6
Chemical Ecology: The Language of Microbiomes
Mark E. Hay,a,* Deanna S. Beatty,a and Frank J. Stewarta
INTRODUCTION
Chemistry is the language of life. Adequately translating this language allows enhanced insight into ecosystem sustainability and function. Most organisms lack eyes and ears, and so must decide whether to mate with, eat, fight, or escape from other organisms based on chemical information. This mode of perception is the basis for most interactions between microorganisms, but also has advantages that select for strong chemical senses among organisms with vision and hearing (Hay, 2009; Huijbers et al., 2012; Puglisi et al., 2014). Chemical cues are useful when vision is not (e.g., in darkness or when prey or predators are hiding), persist longer than most visual or auditory cues, and often provide more nuanced information regarding predators, competitors, mates, etc. (Hay, 2009). Just as biomedical research has cured disease by understanding chemical cues and signals within cells and tissues, an understanding of chemical ecology provides insight into ways to avoid and cure ecological collapse, such as when coral reefs produce metabolites to defend against consumers as discussed in the next paragraph. Because our understanding of chemical ecology is best developed for macroorganisms that can be manipulated in field experiments, we first provide an overview of how chemically mediated interactions affect populations, communities, and ecosystems of marine macroorganisms, and then show that the same processes structure interactions within marine microbiomes.
CORAL REEFS AS A MACROEXAMPLE
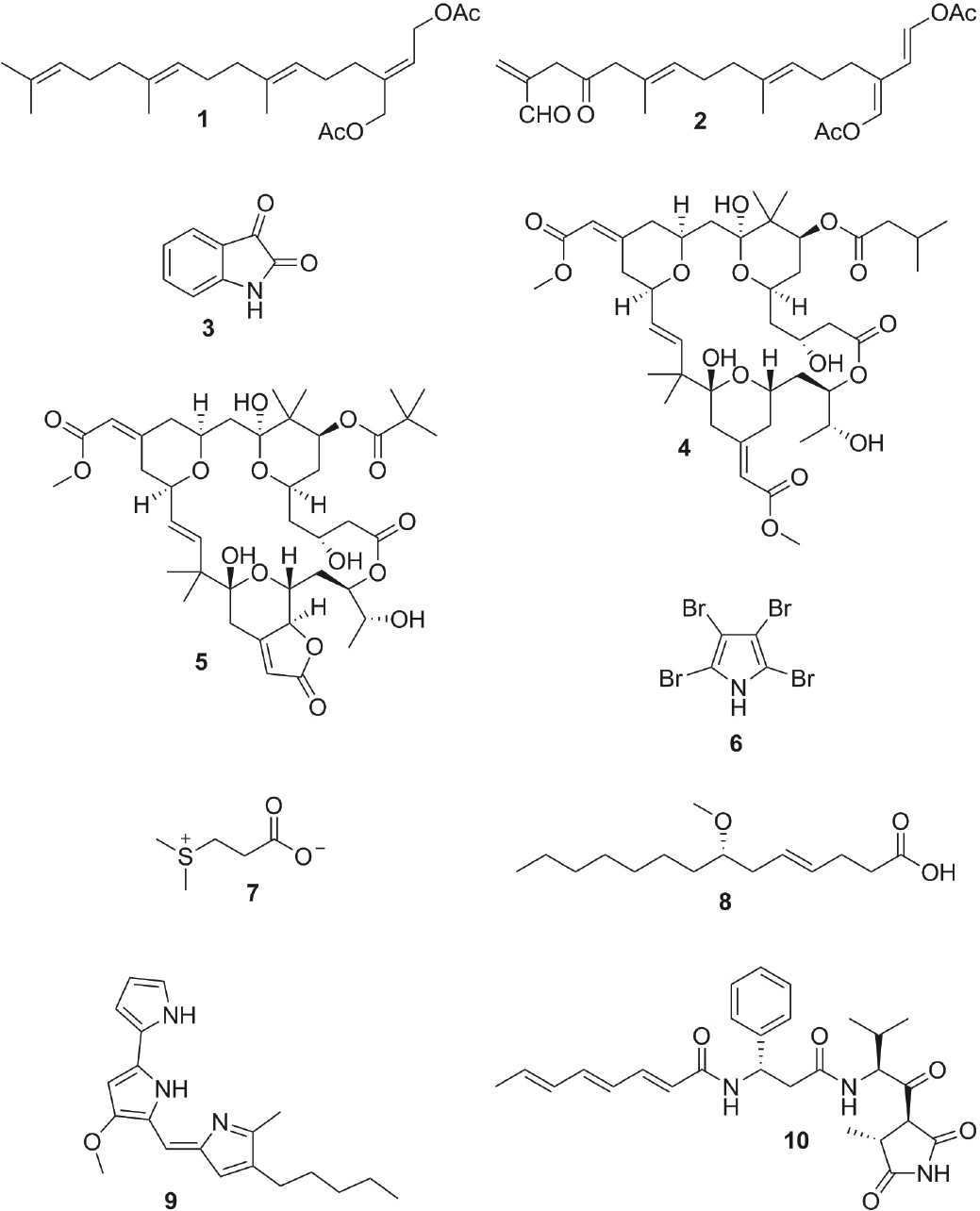
On coral reefs, seaweeds and soft-bodied invertebrates (e.g., sponges, soft corals) commonly produce secondary metabolites that function as defenses against consumers (Hay, 2009; Puglisi et al., 2014) and allelopathic agents against competitors (Rasher et al., 2011; Puglisi et al., 2014). As an example, the green seaweed Chlorodesmis fastigiata produces acetylated diterpenes (see Figure 6-1: compounds 1, 2) that begin to kill corals within days of contact (Rasher et al., 2011). As a countermeasure, the coral Acropora nasuta detects the chemistry of this seaweed within minutes of contact, and sends a chemical signal to mutualistic fishes that trim the seaweed until it no longer touches the coral (Dixson and Hay, 2012). Similar interactions occur at the reef scale. Corals use chemical cues to selectively recruit juvenile fishes to desirable, coral-dominated reefs, while cues from seaweeds are used
