospoviruses are transmitted in a circulative-propagative manner exclusively by thrips vectors, including Frankliniella occidentalis, the western flower thrips (Whitfield et al., 2005). Efficient transmission to plants requires that thrips acquire TSWV during the larval stages to transmit as adults. When vector competent larval thrips feed on infected tissue, the virus enters the insect midgut, initiates a high titer infection in the gut, and then disseminates to the salivary gland tissues. The virus traverses several membrane barriers en route from the vector midgut to salivary glands (Kritzman et al., 2002; Nagata et al., 2002;
Moritz et al., 2004), and virus titer was documented to be positively correlated with the number of TSWV transmission events by individual female and male thrips (Rosenberg et al., 2009). Collectively, these studies highlight the importance of virus accumulation and spread in the vector as quantitative determinants of a successful transmission event.
The structure of the TSWV virion is characteristic of members of the family Bunyaviridae, and the virion is spherical and composed of an outer membrane envelope derived from the host. Two glycoproteins (GPs) are embedded in the membrane and project from the surface. The GPs are designated GN and GC and thrips transmission of TSWV maps to the M segment, the viral RNA that encodes the GPs (Sin et al., 2005).
Due to the unique biology of the TSWV–thrips interaction, there is a narrow window of opportunity for virus acquisition during larval development that is a good target for blocking virus entry. Defining the molecular determinants of a plant virus–vector interaction enabled the development of novel virus control strategies that aim to specifically disrupt the interaction. TSWV acquisition is mediated by the molecular interaction between the virus membrane glycoprotein GN, which serves as a viral attachment protein, and the thrips midgut. Previously, we found that an exogenously-applied soluble form of GN (GN-S) inhibits TSWV binding, acquisition (Whitfield et al., 2004), and transmission to a plant host (Whitfield et al., 2008). We generated transgenic tomato plants expressing a soluble form of GN and found that thrips that fed on these transgenics had significantly lower virus titers and adult transmission efficiencies than thrips fed on TSWV-infected non-transgenic tomato plants (Montero-Astúa et al., 2014). These results demonstrate that an initial reduction in virus infection of the larval insect midgut can result in a significant decrease in virus titer and transmission over the life-span of the vector.
The inhibition results with GN-S and TSWV are supported by the results of research with Rice ragged stunt virus, which is a Reovirus that is transmitted in a circulative, propagative manner by rice brown planthoppers (Guoying et al., 1999; Shao et al., 2003). In those experiments, the viral spike protein inhibited virus transmission and insects that were fed a nonstructural virus protein exhibited no transmission inhibition. These results support the concept of disrupting the insect-mediated transmission of viruses via viral attachment proteins. Future work with this transmission-blocking strategy will focus on the spectrum of efficacy, that is, does TSWV GN block other related tospoviruses and transmission by other thrips vectors. This research is important because new tospoviruses of significance to agriculture have been recently described including Soybean vein necrosis-associated virus (SVNaV) and a naturally-occurring interspecies reassortant between Groundnut ringspot virus (GRSV) and Tomato chlorotic spot virus (TCSV) (Zhou et al., 2011; Webster et al., 2011).
Potential for Disruption of Transmission of Noncirculative Viruses
Similar strategies as those described above for circulative viruses may provide reasonable methods for disrupting transmission of non-circulative viruses due to the specific nature of binding and retention documented for these viruses. The non-circulative viruses generally bind to cuticular surfaces of the insect body including the insect stylet and foregut. Many of the stylet-borne viruses are associated with specific regions of the stylet, and virions that are successfully transmitted to host plants are those that bind to the distal tip of the stylet where the food and salivary canals merge (Uzest et al., 2007; Wang et al., 1996).
Work with Cauliflower mosaic virus (CaMV) binding to aphid stylets has also directed attention to the presence of a specialized region of the aphid maxillary stylet termed the “acrostyle,” an electron-dense area where virions of CaMV are specifically retained (Uzet et al., 2010). For the semi-persistent criniviruses, virion bind to the whitefly foregut and the minor coat protein (CPm) is the VAP (Chen et al., 2004). For Cucumber mosaic virus, the coat protein is the primary determinant of aphid transmission and helper proteins have not been identified. Specific regions of the virion including a surface loop and the quasi-threefold axis of symmetry have been shown to be essential for virus transmission by aphids (Liu et al., 2002; Bricault and Perry, 2013). Plants and bacteria have been engineered to produce viral proteins that bind to the insect stylet or foregut. Using these tools, determining if excess helper component or coat protein can compete with virions to saturate binding sites in the vector to subsequently prevent virus attachment is an exciting avenue to pursue for this category of vector-transmitted viruses.
