10
Engineering the Microbiome for Human Health Applications
Timothy K. Lu,a,b,c,d,* Mark Mimee,a,bRobert J. Citorik,a,band Karen Pepperc
INTRODUCTION
The importance of the microbiome for human health and disease has become increasingly clear over the last decade. Metabolism, immunity, and the gut–brain axis are affected by the intimate associations between host and the microbiota, with interactions occurring across multiple body sites: the nasopharynx, oral cavity, respiratory tract, gastrointestinal tract, female reproductive tract, and skin. This new understanding has motivated the development of microbiome-based therapeutics to treat diseases linked to these diverse microbial communities. For example, the genetic engineering of microbes, including natural members of the microbiota, has enabled the design of microorganisms that sense and treat disease. Beyond individual bacteria, increasing interest has been placed on the study of microbial consortia, interactions between host and microbe, the role of viruses, and the modulation of these processes for therapeutic applications. Despite significant progress in recombinant probiotics, therapeutic microbial consortia, and targeted antimicrobials, translation into clinical applications still faces numerous challenges and unknowns. Here, we discuss recent research opportunities for impacting human health through the microbiome, and potential roadblocks for microbiome-based therapeutics. This work is adapted from our more extensive review on this topic (Mimee et al., 2016).
HARNESSING AND ENGINEERING THE MICROBIOME
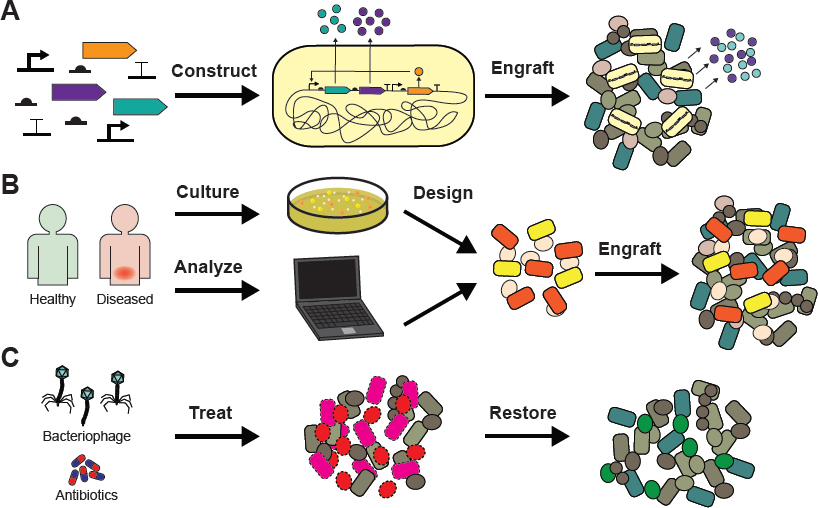
Microbiome-based therapeutics, designed to improve human health by altering the associated microbial communities, may employ modulatory, additive, or subtractive approaches. Modulatory therapies involve altering the composition or activity of the endogenous microbiota via the administration of nonliving agents or prebiotics (for a review of prebiotics, see Frei et al., 2015). Additive therapies supplement the microbiota with natural or engineered microorganisms (de Moreno de LeBlanc and LeBlanc, 2014; Derrien and van Hylckama Vlieg, 2015; Varankovich et al., 2015; Marchesi et al., 2016; see Figures 10-1A and 10-1B), given either individually or as
___________________
a Microbiology Program, Massachusetts Institute of Technology.
b Synthetic Biology Center, Massachusetts Institute of Technology.
c Department of Biological Engineering, Massachusetts Institute of Technology.
d Department of Electrical Engineering and Computer Science, Massachusetts Institute of Technology.
* Corresponding Author: timlu@mit.edu.
SOURCE: Reprinted from Advanced Drug Delivery Reviews, 105, Mark Mimee, Robert J. Citorik, and Timothy K. Lu, “Microbiome therapeutics—Advances and challenges,” 44-54, Copyright 2016, with permission from Elsevier.
collections of strains. Subtractive therapies aim to modulate host interactions by eliminating specific members of the microbiome (see Figure 10-1C). In the future, additive and subtractive approaches may be used together to achieve greater effects on the microbiome.
Additive Approaches
Numerous health benefits have been attributed to natural, human-associated microbes. Lactobacillus spp., Escherichia coli, and Bifidobacterium spp. have the potential to treat a variety of diseases (Ritchie and Romanuk, 2012; Fujiya et al., 2014; Cuello-Garcia et al., 2015; Zuccotti et al., 2015) and, indeed, can be found in over-the-counter probiotics. The recombinant expression of therapeutic biomolecules from engineered microbes may increase these benefits and help prevent infection, resolve inflammation, and treat metabolic disorders. Bacteria could be developed to deliver drugs at the site of disease, enhancing bioavailability and reducing drug inactivation. Furthermore, these bacteria could be outfitted with sensors that detect disease biomarkers and trigger on-demand drug release. Fully autonomous, “smart” cell-based therapeutics for restoring the health of a human host have not yet been advanced into the clinic, but the requisite technologies are available. Below, we discuss examples of microbes being used, either individually or as consortia, to treat disease. A major challenge in creating microbiota-based therapeutics is the identification and customization of bacterial communities to address complex human diseases, despite the diversity of human-associated microbiota.
One application of engineered bacteria is to treat bacterial and viral infections. The normal flora present in healthy individuals can resist host colonization by pathogens, and cellular engineering can augment such resistance. E. coli Nissle 1917, a probiotic strain, has been designed to inhibit the virulence of Vibrio cholerae within infant mouse models (Duan and March, 2010). V. cholerae depends on quorum sensing to coordinate the expression of certain virulence factors with cell density. The administration of E. coli engineered to interfere with this quorum sensing system resulted in the increased survival of infected mice, along with a decreased bacterial burden and cholera toxin expression. In another example, genetically modified Lactobacillus jensenii prevented transmission of chimeric simian/human immunodeficiency virus (SHIV) in a rhesus macaque monkey model. Bacteria were modified to express cyanovirin-N, an antiviral molecule, and decreased both the occurrence of SHIV and peak viral load when administered as a prophylactic treatment (Lagenaur et al., 2011).