___________________
a School of Biological Sciences and Aquatic Chemical Ecology Center, Georgia Institute of Technology.
* Corresponding Author: mark.hay@biology.gatech.edu.
by corals to avoid degraded reefs (Dixson et al., 2014). These fishes and corals are not simply reacting to coral versus seaweed cues, but are responding to chemicals from species that best predict reef health. Thus, chemically mediated behaviors determine consumer–prey and competitive interactions, cue critical mutualisms, determine recruitment patterns, and fundamentally alter the stability and resilience of coral reefs. Given that microbes lack well-developed vision and hearing, chemical cues and signals likely play even larger roles within microbiomes.
THE CHEMICAL ECOLOGY OF MARINE MICROBIOMES
Though not yet broadly recognized, most of microbial ecology is chemical ecology. Integrating these fields is challenging due to the difficulty of rigorously exploring chemically mediated interactions among microbes under natural conditions. Many marine microbes are not yet culturable, especially those living in association with specific hosts. Furthermore, the behavior of cultured microbes may not reflect behaviors in natural multispecies microbiomes. Microbes may express chemical or other traits only in response to environmental cues that may be missing under culture conditions (Moree et al., 2012). As an example, the phytoplankton Phaeocystis globosa chemically senses conspecifics being attacked, identifies the attacker as a ciliate or copepod via chemical cues, and alters its traits to reduce susceptibility to that consumer (Long et al., 2007). Similarly, many microbes thrive only in a biodiverse community of species, which often cannot be replicated in culture. For example, microbial degradation of complex hydrocarbons involves a cascade of interacting microbes that consume the metabolites of their neighbor as the hydrocarbon is successively degraded (McGenity et al., 2012). Thus, insights from simplified culture conditions may not apply to the more complex conditions in nature (Lopanik, 2014). Despite these challenges, an understanding of microbiome chemical ecology is developing (Weitz et al., 2013).