Disruption of Other Insect-Borne Plant Pathogens
Much like with plant viruses, recent work has focused on blocking transmission of other arthropod-borne plant pathogens. The plant pathogenic bacterium, Xylella fastidiosa, is transmitted by hemipteran (leafhopper) vectors and is retained in the vector foregut. Unlike non-circulative plant viruses, X. fastidiosa cells attach to the foregut and replicate in the insect and this is termed non-circulative propagative transmission. Like plant viruses, the bacterial cells that attach to the foregut are lost during insect molts. Progress has been made toward identifying the bacterial components of the interaction, pointing to afimbrial adhesins as playing a major role in pathogen attachment to the vector (Killiny and Almeida, 2009). Complementary studies using antibodies to various bacterial cell-associated proteins and molecules confirmed the role of afimbrial adhesins (carbohydrate-binding proteins) in transmission (Killiny et al., 2012). Additionally, competition assays with lectins and carbohydrates confirmed the importance of these host carbohydrate-bacterial protein interactions in X. fastidiosa transmission by leafhopper vectors. Exogenous application of excess amounts of N-acetylgucosamine carbohydrates reduced pathogen transmission
indicating that this specific carbohydrate may be a vector component recognized by X. fastidiosa, facilitating binding of bacteria to the vector foregut (Killiny et al., 2012). Conversely, competition experiments with carbohydrate-binding proteins (lectins) also disrupted the interaction presumably by binding the carbohydrates on the foregut attachment site and blocking X. fastidiosa attachment. This work supports the hypothesis that pathogen retention in insect vectors is mediated by specific interactions and highlights commonalities in vector transmission of diverse types of pathogens.
Potential Use of RNAi for Disruption of Plant Virus Transmission
RNAi is an insect control approach that can also be used to directly target insect vectors and is considered to be the basis for the next generation of transgenic plants designed for insect pest control (Gordon and Waterhouse, 2007). RNAi is an attractive option for plant-feeding insects because dsRNA can be delivered via transgenic expression in plants, transiently-expressed by viral vectors (i.e., attenuated plant viruses), or exogenously-applied by soil drench or foliar sprays to plants. For insect control, silencing of an insect gene by endogenous RNAi cellular machinery is triggered by delivery of dsRNA of the same sequence to the gene transcript into the cell. Once inside, dsRNA is recognized by Dicer, an RNAse III enzyme that cleaves the RNA into short interfering RNAs (siRNAs). The siRNAs are incorporated into the RNA-induced silencing complex (RISC) which then targets the degradation of homologous transcript sequences by the activity of the endonuclease, Argonaute. In insects, systemic silencing occurs when the dsRNA signal spreads in the body of the insect by cellular uptake mechanisms that are still under investigation for various insect pest species. For example, several well-characterized insect genomes have SID orthologs, multispanning transmembrane proteins essential for systemic RNAi in Caenorhabditis elegans, while other mechanisms including endocytosis (V-ATPases) and scavenger receptor-mediated and other pattern recognition receptor-mediated uptake, that is, innate immunity, have been proposed (Huvenne and Smagghe, 2010; Winston et al., 2002; Feinberg and Hunter, 2003). Functional analysis of genes implicated in these cellular processes in insects including virus vectors remains a need. To add to the mystery, to date, there is no evidence of the presence of RNA-dependent RNA polymerase in insects and therefore amplification of the RNAi signal, a process that leads to transitive RNAi as documented for ticks, C. elegans, and plants (Gordon and Waterhouse, 2007).
The success of RNAi varies depending on the insect species and developmental stage targeted, dsRNA delivery method, length and concentration of dsRNA input, and gene target (Scott et al., 2013). An effective target for several insect vectors of plant viruses has been the V-ATPase gene and has been shown to reduce insect life-span and egg production (Yao et al., 2013; Khan et al., 2013). Genome and transcriptome-wide data-mining projects will likely aid in the
identification of additional RNAi gene targets unique to vector insects, thereby addressing concerns of off-target effects. In addition, the discovery of essential interactions for vector transmission may yield additional targets for pest control and reduction of virus transmission by RNAi. Targeting the vector components is challenging because of the relatively large genome sizes and ploidy of insect vectors (Hanrahan and Johnston, 2011; Jacobson et al., 2013), and vector transmission strategies among vector–virus systems are vastly different, and thus insect gene targets are likely more diverse and may vary between tissue types (i.e., receptors in guts vs. salivary glands) in the same insect vector. Despite challenges of working with non-model insects, significant progress has been made in defining vector molecules that interact with viruses.