Fecal microbiota transplant—consisting of stool derived from healthy donors then infused to diseased patients—has greater than 90% efficacy in resolving recurrent Clostridium difficile infections (Kassam et al., 2013), which is greater than antibiotic treatment alone (van Nood et al., 2013). Safety concerns about introducing pathogens and exacerbating disease have led to a regulatory framework and stringent donor screening guidelines. Determining the minimal subset of microbes needed to achieve therapeutic efficacy may mitigate safety concerns and increase treatment reliability (Petrof et al., 2013).
Fecal microbiota transplants may prove effective for treating inflammatory bowel disease (IBD) (Ratner, 2015); already, early trials have shown some success (Ianiro et al., 2014; Moayyedi et al., 2015). In addition, recombinant bacterial therapies may provide cheaper and less invasive treatments for chronic inflammatory diseases (Wlodarska et al., 2015). Lactococcus lactis has been engineered to secrete interleukin-10, an important anti-inflammatory cytokine, reducing pathology and suppressing pro-inflammatory cytokine secretion in mouse models of colitis (Steidler et al., 2000). Microbial expression of other anti-inflammatory cytokines—such as transforming growth factor-β1 (Hamady et al., 2011), antitumor necrosis factor α nanobodies (Vandenbroucke et al., 2010), and the tissue repair factor keratinocyte growth factor-2 (Hamady et al., 2010)—protected against colitis in mouse models of IBD. In addition to cytokines, the protease inhibitor Elafin, when produced by lactic acid bacteria, restored the proteolytic homeostasis disrupted in mouse colitis models and protected against inflammation (Motta et al., 2012). Despite these preclinical studies, these approaches have yet to show efficacy in humans, perhaps due to the challenge of expressing therapeutic molecules in the right place at the right time and at high enough levels to be effective.
Metabolic diseases, such as obesity and diabetes, are also being addressed by delivering engineered microbes into the host microbiota. E. coli designed to synthesize precursors of appetite-suppressing lipids reduced obesity in mice fed a high-fat diet, and effects lasted weeks after bacterial treatment ended (Chen et al., 2014). GLP-1, a protein that induces the conversion of intestinal epithelial cells into insulin-producing cells, expanded the numbers of insulin-producing intestinal cells and reduced hyperglycemia when delivered by Lactobacillus gasseri in a rat model (Duan et al., 2015).
Hyperammonemia is another metabolic condition for which engineering the microbiota may prove effective. In the gut, bacterial ureases convert urea made by the liver to ammonia and carbon dioxide. Hyperammonemia occurs when too much ammonia accumulates systemically, and leads to neurotoxicity and encephalopathy in people with liver disease. In mouse models, reconstituting the microbiota altered community-wide urea metabolism (Shen et al., 2015). When the endogenous microbiota was depleted and a defined microbial community exhibiting low urease activity was transplanted, urease levels remained stable for months (Shen et al., 2015). The redefined microbiota enhanced survival and reduced cognitive defects associated with hyperammonemia in a hepatic injury model. Thus, modifying an existing microbial community can protect against metabolic diseases. Furthermore, microbes have been genetically engineered to degrade ammonia and shown to reduce systemic ammonia levels when fed to mice (Nicaise et al., 2008). Such therapies are currently being developed by companies for clinical trials (Synlogic, 2017).
Subtractive Approaches
Subtractive therapies aim to eliminate deleterious members of the microbiome (see Figure 10-2C) using mechanisms such as antibiotics, chemicals, peptides, and bacteriophages. Antibiotics, a key example of subtractive therapies, often have the undesirable effect of killing a broad set of microbes outside of the desired target. This can result in severe side effects, such as increased susceptibility to bacterial pathogens, including Clostridium difficile. Future subtractive therapies for the microbiome should be much more specific in targeting activity.
One strategy for highly specific subtractive therapies uses phages, which are natural viral parasites that infect bacteria, often killing the bacterial host in the process of producing phage progeny. The growing threat of antibiotic-resistant pathogens has rekindled interest in phage therapy (Reardon, 2014; Kingwell, 2015), especially considering that phages often specifically attack only one or a few cell types of bacteria and, thus, could be employed as more targeted antimicrobials.
Phages naturally shape host-associated bacterial populations (Mills et al., 2013). Metagenomic studies of the fecal virome of healthy and diseased people have revealed phage diversity, variability, and stability (Reyes et al., 2010), as well as changes associated with diet (Minot et al., 2011), IBD (Norman et al., 2015), or antibiotic treatment (Modi et al., 2013). High interpersonal variation in the composition of the viral community, but low intrapersonal diversity dominated by temperate, potentially dormant, phages was seen in a study of monozygotic twins and their mothers (Reyes et al., 2010). Diet can affect both the bacterial population of the gut and the viral community; individuals on the same diet displayed convergence in the phages they carried (Minot et al., 2011). IBD can also coincide with changes in both populations, as reduced bacterial diversity was found alongside increased bacteriophage richness (Norman et al., 2015). Germ-free mice were seeded with a defined, 15-member commensal community from humans and then challenged orally with virus-like particles from healthy donors. In these mice,
SOURCE: Reprinted from Advanced Drug Delivery Reviews, 105, Mark Mimee, Robert J. Citorik, and Timothy K. Lu, “Microbiome therapeutics—Advances and challenges,” 44-54, Copyright 2016, with permission from Elsevier.
an increase in specific phages correlated with a transient decrease in specific bacteria. Some phages and bacteria exhibited fluctuating population dynamics, and a potentially critical observation was that phage resistance seemed due to ecological factors rather than genetic ones (Reyes et al., 2013).
In addition to using natural phage isolates, phages can be modified to carry extra or alternative functions to expand their utility. Immunoglobulin-like protein domains on the capsids of certain phages’ exterior enhance association with mucus (Barr et al., 2013), a mechanism that could potentially be used to localize phage to particular parts of the body or to extend residence time in the gut. Host range can be reprogrammed to alter the bacterial targets (Ando et al., 2015) and genes can be inserted to improve the killing of biofilms (Lu and Collins, 2007). Additionally, phages have been used to deliver DNA to bacteria that reverses antibiotic resistance (Lu and Collins, 2009; Edgar et al., 2012) or to achieve nonspecific (Westwater et al., 2003; Hagens et al., 2004; Krom et al., 2015) or sequence-specific (Bikard et al., 2014; Citorik et al., 2014) antimicrobial activity toward targeted cells. New tools such as CRISPR-Cas (Kiro et al., 2014) genome editing and construction methods, including Gibson (Gibson et al., 2009) and yeast (Ando et al., 2015) assembly, will facilitate future engineering efforts. Phages as therapeutics for microbiota-related diseases represent a promising area of investigation, and using them as tools to alter microbial communities could enable systematic probing of these populations for discovery and validation in the study of health and the microbiome.