Microbes commonly produce secondary metabolites that defend their hosts (Lopanik, 2014). Embryos of the shrimp Palaemon macrodactylus are remarkably resistant to fungal attack because they are covered by an Alteromonas sp. bacterium that produces 2,3-indolinedione (isatin; see Figure 6-1, compound 3), which suppresses pathogenic fungi (Gil-Turnes et al., 1989). Embryos stripped of bacteria using antibiotics die within 4 days due to fungal attack. In contrast, survival is high for control embryos not treated with antibiotics, treated with antibiotics but then reexposed to the bacterium, or treated with antibiotics and exposed only to isatin.
The marine bryozoan Bugulia neritina provides another example (Lopanik, 2014). Bioassay-guided fractionation demonstrates that bryostatins 10 and 20 (see Figure 6-1, compounds 4, 5) defend B. neritina larvae from predators. These compounds are produced by a γ-Proteobacterium, Candidatus Endobugula sertula, associated with the larvae. Stripping Buguila of bacteria using antibiotics causes concentrations of bryostatins to decline by 99%, and larvae to become palatable.
As a final example, on reefs in Papua New Guinea, bright red isopods live in conspicuous groups on reef surfaces exposed to fish predators (Lindquist et al., 2005). The red coloration is from cyanobacteria covering their carapace, and fish reject them as food. Isopods kept in the dark for two days supported fewer cyanobacteria and became palatable to fishes. Chemical extracts of normal, cyanobacteria-covered isopods deterred fish feeding, suggesting that cyanobacterial metabolites defended the isopods.
Microbially produced chemical defense of hosts is also suggested for ascidians, sponges, fish, squid egg capsules, seaweeds, corals, and a host of other marine organisms (Piel, 2009; Weitz et al., 2013; Lopanik, 2014). Although chemical cues from hosts likely affect microbial colonization (McFall-Ngai, 2014), it is rare that we understand how mutualistic microbes are recruited, maintained, cued to produce appropriate metabolites, or prevented from being displaced by nonmutualistic microbes. As methods, concepts, and appreciation for the omnipresent importance of microbiomes become better developed, we anticipate dramatic growth in our understanding of microbiome ecology, and the chemical mediation of microbiome organization and function. Below, we use corals as an example of likely challenges and opportunities.
THE ORGANIZATION AND FUNCTION OF CORAL MICROBIOMES
Corals have historically been understood as a mutualistic association between invertebrate animals and dinoflagellates, which provide much of the corals’ energetic needs via photosynthesis. More recently, corals have been
recognized as a complex mutualism between the coral animal, Symbiodinium, and a diverse assemblage of bacteria and archaea. Microbes play critical roles from birth to death in corals. At the earliest stages, the larvae of some corals acquire commensal microbes from the parent, while larvae of other species acquire microbes only from the environment (Sharp and Ritchie, 2012). Following a planktonic dispersal phase, larvae of several coral species preferentially settle on certain species of crustose coralline algae (CCA). Pseudoaltermonas bacteria growing on CCA produce tetrabromopyrrole (see Figure 6-1, compound 6), which induces metamorphosis and larval settlement in numerous corals (Sneed et al., 2014; Thompson et al., 2015). Once settled, corals develop a microbiome whose members aid in coral nutrition and produce antibiotics to defend against harmful microbes (Ritchie, 2006). For example, coral bacteria of the genus Exiguobacterium produce a small molecular weight compound(s) that reduces growth of the coral pathogen Serratia marcescens by inhibiting catabolism of coral mucus (Krediet et al., 2013). Hydrophobic compounds on coral surfaces also inhibit biofilm formation of S. marcescens, and bacteria from healthy coral reproduce this activity, as well as suppress pathogenic S. marcescens (Alagely et al., 2011). Thus, microbiomes play vital roles from recruitment through senescence.