Virus-Interacting Proteins for Circulative, Propagative Viruses
Propagative viruses have been the focus of proteomics examination of viral–vector interactions. A common theme emerging is the involvement of insect proteins associated with virus transport and dissemination in the vector. A yeast two-hybrid screen with the reovirus, Southern rice black-streaked dwarf virus (SRBSDV), and the planthopper vector, Sogatella furcifera, revealed 153 putative interactions between P7-1, the major nonstructural viral protein that induces host cell tubular structures that serve as conduits for virus movement, and the insect vector (Mar et al., 2014). Key partners identified in the screen are consistent with the role that P7-1 is thought to play in virus movement (cytoskeletal network and endomembrane system) and the potential insect host response to infection (ubiquitin proteasome and nervous system). Experiments with the tenuivirus Rice stripe virus (RSV) and the vector, Laodelphax striatellus, revealed insect proteins that also provide insight into the biology of tissue tropism in the vector (Huo et al., 2014). Using a yeast two hybrid screen, vitellogenin, the major yolk protein precursor of egg-laying animals, was identified to interact with the RSV major nucleocapsid protein, pc3. In this study, RNAi-knockdown of vitellogenin transcripts demonstrated the importance of the protein in transovarial transmission of the virus. These findings support the hypothesis that RSV directly binds and ‘hijacks’ the vitellogenin transport route to enter L. striatellus oocytes.
In a separate study, vitellogenin was the most abundant transcript in RSV-infected planthoppers in a global gene expression analysis indicating that in addition to possibly capitalizing on the natural route of vitellogenin into oocytes, induced expression of this protein may enable more efficient transovarial passage of the virus (Zhang et al., 2010). Proteomic analysis of F. occidentalis in response to TSWV infection identified 26 protein spots that varied in density between the virus-infected and non-infected larval thrips (Badillo-Vargas et al., 2012). The differential proteins included nine proteins that are putatively associated with steps in the viral infection cycle of entry and exit from insect vector cells (i.e., localization to membranes and protein transport) and 14 proteins associated with
response to stimuli, including nine that are potentially involved in the defense response to TSWV infection. Virus-interacting proteins and proteins that respond to virus infection provide potential targets for disruption of transmission and/or for silencing by RNAi.
Virus-Interacting Proteins for Circulative, Nonpropagative Viruses
A variety of virus-binding and responsive proteins have been identified using a combination of protein-protein interaction discovery methods including virus overlay, yeast two-hybrid, co-immunoprecipitation, and proteomics. For luteovirids transmitted by aphids, the development of aphid offspring from crosses between efficient and inefficient vectors, Schizaphis graminum, enabled the genetic mapping of barriers to transmission (gut vs. salivary gland) and provided a resource for identification of proteins associated with vector competence (Burrows et al., 2006; Yang et al., 2008; Tamborindeguy et al., 2013). Virus overlays and proteomics both revealed the importance of cyclophilins which are proposed to play a role in virus transcytosis through the insect gut and possibly salivary gland tissues. The presence and abundance of cyclophilin proteins and isoforms has been associated with vector competence in the S. graminum F2 genotypes of genetic crosses biotypes and field collected biotypes (Tamborindeguy et al., 2013). Other proteins identified to interact with viruses in the genera Luteovirus and Polerovirus include cytoskeletal proteins such as actin and tubulin; additional endocyctic pathway proteins RACK1, GAPDH3, and luciferase-like proteins; and cuticular proteins with chitin-bind 4 domains (Burrows et al., 2006; Yang et al., 2008; Tamborindeguy et al., 2013; Seddas et al., 2004; Cilia et al., 2011). The peptide (GBP3.1) that was found to bind pea aphid guts and reduce the PEMV delivery into the insect hemocoel was also found to bind to a large aphid protein that was identified as an aminopeptidase, a candidate receptor for entry of luteo-viruses into the gut of the aphid vector (Chougule et al., 2013).
Virus-Interacting Proteins for Noncirculative Viruses
Vector proteins have also been implicated in binding and transmission of non-circulative viruses. Virus overlay assays identified two types of green peach aphid (M. persicae) proteins that bound to the helper component protein of potyviruses. Zucchini yellow mosaic virus (ZYMV) helper component protein (HC-Pro) bound to cuticular proteins extracted from whole aphid bodies (Dombrovsky et al., 2010), and using a similar approach but enriching for aphid heads (the stylet is site of virus attachment) ribosomal S2 proteins bound to Tobacco etch virus (TEV) HC-Pro (Fernandez-Calvino et al., 2006). Because non-circulative viruses bind to specialized regions of the insect stylet or foregut, it is hypothesized that once virus binding partners in the vector are identified, strategies to saturate binding sites can be deployed to prevent viral protein binding and subsequent
transmission. Verifying virus–vector protein interactions in vivo is an essential step in documenting the validity of interactions identified under in vitro conditions or using heterologous systems such as yeast two-hybrid screens. The use of RNAi and advanced imaging technologies will be an important component of interaction studies and has already proven to be useful for validation of interactions identified within vectors of propagative viruses.