OUTLOOK FOR MICROBIOTA-BASED THERAPEUTICS
The development of microbiota-based therapeutics has been accelerated by progress in synthetic biology and our understanding of host-associated microbial consortia. However, numerous challenges arise in bringing this work to the clinic. Many advances in microbiome therapeutics have been validated using rodent models, but the ability to generalize these findings to humans has yet to be comprehensively tested. In addition, the development of fully autonomous cellular therapies requires biosensors that are clinically relevant biosensors and genetic circuits that are robust. Finally, the translation of basic research to clinical applications depends on setting up regulatory frameworks to address unique issues with living therapeutics.
Stable Engraftment
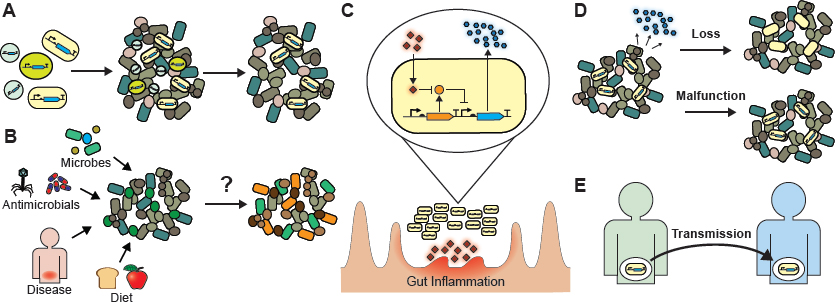
Various organisms can provide chassis for cell-based therapies but may have differential effects based on their affinity for specific environments, their ability to engraft, and their inherent biological effects (see Figure 10-2A). Thorough characterization of a species’ suitability for a given disease will help to determine the choice of microbial chassis. Chassis currently used for cell-based therapies include E. coli Nissle 1917 (Duan et al., 2010; Vandenbroucke et al., 2010), L. lactis (Braat et al., 2006; Vandenbroucke et al., 2010; Takiishi et al., 2012; Limaye et al., 2013), Lactobacillus spp. (Lagenaur et al., 2011; Motta et al., 2012; Duan et al., 2015), and Bacteroides spp. (Hamady et al., 2010, 2011; Mimee et al., 2015). Bacteroides spp. live in the cecum and colon while E. coli and Lactobacillus spp. are enriched in the small intestine (Donaldson et al., 2016). Some species preferably colonize the intestinal lumen while others live in the mucosal layer (Nava et al., 2011; Earle et al., 2015; Li et al., 2015). Choosing the organism best suited for therapy will depend on the biogeography of disease.
Stable colonization of recombinant microbes or microbial consortia may not be necessary if bacterial cells can enact their intended therapeutic functions while they transit through the intestine. For instance, L. lactis, which does not colonize the mammalian intestine, is serving as a chassis for therapeutic protein production (Braat et al., 2006; Vandenbroucke et al., 2010; Hamady et al., 2011; Takiishi et al., 2012; Limaye et al., 2013; Robert et al., 2014). E. coli Nissle 1917, another commonly used chassis, shows great variability in colonization capacity: Less than 50% of volunteers became decolonized 2 weeks after treatment was stopped, whereas, after 6 months, the probiotic was detected by polymerase chain reaction in the stool of only 17.5% of volunteers (Joeres-Nguyen-Xuan et al., 2010).
Nevertheless, stable colonization of bacterial therapies into the endogenous microbiota has the potential to improve treatment efficacy and allow for long-term and fully autonomous therapies that sense and respond to a given disease state. To develop long-term cell-based therapies, invasion, resilience, and succession mechanisms
in host-associated microbial ecosystems must be understood (see Figure 10-2B). Long-term therapies may require organisms that are naturally resilient to environmental perturbations and abundant in the environments of interest, such as Bacteroides spp. Pairing additive approaches with subtractive or modulatory ones could improve the engraftment of strains into the microbiome. Bacteriophages and other targeted antimicrobials could make way for therapeutic microbes by eliminating incompatible partners, and dietary supplementation with prebiotics could be used to introduce new members of the microbiota.
Some of these species may confer additional health benefits. For example, Faecalibacterium prausnitzii, B. fragilis, and bacteria from Clostridium clusters IV and XIVa naturally protect against inflammation (Mazmanian et al., 2008; Sokol et al., 2008; Atarashi et al., 2011), whereas E. coli is enriched in an inflamed gut (Gevers et al., 2014). Furthermore, new methods for engineering currently intractable organisms would increase the range of possibilities for cell-based therapies. Recent work has extended genetic tools to Bacteroides spp. (Mimee et al., 2015) in addition to those already in place for E. coli and lactic acid bacteria. Efficient genetic techniques for manipulating group IV and XIVa clostridia and F. prausnitzii would accelerate progress.
Development of Clinically Relevant Sensors
A well-characterized library of biosensors that dynamically respond to environmental perturbations is needed for the further development of autonomous cell-based therapies (see Figure 10-2C). Synthetic biologists are developing a range of genetic parts to sense environmental signals and regulate gene expression. Biosensors with luminescent, fluorescent, or colorimetric outputs can be transiently transcriptionally regulated or permanently coupled to genomic alterations (Bonnet et al., 2013; Siuti et al., 2013; Farzadfard and Lu, 2014; Mimee et al., 2015). Biosensors have been found by mining genome databases and the scientific literature. However, the next generation of novel biosensors can be developed by directed evolution, as has been achieved with enzymatic substrate specificity (Ellefson et al., 2014) or the promoter specificity of RNA polymerases (Esvelt et al., 2011). DNA-binding and ligand-binding domains have been incorporated into hybrid transcription factors, expanding the variety of available sensors (Chou and Keasling, 2013; Shis et al., 2014; Chan et al., 2016).
Further work in this area awaits a generalized approach for the de novo discovery of sensors for clinically relevant biomarkers. Biosensor discovery paired with metabolomic studies could be used to assay biomarker concentrations inside the body rather than in ex vivo samples. Engineered microbes could thus provide a new class of diagnostics. The localized production of medicines could also be set in motion by these sensors, as needed to treat disease on demand.