When corals are stressed by warming, competition from seaweeds, or other factors, the corals’ defensive microbiome can become destabilized, leading to “dysbiosis,” and in some cases loss of defenses (Ritchie, 2006; Barott and Rohwer, 2012). Dysbiosis is chemically mediated. For example, heat-stressed corals produce excess dimethylsulfoniopropionate (see Figure 6-1, compound 7), and pathogens such as Vibrio coralliilyticus chemotax this compound (Garren et al., 2014). Dissolved organic carbon released by nearby seaweeds can also selectively stimulate the growth of microbes enriched in virulence factors (Nelson et al., 2013). Several examples show that once harmful microbes escape control within the microbiome, they may chemically attack the coral (Puglisi et al., 2014). Notably, lyngbic acid–producing cyanobacteria dominate the polymicrobial consortium that causes black band disease (BBD). Lyngbic acid (see Figure 6-1, compound 8) inhibits quorum sensing and is associated with a loss of commensals and an increase in coral microbiome diversity (Meyer et al., 2016). Serratia marcescens, which causes white pox disease, also produces compounds with antimicrobial properties. For example, the pigment prodigiosin (see Figure 6-1, compound 9), which gives S. marcescens its red color, acts against a broad range of bacteria, potentially poisoning the corals’ mutualistic microbes, and allows S. marcescens exclusive access to the coral resource. Similarly, Vibrio coralliilyticus produces the antibacterial compound andrimid (see Figure 6-1, compound 10), which may aid in this bacterium’s dominance in diseased corals. These, and other examples, are based largely on compounds investigated from pure cultures. The natural functions of these compounds in coral microbiomes, where microbes generally occur at lower densities and within a polyculture, remain inadequately understood.
CHALLENGES AND OPPORTUNITIES
A major challenge for marine microbiome research is that we know too little about the functional roles of microbes within complex communities, the chemical mechanisms operating within natural microbiomes, and the ecological significance of microbiome compositional change. For example, a change in microbiome composition is often interpreted as dysbiosis when it might be a fitness-enhancing acclimation to altered environmental conditions. Egan and Gardiner provide an overview of conceptual and methodological challenges (Egan and Gardiner, 2016). First, how do we separate cause from effect and differentiate between the causative pathogens versus the opportunistic microbial invaders of dead or decaying tissues? Causative agents may be early invaders associated with an asymptomatic host state, whereas the disease state is associated with opportunistic secondary invaders or “detritivores.” Second, for several diseases there may be no single causative agent, but rather a consortium of agents, potentially with complex interacting chemical profiles; for example, BBD involves cyanobacteria, archaea, sulfur-cycling, and heterotrophic bacteria. Third, mutualistic or commensal bacteria may become pathogenic when the host is stressed; therefore, the chemical ecology of the interacting microbes is likely context dependent. Finally, the conditional switch from mutualist to enemy may interact with host genetics, immune responses, and environmental factors to cause dysbiosis that is expressed as a disease. Given these issues, we need to understand host–microbiome chemistry, and variations thereof, over a broad range of community states and environmental factors.
Most of the focus on interactions within microbiomes has been on dysbiosis, competition, or mutualism
as opposed to predation, parasitism, or other interactions. Studies of many communities find strong effects of predation (Estes et al., 2011; Ohgushi et al., 2012), and of pathogens minimally impacting a vectoring species but having large impacts on other species (Parker et al., 2015). Such interactions likely affect microbiomes as well. For example, the predatory bacterium Halobacteriovorax occurs in 80% of some coral microbiomes and consumes the coral pathogens Vibrio corallyticus and V. harveyii (Webster et al., 2013). Eating pathogens may be as effective as poisoning them, and both interactions are probably chemically mediated. Similarly, some corals might use biological warfare by vectoring microbes that cause disease in other corals. As a possible example, Campylobacteraceae bacteria occur on healthy Acropora, despite this group often being associated with disease in corals such as Montastrea (Chu and Vollmer, 2016). Could these bacteria advantage Acropora by infecting and suppressing neighboring Montastrea?