Closing Remarks
Our global community faces the mounting threat of newly emergent and re-emerging viruses on the world food supply. Of those viruses, 70 percent of plant-infecting viruses are transmitted from one host to another by arthropod vectors (Hogenhout et al., 2008). As the human population increases from 7 billion to a predicted 9 billion by 2050, it is crucial that plant science research aims to bolster food security by developing reliable, safe, and sustainable plant virus control strategies. The identification of unique steps in the viral infection cycle that enable the design of molecules that interfere with the infection process is a promising approach for virus disease control. The development of new biotechnology-based strategies to reduce transmission by vectors and to decrease vector populations is attractive because they target pathways in the transmission cycle. However, the long-term effectiveness of these control methods relies on their judicious use and incorporation into existing virus-control and vector-control regimes. One approach to increase the durability of the new biotech-based control strategies would be to “stack” these novel traits with traditional virus and vector resistance genes or combine multiple biotech approaches such as deploying transgenic plants that co-express a viral protein to block virus acquisition and dsRNA hairpins to target vital genes in vector populations. The integration of new technologies with traditional integrated pest management strategies (IPM) such as altering planting date and reflective mulches to reduce vector landing rates will also extend the shelf-life of biotechnological traits. This is particularly important for managing resistance to viruses as they have great potential for genetic change and have been shown to rapidly overcome single-target resistance strategies (Lafforgue et al., 2011; Lopez et al., 2011).
Other promising strategies that deserve further exploration for vectors of plant viruses include, firstly, insect transgenesis and secondly, microbial manipulation to reduce vector transmission (Alphey, 2014). Transgenic insects expressing viral dsRNA have been shown to elicit RNAi to reduce virus loads and prevent dissemination to the salivary glands (Franz et al., 2006), thus rendering these transgenics refractory to virus. This type of strategy could be applied to circulative, propagative plant viruses. An alternative transgenic approach is population suppression by introduction of a lethal gene into the population. The development of the ‘release of insects carrying a dominant lethal genetic system’ (RIDL) has been highly effective for mosquito vectors of human-infecting pathogens in lab
experiments (Alphey, 2014; Alphey and Alphey, 2014; Massonnet-Bruneel et al., 2013).
There are several field studies that have examined the effectiveness and persistence of the transgenic insect strategy (Harris et al., 2012; Lacroix et al., 2012). For insect vectors that reproduce sexually, the RIDL technology could provide new ways to reduce plant vector populations. Metagenomics and microbiome studies have directed attention to the influence of microbes on multiple biotic processes and ecological interactions, including virus transmission. The use of microbial manipulation to alter vector competence and/or capacity is an emerging field of study. For example, the mosquito-Wolbachia system has led to exciting findings and several examples of pathogen reduction and stimulation have been documented (Martinez et al., 2014; van den Hurk et al., 2012; Dodson et al., 2014). Wolbachia and other endosymbionts are commonly found in plant-feeding insects and the potential role of microbes in transmission by plant virus vectors warrants further exploration and manipulation (Zhang et al., 2010; Bing et al., 2014; see also Pinheiro et al., 2015). With the increase in tools available for the control of viruses and their vectors, the next phase of this research is to move discoveries from the lab to the field. With the vast array of tools available and the collaborative networks that can be developed, it appears that virologists will be up for the challenge of feeding a growing global population while keeping our environment healthy and productive.
Acknowledgements
This work was supported by the USDA NIFA/AFRI grant numbers 2012-68004-20166 and 2007-35319-18326 and by the National Science Foundation CAREER grant IOS-0953786. Contribution no. 15-281-J from the Kansas Agricultural Experiment Station.
References
Adams, M.J., J.F. Antoniw, and J.G.L. Mullins. 2001. Plant virus transmission by plasmodiophorid fungi is associated with distinctive transmembrane regions of virus-encoded proteins. Archives of Virology 146(6):1139–1153.
Alphey, L. 2014. Genetic control of mosquitoes. Annual Review of Entomology 59:205–224.