Relevance, Robustness, and Stability of Genetic Circuits
The genetic circuits needed to implement sense-and-respond bacterial therapeutics are usually prototyped in optimal in vitro growth conditions, but once inside the body, they may behave differently. Cellular therapies may not last sufficiently long or may not withstand changing environments (see Figure 10-2D). More sophisticated in vitro systems reflecting the conditions of the endogenous microbiota are needed, particularly ones that can sustain host cells together with multispecies bacterial communities. Single (McDonald et al., 2013; Auchtung et al., 2015) and multistage (Van den Abbeele et al., 2013) chemostats used to culture fecal samples could help elucidate the impact of interbacterial interactions on genetic circuits. Organoid (Lukovac et al., 2014), three-dimensional intestinal scaffolds (Costello et al., 2014), and gut-on-a-chip (Kim et al., 2016) models could be used to predict interactions between the host and bacteria.
Long-term therapeutics pose challenges because gene circuit function is generally assessed on time scales of less than 24 hours in vitro, whereas cellular therapies may need to operate for weeks to months, which may not be possible if mutations occur that inactivate the desired behaviors (Ceroni et al., 2015). In addition, in vitro evolution experiments have revealed that engineered bacteriophages may lose their function over time (Gladstone et al., 2012; Springman et al., 2012). Thus, strategies to sustain the activity of therapeutics within the competitive microbiota environment are of major importance (Ceroni et al., 2015).
Regulation, Safety, and Biocontainment
A regulatory framework that can be used to address the safety and biocontainment issues of cell-based therapies should be established to guide the field toward real-world applications (see Figure 10-2E). Existing probiotics and bacteria already employed in food production are classified by the U.S. Food and Drug Administration as generally safe organisms. The safety of other organisms proposed for microbiota-based therapies—including natural commensal organisms such as clostridial or Bacteroides species—needs to be evaluated in well-controlled clinical trials. Furthermore, the capability of these bacteria to stably colonize host-associated environments poses unique challenges for modeling the pharmacokinetics and pharmacodynamics of their therapeutic effects. Another potential question is the extent to which DNA will be transferred between recombinant and natural organisms. Strategies for recoding the genetic code may enhance the isolation of heterologous genetic constructs from natural systems (Lajoie et al., 2013).
Finally, most genetically modified organisms created in the laboratory are less fit than the wild type strain from which they were derived (Ceroni et al., 2015), so even if they escape their specific environments, their engineered functions may be degraded over time. To further contain genetically modified constructs, they could be eliminated through DNA degradation devices (Caliando and Voigt, 2015) or kill switches (Wright et al., 2015; Chan et al., 2016). The dissemination of recombinant cells can also be reduced by using auxotrophic microbes that do not replicate in the absence of a specific chemical (Steidler et al., 2003); auxotrophy has been used for biocontainment in early clinical trials of recombinant microbes (Braat et al., 2006; Limaye et al., 2013). Auxotrophs that depend on synthetic chemicals, rather than natural chemicals, may further enhance the biocontainment of these strains (Lajoie et al., 2013; Ostrov et al., 2016).
CONCLUSIONS
Therapeutics targeting the human microbiome are undergoing rapid development and attracting broad interest due to their potential benefits. Current additive and subtractive strategies to manipulate the human microbiome include engineering bacteria to produce therapeutic molecules, constituting natural or artificial consortia to modulate the host, and applying selective antimicrobials. Challenges in creating microbiome therapeutics include engineering microbial therapies that are well adapted to specific environments in the body or able to achieve stable colonization, discovering or constructing clinically relevant biosensors, engineering robust and effective synthetic gene circuits that can function in vivo, and establishing regulatory frameworks to account for safety and biocontainment concerns in addition to therapeutic efficacy. Given the deep interactions between host and microbe that are being uncovered, we envision that various approaches to engineering the microbiome have the potential to transform the treatment of challenging human diseases.
ACKNOWLEDGMENTS
This work was supported by the Center for Microbiome Informatics and Therapeutics, the Defense Advanced Research Projects Agency, the National Institutes of Health (DP2 OD008435, P50 GM098792, R01 EB017755), the Office of Naval Research (N00014-13-1-0424), the National Science Foundation (MCB-1350625), and the Defense Threat Reduction Agency (HDTRA1-14-1-0007, HDTRA1-15-1-0050). M.M. is a Howard Hughes Medical Institute Student Research fellow. T.K.L. is a co-founder of Synlogic and Eligo Biosciences, which are pursuing some of the approaches described here.
REFERENCES
Ando, H., S. Lemire, D. P. Pires, and T. K. Lu. 2015. Engineering modular viral scaffolds for targeted bacterial population editing. Cell Syst (3):187-196. doi: 10.1016/j.cels.2015.08.013.
Atarashi, K., T. Tanoue, T. Shima, A. Imaoka, T. Kuwahara, Y. Momose, G. Cheng, S. Yamasaki, T. Saito, Y. Ohba, T. Taniguchi, K. Takeda, S. Hori, I. I. Ivanov, Y. Umesaki, K. Itoh, and K. Honda. 2011. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331(6015):337-341. doi:10.1126/science.1198469.
Auchtung, J. M., C. D. Robinson, and R. A. Britton. 2015. Cultivation of stable, reproducible microbial communities from different fecal donors using minibioreactor arrays (MBRAs). Microbiome 3:42. doi: 10.1186/s40168-015-0106-5.
Barr, J. J., R. Auro, M. Furlan, K. L. Whiteson, M. L. Erb, J. Pogliano, A. Stotland, R. Wolkowicz, A. S. Cuttin, K. S. Doran, P. Salamon, M. Youle, and F. Rohwer. 2013. Bacteriophage adhering to mucus provide a non–host-derived immunity. Proc Natl Acad Sci USA 110(26):10771-10776. doi: 10.1073/pnas.1305923110.
Bikard, D., C. W. Euler, W. Jiang, P. M. Nussenzweig, G. W. Goldberg, X. Duportet, V. A. Fischetti, and L. A. Marraffini. 2014. Exploiting CRISPR-Cas nucleases to produce sequence-specific antimicrobials. Nat Biotechnol 32(11):1146-1150. doi: 10.1038/nbt.3043.
Bonnet, J., P. Yin, M. E. Ortiz, P. Subsoontorn, and D. Endy. 2013. Amplifying genetic logic gates. Science 340(6132):599-603. doi: 10.1126/science.1232758.
Braat, H., P. Rottiers, D. W. Hommes, N. Huyghebaert, E. Remaut, J. P. Remon, S. J. van Deventer, S. Neirynck, M. P. Peppelenbosch, and L. Steidler. 2006. A phase I trial with transgenic bacteria expressing interleukin-10 in Crohn’s disease. Clin Gastroenterol Hepatol 4(6):754-759. doi: 10.1016/j.cgh.2006.03.028.