Finally, if the suggested, but often undemonstrated, chemically mediated interactions structuring microbiomes and microbe–host interactions are constrained by environmental conditions, it is possible that global change will destabilize this chemical language. As an example, coral settlement cues from CCA-associated microbes are lost under ocean acidification due to shifts in microbial communities (Welsh et al., 2016). Thus, ocean acidification and warming could compromise chemical information gathering, altering the organization and stability of communities.
The importance of microbiomes as mediators of individual health, population regulation, community structure, and ecosystem function is becoming increasingly clear (Welsh et al., 2016). Given their sensory modalities, the major interactions within microbiomes, and between microbiomes and their hosts, must be chemically mediated. Finding, carefully describing, and correctly interpreting the ecological and evolutionary insights that can be gained from this chemical Rosetta stone to microbial language will be a great challenge, but also a tremendous opportunity to gain insight into the players that structure much of our biotic world.
REFERENCES
Alagely, A., C. J. Krediet, K. B. Ritchie, and M. Teplitski. 2011. Signaling-mediated cross-talk modulates swarming and biofilm formation in a coral pathogen Serratia marcescens. ISME J 5:1609-1620.
Barott, K. L., and F. L. Rohwer. 2012. Unseen players shape benthic competition on coral reefs. Trends Microbiol 20:621-628.
Chu, N. D., and S. V. Vollmer. 2016. Caribbean corals house shared and host-specific microbial symbionts over time and space. Environ Microbiol Rep 8:493-500.
Dixson, D. L., and M. E. Hay. 2012. Corals chemically cue mutualistic fishes to remove competing seaweeds. Science 338:804-807.
Dixson, D. L., D. Abrego, and M. E. Hay. 2014. Reef ecology. Chemically mediated behavior of recruiting corals and fishes: A tipping point that may limit reef recovery. Science. 345:892-897.
Egan, S., and M. Gardiner. 2016. Microbial dysbiosis: Rethinking disease in marine ecosystems. Front Microbiol 7:991.
Estes, J. A., J. Terborgh, J. S. Brashares, M. E. Power, J. Berger, W. J. Bond, S. R. Carpenter, T. E. Essington, R. D. Holt, J. B. Jackson, R. J. Marquis, L. Oksanen, T. Oksanen, R. T. Paine, E. K. Pikitch, W. J. Ripple, S. A. Sandin, M. Scheffer, T. W. Schoener, J. B. Shurin, A. R. Sinclair, M. E. Soulé, R. Virtanen, and D. A. Wardle. 2011. Trophic downgrading of planet Earth. Science 333:301-306.
Garren, M., K. Son, J. B. Raina, R. Rusconi, F. Menolascina, O. H. Shapiro, J. Tout, D. G. Bourne, J. R. Seymour, and R. Stocker. 2014. A bacterial pathogen uses dimethylsulfoniopropionate as a cue to target heat-stressed corals. ISME J 8:999-1007.
Gil-Turnes, M. S., M. E. Hay, and W. Fenical. 1989. Symbiotic marine bacteria chemically defend crustacean embryos from a pathogenic fungus. Science 246:116-118.
Hay, M. E. 2009. Marine chemical ecology: Chemical signals and cues structure marine populations, communities, and ecosystems. Annu Rev Mar Sci 1:193-212.
Huijbers, C. M., I. Nagelkerken, P. A. Lössbroek, I. E. Schulten, A. Siegenthaler, M. W. Holderied, and S. D. Simpson. 2012. A test of the senses: Fish select novel habitats by responding to multiple cues. Ecology 93:46-55.
Krediet, C. J., K. B. Ritchie, V. J. Paul, and M. Teplitski. 2013. Coral-associated micro-organisms and their roles in promoting coral health and thwarting diseases. Proc R Soc B Lond [Biol] 280:20122328.
Lindquist, N., P. H. Barber, and J. B. Weisz. 2005. Episymbiotic microbes as food and defence for marine isopods: Unique symbioses in a hostile environment. Proc R Soc B 272:1209-1216.