Alphey, L., and N. Alphey. 2014. Five things to know about genetically modified (GM) insects for vector control. PLoS Pathogens 10:e1003909.
Ammar, E., C. Tsai, A.E. Whitfield, M.G. Redinbaugh, and S.A. Hogenhout. 2009. Cellular and molecular aspects of Rhabdovirus interactions with insect and plant hosts. Annual Review of Entomology 54:447–468.
Andret-Link, P., C. Schmitt-Keichinger, G. Demangeat, V. Komar, and M. Fuchs. 2004. The specific transmission of grapevine fanleaf virus by its nematode vector Xiphinema index is solely determined by the viral coat protein. Virology 320:12–22.
Badillo-Vargas, I.E., D. Rotenberg, Y. Hiromasa, J.M. Tomich, and A.E. Whitfield. 2012. Proteomic analysis of Frankliniella occidentalis and differentially-expressed proteins in response to tomato spotted wilt virus infection. Journal of Virology 86(16):8793–8809.
Bing, X., W. Xia, J. Gui, G. Yan, X. Wang, and S. Liu. 2014. Diversity and evolution of the Wolbachia endosymbionts of Bemisia (Hemiptera: Aleyrodidae) whiteflies. Ecology and Evolution 4(13):2714–2737.
Blanc, S., M. Drucker, and M. Uzest. 2014. Localizing viruses in their insect vectors. Annual Review of Phytopathology 52:403–425.
Bonning, B.C., N. Pal, S. Liu, Z. Wang, S. Sivakumar, P.M. Dixon, G.F. King, and W.A. Miller. 2014. Toxin delivery by the coat protein of an aphid-vectored plant virus provides plant resistance to aphids. Nature Biotechnology 32(1):102–105.
Brault, V., J. Mutterer, D. Scheidecker, M.T. Simonis, E. Herrbach, K. Richards, and V. Ziegler-Graff. 2000. Effects of point mutations in the readthrough domain of the beet western yellows virus minor capsid protein on virus accumulation in plants and on transmission by aphids. Journal of Virology 74(3):1140–1148.
Bricault, C.A., and K.L. Perry. 2013. Alteration of intersubunit acid–base pair interactions at the quasi-threefold axis of symmetry of cucumber mosaic virus disrupts aphid vector transmission. Virology 440(2):160–170.
Burrows, M.E., M.C. Caillaud, D.M. Smith, E.C. Benson, F.E. Gildow, and S.M. Gray. 2006. Genetic regulation of Polerovirus and Luteovirus transmission in the aphid Schizaphis graminum. Phytopathology 96(8):828–837.
Chen, A.Y.S., G.P. Walker, D. Carter, and J.C.K. Ng. 2011. A virus capsid component mediates virion retention and transmission by its insect vector. Proceedings of the National Academy of Sciences of the United States of America 108:16777–16782.
Chougule, N.P., H. Li, S. Liu, L.B. Linz, K.E. Narva, T. Meade, and B.C. Bonning. 2013. Retargeting of the Bacillus thuringiensis toxin Cyt2Aa against hemipteran insect pests. Proceedings of the National Academy of Sciences of the United States of America 110:465–8470.
Cilia, M., C. Tamborindeguy, T. Fish, K. Howe, T.W. Thannhauser, and S. Gray. 2011. Genetics coupled to quantitative intact proteomics links heritable aphid and endosymbiont protein expression to circulative polerovirus transmission. Journal of Virology, 85:2148–2166.
Dodson, B.L., G.L. Hughes, O. Paul, A.C. Matacchiero, L.D. Kramer, and J.L. Rasgon. 2014. Wolbachia enhances West Nile virus (WNV) infection in the mosquito Culex tarsalis. PLoS Neglected Tropical Diseases e29658.
Dombrovsky, A., N. Gollop, S. Chen, N. Chejanovsky, and B. Raccah. 2007. In vitro association between the helper component-proteinase of zucchini yellow mosaic virus and cuticle proteins of Myzus persicae. Journal of General Virology, 88:1602–1610.
Feinberg, E.H., and C.P. Hunter. 2003. Transport of dsRNA into cells by the transmembrane protein SID-1. Science 301:1545–1547.
Fernandez-Calvino, L., E. Goytia, D. Lopez-Abella, A. Giner, M. Urizarna, L. Vilaplana, and J. Jose Lopez-Moya. 2010. The helper-component protease transmission factor of tobacco etch potyvirus binds specifically to an aphid ribosomal protein homologous to the laminin receptor precursor. Journal of General Virology 91(11):2862–2873.