Caliando, B. J., and C. A. Voigt. 2015. Targeted DNA degradation using a CRISPR device stably carried in the host genome. Nat Commun 6:6989. doi: 10.1038/ncomms7989.
Ceroni, F., R. Algar, G. B. Stan, and T. Ellis. 2015. Quantifying cellular capacity identifies gene expression designs with reduced burden. Nat Methods 12(5):415-418. doi: 10.1038/nmeth.3339.
Chan, C. T., J. W. Lee, D. E. Cameron, C. J. Bashor, and J. J. Collins. 2016. “Deadman” and “Passcode” microbial kill switches for bacterial containment. Nat Chem Biol 12(2):82-86. doi: 10.1038/nchembio.1979.
Chen, Z., L. Guo, Y. Zhang, R. L. Walzem, J. S. Pendergast, R. L. Printz, L. C. Morris, E. Matafonova, X. Stien, L. Kang, D. Coulon, O. P. McGuinness, K. D. Niswender, and S. S. Davies. 2014. Incorporation of therapeutically modified bacteria into gut microbiota inhibits obesity. J Clin Invest 124(8):3391-3406. doi: 10.1172/JCI72517.
Chou, H. H., and J. D. Keasling. 2013. Programming adaptive control to evolve increased metabolite production. Nat Commun 4:2595. doi: 10.1038/ncomms3595.
Citorik, R. J., M. Mimee, and T. K. Lu. 2014. Sequence-specific antimicrobials using efficiently delivered RNA-guided nucleases. Nat Biotechnol 32(11):1141-1145. doi: 10.1038/nbt.3011.
Costello, C. M., R. M. Sorna, Y. L. Goh, I. Cengic, N. K. Jain, and J. C. March. 2014. 3-D intestinal scaffolds for evaluating the therapeutic potential of probiotics. Mol Pharm 11(7):2030-2039. doi: 10.1021/mp5001422.
Cuello-Garcia, C. A., J. L. Brozek, A. Fiocchi, R. Pawankar, J. J. Yepes-Nunez, L. Terracciano, S. Gandhi, A. Agarwal, Y. Zhang, and H. J. Schunemann. 2015. Probiotics for the prevention of allergy: A systematic review and meta-analysis of randomized controlled trials. J Allergy Clin Immunol 136(4):952-961. doi: 10.1016/j.jaci.2015.04.031.
de Moreno de LeBlanc, A., and J. G. LeBlanc. 2014. Effect of probiotic administration on the intestinal microbiota, current knowledge and potential applications. World J Gastroenterol 20(44):16518-16528. doi: 10.3748/wjg.v20.i44.16518.
Derrien, M., and J. E. van Hylckama Vlieg. 2015. Fate, activity, and impact of ingested bacteria within the human gut microbiota. Trends Microbiol 23(6):354-366. doi:10.1016/j.tim.2015.03.002.
Donaldson, G. P., S. M. Lee, and S. K. Mazmanian. 2016. Gut biogeography of the bacterial microbiota. Nat Rev Microbiol 14(1):20-32. doi: 10.1038/nrmicro3552.
Duan, F., and J. C. March. 2010. Engineered bacterial communication prevents Vibrio cholerae virulence in an infant mouse model. Proc Natl Acad Sci USA 107(25):11260-11264. doi:10.1073/pnas.1001294107.
Duan, F. F., J. H. Liu, and J. C. March. 2015. Engineered commensal bacteria reprogram intestinal cells into glucose-responsive insulin-secreting cells for the treatment of diabetes. Diabetes 64(5):1794-1803. doi: 10.2337/db14-0635.
Earle, K. A., G. Billings, M. Sigal, J. S. Lichtman, G. C. Hansson, J. E. Elias, M. R. Amieva, K. C. Huang, and J. L. Sonnenburg. 2015. Quantitative imaging of gut microbiota spatial organization. Cell Host Microbe 18(4):478-488. doi: 10.1016/j. chom.2015.09.002.
Edgar, R., N. Friedman, S. Molshanski-Mor, and U. Qimron. 2012. Reversing bacterial resistance to antibiotics by phage-mediated delivery of dominant sensitive genes. Appl Environ Microbiol 78(3):744-751. doi: 10.1128/AEM.05741-11.
Ellefson, J. W., A. J. Meyer, R. A. Hughes, J. R. Cannon, J. S. Brodbelt, and A. D. Ellington. 2014. Directed evolution of genetic parts and circuits by compartmentalized partnered replication. Nat Biotechnol 32(1):97-101. doi: 10.1038/nbt.2714.
Esvelt, K. M., J. C. Carlson, and D. R. Liu. 2011. A system for the continuous directed evolution of biomolecules. Nature 472(7344):499-503. doi: 10.1038/nature09929.
Farzadfard, F., and T. K. Lu. 2014. Synthetic biology. Genomically encoded analog memory with precise in vivo DNA writing in living cell populations. Science 346(6211):1256272. doi: 10.1126/science.1256272.
Frei, R., M. Akdis, and L. O’Mahony. 2015. Prebiotics, probiotics, synbiotics, and the immune system: Experimental data and clinical evidence. Curr Opin Gastroenterol 31(2):153-158. doi:10.1097/MOG.0000000000000151.
Fujiya, M., N. Ueno, and Y. Kohgo. 2014. Probiotic treatments for induction and maintenance of remission in inflammatory bowel diseases: A meta-analysis of randomized controlled trials. Clin J Gastroenterol 7(1):1-13. doi: 10.1007/s12328-013-0440-8.
Gevers, D., S. Kugathasan, L. A. Denson, Y. Vazquez-Baeza, W. Van Treuren, B. Ren, E. Schwager, D. Knights, S. J. Song, M. Yassour, X. C. Morgan, A. D. Kostic, C. Luo, A. Gonzalez, D. McDonald, Y. Haberman, T. Walters, S. Baker, J. Rosh, M. Stephens, M. Heyman, J. Markowitz, R. Baldassano, A. Griffiths, F. Sylvester, D. Mack, S. Kim, W. Crandall, J. Hyams, C. Huttenhower, R. Knight, and R. J. Xavier. 2014. The treatment-naïve microbiome in new-onset Crohn’s disease. Cell Host Microbe 15(3):382-392. doi: 10.1016/j.chom.2014.02.005.