Long, J. D., G. W. Smalley, T. Barsby, J. T. Anderson, and M. E. Hay. 2007. Chemical cues induce consumer-specific defenses in a bloom-forming marine phytoplankton. Proc Natl Acad Sci USA 104:10512-10517.
Lopanik, N. B. 2014. Chemical defensive symbioses in the marine environment. Funct Ecol 28:328:340.
McFall-Ngai, M. J. 2014. The importance of microbes in animal development: Lessons from the squid-vibrio symbiosis. Annu Rev Microbiol 68:177-194.
McGenity, T. J., B. D. Folwell, B. A. McKew, and G. O. Sanni. 2012. Marine crude-oil biodegradation: A central role for interspecies interactions. Aquat Biosyst 8:10.
Meyer, J. L., S. P. Gunasekera, R. M. Scott, V. J. Paul, and M. Teplitski. 2016. Microbiome shifts and the inhibition of quorum sensing by black band disease cyanobacteria. ISME J 10:1204-1216.
Moree, W. J., V. V. Phelan, C. H. Wu, N. Bandeira, D. S. Cornett, B. M. Duggan, and P. C. Dorrestein. 2012. Interkingdom metabolic transformations captured by microbial imaging mass spectrometry. Proc Natl Acad Sci USA 109:13811-13816.
Nelson, C. E., S. J. Goldberg, K. L. Wegley, A. F. Haas, J. E. Smith, F. Rohwer, and C. A. Carlson. 2013. Coral and macroalgal exudates vary in neutral sugar composition and differentially enrich reef bacterioplankton lineages. ISME J 7:962-979.
Ohgushi, T., O. Schmitz, and R. D. Holteds. 2012. Trait-Mediated Indirect Interactions: Ecological and Evolutionary Perspectives. New York: Cambridge University Press.
Parker, I. M., M. Saunders, M. Bontrager, A. P. Weitz, R. Hendricks, Magarey, K. Suiter, and G. S. Gilbert. 2015. Phylogenetic structure and host abundance drive disease pressure in communities. Nature 520:542-544.
Piel, J. 2009. Metabolites from symbiotic bacteria. Nat Prod Rep 21:519-538.
Puglisi, M. P., J. M. Sneed, K. H. Sharp, R. Ritson-Williams, and V. J. Paul. 2014. Marine chemical ecology in benthic environments. Nat Prod Rep 11:1510-1553.
Rasher, D. B., E. P. Stout, S. Engel, J. Kubanek, and M. E. Hay. 2011. Macroalgal terpenes function as allelopathic agents against reef corals. Proc Natl Acad Sci USA 108:17726-17731.
Ritchie, K. B. 2006. Regulation of microbial populations by coral surface mucus and mucus-associated bacteria. Mar Ecol Prog Ser 322:1-14.
Sharp, K. H., and K. B. Ritchie. 2012. Multi-partner interactions in corals in the face of climate change. Biol Bull 223:66-77.
Sneed, J. M., K. H. Sharp, K. B. Ritchie, and V. J. Paul. 2014. The chemical cue tetrabromopyrrole from a biofilm bacterium induces settlement of multiple Caribbean corals. Proc R Soc B 281:20133086.
Thompson, J. R., H. E. Rivera, C. J. Closek, and M. Medina. 2015. Microbes in the coral holobiont: Partners through evolution, development, and ecological interactions. Front Cell Infect Microbiol 4:176.
Webster, N. S., S. Uthicke, E. S. Botte, F. Flores, and A. P. Negri. 2013. Ocean acidification reduces induction of coral settlement by crustose coralline algae. Glob Change Biol 19:303-315.
Weitz, M., K. Duncan, N. V. Patin, and P. R. Jensen. 2013. Antagonistic interactions mediated by marine bacteria: The role of small molecules. J Chem Ecol 39:879-891.
Welsh, R. M., J. R. Zaneveld, S. M. Rosales, J. P. Payet, D. E. Burkepile, and R. V. Thurber. 2016. Bacterial predation in a marine host-associated microbiome. ISME J 10:1540-1544.