Franz, A.W.E., I. Sanchez-Vargas, Z.N. Adelman, C.D. Blair, B.J. Beaty, A.A. James, and K.E. Olson. 2006. Engineering RNA interference-based resistance to dengue virus type 2 in genetically modified Aedes aegypti. Proceedings of the National Academy of Sciences of the United States of America 103(11):4198–4203.
Gordon, K.H.J., and P.M. Waterhouse. 2007. RNAi for insect-proof plants. Nature 25(11) 1231–1232.
Govier, D.A., and B. Kassanis. 1974. Virus-induced component of plant sap needed when aphids acquire potato virus-Y from purified preparations. Virology 61:420–426.
Gray, S., M. Cilia, and M. Ghanim. 2014. Circulative “Nonpropagative” virus transmission: An orchestra of virus-, insect-, and plant-derived instruments. Advances in Virus Research 89:141–199.
Guoying, Z., L. Xiongbin, and L. Huijuan. 1999. Rice ragged stunt oryzavirus: Role of the viral spike protein in transmission by the insect vector. Annals of Applied Biology 135(3):573–578.
Hanrahan, S.J., and J.S. Johnston. 2011. New genome size estimates of 134 species of arthropods. Chromosome Research 19(6):809–823.
Harris, A.F., A.R. McKemey, D. Nimmo, Z. Curtis, I. Black, S.A. Morgan, M.N. Oviedo, R. Lacroix, N. Naish, N.I. Morrison, et al. 2012. Successful suppression of a field mosquito population by sustained release of engineered male mosquitoes. Nature Biotechnology 30:828–830.
Hogenhout, S.A., E.D. Ammar, A.E. Whitfield, and M.G. Redinbaugh. 2008. Insect vector interactions with persistently transmitted viruses. Annual Review of Phytopathology 46:327–359.
Huo, Y., L. Wenwen, Z. Fujie, C. Xiaoying, L. Li, L. Qifei, Z. Yijun, W. Taiyun, F. Rongxiang, and X. Wang. 2014. Transovarial transmission of a plant virus is mediated by vitellogenin of its insect vector. PLoS Pathogens 10(3):e1003949.
Huvenne, H., and G. Smagghe. 2010. Mechanisms of dsRNA uptake in insects and potential of RNAi for pest control: A review. Journal of Insect Physiology 56:227–235.
Jacobson, A.L., J.S. Johnston, D. Rotenberg, A.E. Whitfield, W. Booth, E.L. Vargo, and G.G. Kennedy. 2013. Genome size and ploidy of Thysanoptera. Insect Molecular Biology 22(1):12–17.
Khan, A.M., M. Ashfaq, Z. Kiss, A.A. Khan, S. Mansoor, and B.W. Falk. 2013. Use of recombinant tobacco mosaic virus to achieve RNA interference in plants against the citrus mealybug, Planococcus citri (Hemiptera: Pseudococcidae). PLoS One 8(9):e73657.
Killiny, N., A. Rashed, and R.P.P. Almeida. 2012. Disrupting the transmission of a vector-borne plant pathogen. Applied and Environmental Microbiology 78(3):638–643.
Killiny, N., and R.P.P. Almeida. 2009. Xylella fastidiosa afimbrial adhesins mediate cell transmission to plants by leafhopper vectors. Applied and Environmental Microbiology 75(2):521–528.
Kritzman, A., A. Gera, B. Raccah, J.W.M. van Lent, and D. Peters. 2002. The route of tomato spotted wilt virus inside the thrips body in relation to transmission efficiency. Archives of Virology 147(11):2143–2156.
Lacroix, R., A.R. McKemey, N. Raduan, L.K. Wee, W.H. Ming, T.G. Ney, S.A.A. Rahidah, S. Salman, S. Subramaniam, O. Nordin, et al. 2012. Open field release of genetically engineered sterile male Aedes aegypti in Malaysia. PLoS One 7:e42771.
Lafforgue, G., F. Martinez, J. Sardanyes, F. de la Iglesia, Q. Niu, S. Lin, R.V. Solé, N. Chua, J.-A. Daròs, and S.F. Elena. 2011. Tempo and mode of plant RNA virus escape from RNA interference-mediated resistance. Journal of Virology 85:9686–9695.
Liu, S., S. Sivakumar, Z. Wang, B.C. Bonning, and W.A. Miller. 2009. The readthrough domain of pea enation mosaic virus coat protein is not essential for virus stability in the hemolymph of the pea aphid. Archives of Virology 154(3) 469–479.
Liu, S., S. Sivakumar, W.O. Sparks, W.A. Miller, and B.C. Bonning. 2010. A peptide that binds the pea aphid gut impedes entry of pea enation mosaic virus into the aphid hemocoel. Virology 401(1):107–116.