Gibson, D. G., L. Young, R. Y. Chuang, J. C. Venter, C. A. Hutchison, 3rd, and H. O. Smith. 2009. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6(5):343-345. doi:10.1038/nmeth.1318.
Gladstone, E. G., I. J. Molineux, and J. J. Bull. 2012. Evolutionary principles and synthetic biology: Avoiding a molecular tragedy of the commons with an engineered phage. J Biol Eng 6(1):13. doi:10.1186/1754-1611-6-13.
Hagens, S., A. Habel, U. von Ahsen, A. von Gabain, and U. Blasi. 2004. Therapy of experimental pseudomonas infections with a nonreplicating genetically modified phage. Antimicrob Agents Chemother 48(10):3817-3822. doi: 10.1128/AAC.48.10.3817-3822.2004.
Hamady, Z. Z., N. Scott, M. D. Farrar, J. P. Lodge, K. T. Holland, T. Whitehead, and S. R. Carding. 2010. Xylan-regulated delivery of human keratinocyte growth factor-2 to the inflamed colon by the human anaerobic commensal bacterium Bacteroides ovatus. Gut 59(4):461-469. doi: 10.1136/gut.2008.176131.
Hamady, Z. Z., N. Scott, M. D. Farrar, M. Wadhwa, P. Dilger, T. R. Whitehead, R. Thorpe, K. T. Holland, J. P. Lodge, and S. R. Carding. 2011. Treatment of colitis with a commensal gut bacterium engineered to secrete human TGF-beta1 under the control of dietary xylan 1. Inflamm Bowel Dis 17(9):1925-1935. doi: 10.1002/ibd.21565.
Ianiro, G., S. Bibbo, F. Scaldaferri, A. Gasbarrini, and G. Cammarota. 2014. Fecal microbiota transplantation in inflammatory bowel disease: Beyond the excitement. Medicine (Baltimore). 93(19):e97. doi: 10.1097/MD.0000000000000097.
Joeres-Nguyen-Xuan, T. H., S. K. Boehm, L. Joeres, J. Schulze, and W. Kruis. 2010. Survival of the probiotic Escherichia coli Nissle 1917 (EcN) in the gastrointestinal tract given in combination with oral mesalamine to healthy volunteers. Inflamm Bowel Dis 16(2):256-262. doi: 10.1002/ibd.21042.
Kassam, Z., C. H. Lee, Y. Yuan, and R. H. Hunt. 2013. Fecal microbiota transplantation for Clostridium difficile infection: Systematic review and meta-analysis. Am J Gastroenterol 108(4):500-508. doi: 10.1038/ajg.2013.59.
Kim, H. J., H. Li, J. J. Collins, and D. E. Ingber. 2016. Contributions of microbiome and mechanical deformation to intestinal bacterial overgrowth and inflammation in a human gut-on-a-chip. Proc Natl Acad Sci USA 113(1):E7-E15. doi: 10.1073/pnas.1522193112.
Kingwell, K. 2015. Bacteriophage therapies re-enter clinical trials. Nat Rev Drug Discov 14(8):515-516. doi: 10.1038/nrd4695.
Kiro, R., D. Shitrit, and U. Qimron. 2014. Efficient engineering of a bacteriophage genome using the type I-E CRISPR-Cas system. RNA Biol 11(1):42-44. doi: 10.4161/rna.27766.
Krom, R. J., P. Bhargava, M. A. Lobritz, and J. J. Collins. 2015. Engineered phagemids for nonlytic, targeted antibacterial therapies. Nano Lett 15(7):4808-4813. doi: 10.1021/acs.nanolett.5b01943.
Lagenaur, L. A., B. E. Sanders-Beer, B. Brichacek, R. Pal, X. Liu, Y. Liu, R. Yu, D. Venzon, P. P. Lee, and D. H. Hamer. 2011. Prevention of vaginal SHIV transmission in macaques by a live recombinant Lactobacillus. Mucosal Immunol 4(6):648-657. doi: 10.1038/mi.2011.30.
Lajoie, M. J., A. J. Rovner, D. B. Goodman, H. R. Aerni, A. D. Haimovich, G. Kuznetsov, J. A. Mercer, H. H. Wang, P. A. Carr, J. A. Mosberg, N. Rohland, P. G. Schultz, J. M. Jacobson, J. Rinehart, G. M. Church, and F. J. Isaacs. 2013. Genomically recoded organisms expand biological functions. Science 342(6156):357-360. doi:10.1126/science.1241459.
Li, H., J. P. Limenitakis, T. Fuhrer, M. B. Geuking, M. A. Lawson, M. Wyss, S. Brugiroux, I. Keller, J. A. Macpherson, S. Rupp, B. Stolp, J. V. Stein, B. Stecher, U. Sauer, K. D. McCoy, and A. J. Macpherson. 2015. The outer mucus layer hosts a distinct intestinal microbial niche. Nat Commun 6:8292. doi:10.1038/ncomms9292.
Limaye, S. A., R. I. Haddad, F. Cilli, S. T. Sonis, A. D. Colevas, M. T. Brennan, K. S. Hu, and B. A. Murphy. 2013. Phase 1b, multicenter, single blinded, placebo-controlled, sequential dose escalation study to assess the safety and tolerability of topically applied AG013 in subjects with locally advanced head and neck cancer receiving induction chemotherapy. Cancer 119(24):4268-4276. doi:10.1002/cncr.28365.
Lu, T. K., and J. J. Collins. 2007. Dispersing biofilms with engineered enzymatic bacteriophage. Proc Natl Acad Sci USA 104(27):11197-11202. doi: 10.1073/pnas.0704624104.
Lu, T. K., and J. J. Collins. 2009. Engineered bacteriophage targeting gene networks as adjuvants fornantibiotic therapy. Proc Natl Acad Sci USA 106(12):4629-4634. doi:10.1073/pnas.0800442106.
Lukovac, S., C. Belzer, L. Pellis, B. J. Keijser, W. M. de Vos, R. C. Montijn, and G. Roeselers. 2014. Differential modulation by Akkermansia muciniphila and Faecalibacterium prausnitzii of host peripheral lipid metabolism and histone acetylation in mouse gut organoids. mBio 5(4). doi: 10.1128/mBio.01438-14.
Marchesi, J. R., D. H. Adams, F. Fava, G. D. Hermes, G. M. Hirschfield, G. Hold, M. N. Quraishi, J. Kinross, H. Smidt, K. M. Tuohy, L. V. Thomas, E. G. Zoetendal, and A. Hart. 2016. The gut microbiota and host health: A new clinical frontier. Gut 65(2):330-339. doi: 10.1136/gutjnl-2015-309990.
Mazmanian, S. K., J. L. Round, and D. L. Kasper. 2008. A microbial symbiosis factor prevents intestinal inflammatory disease. Nature 453(7195):620-625. doi: 10.1038/nature07008.
McDonald, J. A., K. Schroeter, S. Fuentes, I. Heikamp-Dejong, C. M. Khursigara, W. M. de Vos, and E. Allen-Vercoe. 2013. Evaluation of microbial community reproducibility, stability and composition in a human distal gut chemostat model. J Microbiol Methods 95(2):167-174. doi: 10.1016/j.mimet.2013.08.008.
Mills, S., F. Shanahan, C. Stanton, C. Hill, A. Coffey, and R. P. Ross. 2013. Movers and shakers: Influence of bacteriophages in shaping the mammalian gut microbiota. Gut Microbes 4(1):4-16. doi:10.4161/gmic.22371.
Mimee, M., A. C. Tucker, C. A. Voigt, and T. K. Lu. 2015. Programming a human commensal bacterium, bacteroides thetaiotaomicron, to sense and respond to stimuli in the murine gut microbiota. Cell Syst 1(1):62-71. doi: 10.1016/j.cels.2015.06.001.
Mimee, M., R. J. Citorik, and T. K. Lu. 2016. Microbiome therapeutics—Advances and challenges. Adv Drug Deliv Rev 105(Pt A):44-54. doi: 10.1016/j.addr.2016.04.032.
Minot, S., R. Sinha, J. Chen, H. Li, S. A. Keilbaugh, G. D. Wu, J. D. Lewis, and F. D. Bushman. 2011. The human gut virome: Inter-individual variation and dynamic response to diet. Genome Res 21(10):1616-1625. doi: 10.1101/gr.122705.111.
Moayyedi, P., M. G. Surette, P. T. Kim, J. Libertucci, M. Wolfe, C. Onischi, D. Armstrong, J. K. Marshall, Z. Kassam, W. Reinisch, and C. H. Lee. 2015. Fecal microbiota transplantation induces remission in patients with active ulcerative colitis in a randomized controlled trial. Gastroenterology 149(1):102-109. doi: 10.1053/j.gastro.2015.04.001.
Modi, S. R., H. H. Lee, C. S. Spina, and J. J. Collins. 2013. Antibiotic treatment expands the resistance reservoir and ecological network of the phage metagenome. Nature 499(7457):219-222. doi: 10.1038/nature12212.
Motta, J. P., L. G. Bermudez-Humaran, C. Deraison, L. Martin, C. Rolland, P. Rousset, J. Boue, G. Dietrich, K. Chapman, P. Kharrat, J. P. Vinel, L. Alric, E. Mas, J. M. Sallenave, P. Langella, and N. Vergnolle. 2012. Food-grade bacteria expressing elafin protect against inflammation and restore colon homeostasis. Sci Transl Med 4(158):158ra144. doi: 10.1126/scitranslmed.3004212.
Nava, G. M., H. J. Friedrichsen, and T. S. Stappenbeck. 2011. Spatial organization of intestinal microbiota in the mouse ascending colon. ISME J 5(4):627-638. doi: 10.1038/ismej.2010.161.
Nicaise, C., D. Prozzi, E. Viaene, C. Moreno, T. Gustot, E. Quertinmont, P. Demetter, V. Suain, P. Goffin, J. Deviere, and P. Hols. 2008. Control of acute, chronic, and constitutive hyperammonemia by wild-type and genetically engineered Lactobacillus plantarum in rodents. Hepatology 48(4):1184-1192. doi: 10.1002/hep.22445.
Norman, J. M., S. A. Handley, M. T. Baldridge, L. Droit, C. Y. Liu, B. C. Keller, A. Kambal, C. L. Monaco, G. Zhao, P. Fleshner, T. S. Stappenbeck, D. P. McGovern, A. Keshavarzian, E. A. Mutlu, J Sauk, D. Gevers, R. J. Xavier, D. Wang, M. Parkes, and H. W. Virgin. 2015. Disease-specific alterations in the enteric virome in inflammatory bowel disease. Cell 160(3):447-460. doi: 10.1016/j.cell.2015.01.002.
Ostrov, N., M. Landon, M. Guell, G. Kuznetsov, J. Teramoto, N. Cervantes, M. Zhou, K. Singh, M. G. Napolitano, M. Moosburner, E. Shrock, B. W. Pruitt, N. Conway, D. B. Goodman, C. L. Gardner, G. Tyree, A. Gonzales, B. L. Wanner, J. E. Norville, M. J. Lajoie, and G. M. Church. 2016. Design, synthesis, and testing toward a 57-codon genome. Science 353(6301):819-822. doi: 10.1126/science.aaf3639.
Petrof, E. O., G. B. Gloor, S. J. Vanner, S. J. Weese, D. Carter, M. C. Daigneault, E. M. Brown, K. Schroeter, and E. Allen-Vercoe. 2013. Stool substitute transplant therapy for the eradication of Clostridium difficile 9 infection: “RePOOPulating” the gut. Microbiome 1(1):3. doi: 10.1186/2049-2618-1-3.
Ratner, M. 2015. Microbial cocktails join fecal transplants in IBD treatment trials. Nat Biotechnol 33(8):787-788. doi: 10.1038/nbt0815-787.
Reardon, S. 2014. Phage therapy gets revitalized. Nature 510(7503):15-16. doi:10.1038/510015a.
Reyes, A., M. Haynes, N. Hanson, F. E. Angly, A. C. Heath, F. Rohwer, and J. I. Gordon. 2010. Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 466(7304):334-338. doi:10.1038/nature09199.
Reyes, A., M. Wu, N. P. McNulty, F. L. Rohwer, and J. I. Gordon. 2013. Gnotobiotic mouse model of phagebacterial host dynamics in the human gut. Proc Natl Acad Sci USA 110(50):20236-20241. doi: 10.1073/pnas.1319470110.
Ritchie, M. L., and T. N. Romanuk. 2012. A meta-analysis of probiotic efficacy for gastrointestinal diseases. PloS One 7(4):e34938. doi: 10.1371/journal.pone.0034938.