Liu, S.J., X.H. He, G. Park, C. Josefsson, and K.L. Perry. 2002. A conserved capsid protein surface domain of cucumber mosaic virus is essential for efficient aphid vector transmission. Journal of Virology 76(2) 9756–9762.
Lopez, C., J. Aramburu, L. Galipienso, S. Soler, F. Nuez, and L. Rubio. 2011. Evolutionary analysis of tomato Sw-5 resistance-breaking isolates of tomato spotted wilt virus. Journal of General Virology 92(1):210–215.
Lung, M.C.Y., and T.P. Pirone. 1974. Acquisition factor required for aphid transmission of purified cauliflower mosaic-virus. Virology 60:260–264.
Mar, T.T., L. Wenwen, and W. Xifeng. 2014. Proteomic analysis of interaction between P7-1 of Southern rice black-streaked dwarf virus and the insect vector reveals diverse insect proteins involved in successful transmission. Journal of Proteomics 102:83–97.
Martinez, J., B. Longdon, S. Bauer, Y. Chan, W.J. Miller, K. Bourtzis, L. Teixeira, and F.M. Jiggins. 2014. Symbionts commonly provide broad spectrum resistance to viruses in insects: A comparative analysis of Wolbachia strains. PLoS Pathogens 10:e1004369.
Massonnet-Bruneel, B., N. Corre-Catelin, R. Lacroix, R.S. Lees, Kim Phuc Hoang, D. Nimmo, L. Alphey, and P. Reiter. 2013. Fitness of transgenic mosquito Aedes aegypti males carrying a dominant lethal genetic system. PLoS One 8:e62711.
Montero-Astúa, M., D. Rotenberg, A. Leach-Kieffaber, B.A. Schneweis, S. Park, J.K. Park, T.L. German, and A.E. Whitfield. 2014. Disruption of vector transmission by a plant-expressed viral glycoprotein. Molecular Plant Microbe Interactions 27(3):296–304.
Moritz, G., S. Kumm, and L. Mound. 2004 Tospovirus transmission depends on thrips ontogeny. Virus Research 100(1):143–149.
Nagata, T., A.K. Inoue-Nagata, J. van Lent, R. Goldbach, and D. Peters. 2002. Factors determining vector competence and specificity for transmission of tomato spotted wilt virus. Journal of General Virology 83(3):663–671.
Ng, J., and B.W. Falk. 2006: Virus–vector interactions mediating nonpersistent and semipersistent transmission of plant viruses. Annual Review of Phytopathology 44:183–212.
Ohki, T., F. Akita, T. Mochizuki, A. Kanda, T. Sasaya, and S. Tsuda. 2010. The protruding domain of the coat protein of melon necrotic spot virus is involved in compatibility with and transmission by the fungal vector Olpidium bornovanus. Virology 402(1):129–134.
Peter, K.A., D. Liang, P. Palukaitis, and S.M. Gray. 2008. Small deletions in the potato leafroll virus readthrough protein affect particle morphology, aphid transmission, virus movement and accumulation. Journal of General Virology 89(8):2037–2045.
Pinheiro, P., A. Kliot, M. Ghanim, and M. Cilia. 2015. Is there a role for symbiotic bacteria in plant virus transmission by insects? Current Opinion in Insect Science 8:69–78.
Rotenberg, D., N.K.K. Kumar, D.E. Ullman, M. Montero-Astúa, D.K. Willis, T.L. German, and A.E. Whitfield. 2009. Variation in tomato spotted wilt virus titer in Frankliniella occidentalis and its association with frequency of transmission. Phytopathology 99:404–410.
Scott, J.G, K. Michel, L.C. Bartholomay, B.D. Siegfried, W.B. Hunter, G. Smagghe, K.Y. Zhu, and A.E. Douglas. 2013. Towards the elements of successful insect RNAi. Journal of Insect Physiology 59(12):1212–1221.
Seddas, P., S. Boissinot, J. Strub, A. Van Dorsselaer, M. Van Regenmortel, and F. Pattus. 2004. Rack-1, GAPDH3, and actin: Proteins of Myzus persicae potentially involved in the transcytosis of beet western yellows virus particles in the aphid. Virology 325(2):399–412.
Shao, C., J. Wu, G. Zhou, G. Sun, B. Peng, J. Lei, D. Jin, S. Chen, N. Upadhyaya, P. Waterhouse, and Z. Gong. 2003. Ectopic expression of the spike protein of Rice Ragged Stunt Oryzavirus in transgenic rice plants inhibits transmission of the virus to insects. Molecular Breeding 11:295–301.