Robert, S., C. Gysemans, T. Takiishi, H. Korf, I. Spagnuolo, G. Sebastiani, K. Van Huynegem, L. Steidler, S. Caluwaerts, P. Demetter, C. H. Wasserfall, M. A. Atkinson, F. Dotta, P. Rottiers, T. L. Van Belle, and C. Mathieu. 2014. Oral delivery of glutamic acid decarboxylase (GAD)-65 and IL10 by Lactococcus lactis reverses diabetes in recent-onset NOD mice. Diabetes 63(8):2876-2887. doi: 10.2337/db13-1236.
Shen, T. C., L. Albenberg, K. Bittinger, C. Chehoud, Y. Y. Chen, C. A. Judge, L. Chau, J. Ni, M. Sheng, A. Lin, B. J. Wilkins, E. L. Buza, J. D. Lewis, Y. Daikhin, I. Nissim, M. Yudkoff, F. D. Bushman, and G. D. Wu. 2015. Engineering the gut microbiota to treat hyperammonemia. J Clin Invest 125(7):2841-2850. doi: 10.1172/JCI79214.
Shis, D. L., F. Hussain, S. Meinhardt, L. Swint-Kruse, and M. R. Bennett. 2014. Modular, multi-input transcriptional logic gating with orthogonal LacI/GalR family chimeras. ACS Synth Biol 3(9):645-651. doi: 10.1021/sb500262f.
Siuti, P., J. Yazbek, and T. K. Lu. 2013. Synthetic circuits integrating logic and memory in living cells. Nat Biotechnol 31(5):448-452. doi: 10.1038/nbt.2510.
Sokol, H., B. Pigneur, L. Watterlot, O. Lakhdari, L. G. Bermudez-Humaran, J. J. Gratadoux, S. Blugeon, C. Bridonneau, J. P. Furet, G. Corthier, C. Grangette, N. Vasquez, P. Pochart, G. Trugnan, G. Thomas, H. M. Blottiere, J. Dore, P. Marteau, P. Seksik, and P. Langella. 2008. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc Natl Acad Sci USA 105(43):16731-16736. doi: 10.1073/pnas.0804812105.
Springman, R., I. J. Molineux, C. Duong, R. J. Bull, and J. J. Bull. 2012. Evolutionary stability of a refactored phage genome. ACS Synth Biol 1(9):425-430. doi: 10.1021/sb300040v.
Steidler, L., W. Hans, L. Schotte, S. Neirynck, F. Obermeier, W. Falk, W. Fiers, and E. Remaut. 2000. Treatment of murine colitis by Lactococcus lactis secreting interleukin-10. Science 289(5483):1352-1355.
Steidler, L., S. Neirynck, N. Huyghebaert, V. Snoeck, A. Vermeire, B. Goddeeris, E. Cox, J. P. Remon, and E. Remaut. 2003. Biological containment of genetically modified Lactococcus lactis for intestinal delivery of human interleukin 10. Nat Biotechnol 21(7):785-789. doi: 10.1038/nbt840.
Synlogic. 2017. Synlogic Proprietary Synthetic Biotic Receives FDA Orphan Drug Designation. http://www.synlogictx.com/news/press-releases/synlogic-proprietary-synthetic-biotic-receives-fda-orphan-drug-designation (accessed February 2, 2017).
Takiishi, T., H. Korf, T. L. Van Belle, S. Robert, F. A. Grieco, S. Caluwaerts, L. Galleri, I. Spagnuolo, L. Steidler, K. Van Huynegem, P. Demetter, C. Wasserfall, M. A. Atkinson, F. Dotta, P. Rottiers, C. Gysemans, and C. Mathieu. 2012. Reversal of autoimmune diabetes by restoration of antigen-specific tolerance using genetically modified Lactococcus lactis in mice. J Clin Invest 122(5):1717-1725. doi:10.1172/JCI60530.
Van den Abbeele, P., C. Belzer, M. Goossens, M. Kleerebezem, W. M. De Vos, O. Thas, R. De Weirdt, F. M. Kerckhof, and T. Van de Wiele. 2013. Butyrate-producing Clostridium cluster XIVa species specifically colonize mucins in an in vitro gut model. ISME J 7(5):949-961. doi: 10.1038/ismej.2012.158.
van Nood, E., M. G. Dijkgraaf, and J. J. Keller. 2013. Duodenal infusion of feces for recurrent Clostridium difficile. N Eng J Med 368(22):2145. doi: 10.1056/NEJMc1303919.
Vandenbroucke, K., H. de Haard, E. Beirnaert, T. Dreier, M. Lauwereys, L. Huyck, J. Van Huysse, P. Demetter, L. Steidler, E. Remaut, C. Cuvelier, and P. Rottiers. 2010. Orally administered L. lactis secreting an anti-TNF nanobody demonstrate efficacy in chronic colitis. Mucosal Immunol 3(1):49-56. doi: 10.1038/mi.2009.116.
Varankovich, N. V., M. T. Nickerson, and D. R. Korber. 2015. Probiotic-based strategies for therapeutic and prophylactic use against multiple gastrointestinal diseases. Front Microbiol 6:685. doi:10.3389/fmicb.2015.00685.
Westwater, C., L. M. Kasman, D. A. Schofield, P. A. Werner, J. W. Dolan, M. G. Schmidt, and J. S. Norris. 2003. Use of genetically engineered phage to deliver antimicrobial agents to bacteria: An alternative therapy for treatment of bacterial infections. Antimicrob Agents Chemother 47(4):1301-1307.
Wlodarska, M., A. D. Kostic, and R. J. Xavier. 2015. An integrative view of microbiome-host interactions in inflammatory bowel diseases. Cell Host Microbe 17(5):577-591. doi:10.1016/j.chom.2015.04.008.
Wright, O., M. Delmans, G. B. Stan, and T. Ellis. 2015. GeneGuard: A modular plasmid system designed for biosafety. ACS Synth Biol 4(3):307-316. doi: 10.1021/sb500234s.
Zuccotti, G., F. Meneghin, A. Aceti, G. Barone, M. L. Callegari, A. Di Mauro, M. P. Fantini, D. Gori, F. Indrio, L. Maggio, L. Morelli, and L. Corvaglia. 2015. Italian Society of Neonatology. Probiotics for prevention of atopic diseases in infants: Systematic review and meta-analysis. Allergy 70(11):1356-1371. doi:10.1111/all.12700.
This page intentionally left blank.