Sin, S., B.C. McNulty, G.G. Kennedy, and J.W. Moyer 2005. Viral genetic determinants for thrips transmission of tomato spotted wilt virus. Proceedings of the National Academy of Sciences USA 102:5168–5173.
Tamborindeguy, C., M.S. Bereman, S. DeBlasio, D. Igwe, D.M. Smith, F. White, M.J. MacCoss, S.M. Gray, and M. Cilia. 2013. Genomic and proteomic analysis of Schizaphis graminum reveals cyclophilin proteins are involved in the transmission of cereal yellow dwarf virus. PLoS One 8(8):e71620.
Uzest, M., D. Gargani, A. Dombrovsky, C. Cazevieille, D. Cot, and S. Blanc. 2010. The “acrostyle”: A newly described anatomical structure in aphid stylets. Arthropod Structure and Development 39(4):221–229.
Uzest, M., D. Gargani, M. Drucker, E. Hébrard, E. Garzo, T. Candresse, A. Fereres, and S. Blanc. 2007. A protein key to plant virus transmission at the tip of the insect vector stylet. Proceedings of the National Academy of Sciences USA 104:17959–17964.
van den Hurk, A.F., S. Hall-Mendelin, A.T. Pyke, F.D. Frentiu, K. McElroy, A. Day, S. Higgs, and S.L. O’Neill. 2012. Impact of Wolbachia on infection with chikungunya and yellow fever viruses in the mosquito vector Aedes aegypti. PLoS Neglected Tropical Diseases 6(11):e18926.
Wang, L., X. Wei, X. Ye, H. Xu, X. Zhou, S. Liu, and X. Wang. 2014. Expression and functional characterisation of a soluble form of tomato yellow leaf curl virus coat protein. Pest Management Sciences 70(10):1624–1631.
Wang, R.Y., E.D. Ammar, D.W. Thornbury, J.J. Lopez-Moya, and T.P. Pirone. 1996. Loss of potyvirus transmissibility and helper-component activity correlate with non-retention of virions in aphid stylets. Journal General Virology 77(5):861–867.
Webster, C.G., S.R. Reitz, K.L. Perry, and S. Adkins. 2011. A natural M RNA reassortant arising from two species of plant- and insect-infecting bunyaviruses and comparison of its sequence and biological properties to parental species. Virology 413:216–225.
Whitfield, A.E., D.E. Ullman, and T.L. German. 2004. Expression, purification, and characterization of a soluble form of tomato spotted wilt virus glycoprotein GN. Journal of Virology 78(23):13197–13206.
Whitfield, A.E., D.E. Ullman, and T.L. German. 2005. Tospovirus–thrips interactions. Annual Review of Phytopathology 43 459–489.
Whitfield, A.E., N.K.K. Kumar, D. Rotenberg, D.E. Ullman, E.A. Wyman, C. Zietlow, D.K. Willis, and T.L. German. 2008. A soluble form of the tomato spotted wilt virus (TSWV) glycoprotein GN (GN-S) inhibits transmission of TSWV by Frankliniella occidentalis. Phytopathology 98(1):45–50.
Winston, W.M., C. Molodowitch, and C.P. Hunter. 2002: Systemic RNAi in C. elegans requires the putative transmembrane protein SID-1. Science 295(5565):2456–2459.
Yang, X., T.W. Thannhauser, M. Burrows, D. Cox-Foster, F.E. Gildow, and S.M. Gray. 2008. Coupling genetics and proteomics to identify aphid proteins associated with vector-specific transmission of polerovirus (Luteoviridae). Journal of Virology 82(1):291–299.
Yao, J., D. Rotenberg, A. Afsharifar, K. Barandoc-Alviar, and A.E. Whitfield. 2013. Development of RNAi methods for Peregrinus maidis, the corn planthopper. PLoS One 8(8):e70243.
Zhang, F., H. Guo, Z. Zheng, T. Zhou, Y. Zhou, S. Wang, R. Fang, W. Qian, and X. Chen. 2010. Parallel pyrosequencing-based transcriptome analyses of small brown planthopper (Laodelphax striatellus), a vector insect transmitting rice stripe virus (RSV). BMC Genomics 11:303.
Zhou, J., S. Kantartzi, R- Wen, M. Newman, M. Hajimorad, J. Rupe, and I. Tzanetakis. 2011. Molecular characterization of a new tospovirus infecting soybean. Virus Genes 43:289–295.
This page intentionally left blank.


















