6- Towards Data Attribution and Citation in the Life Sciences
Philip Bourne1
University of California at San Diego
My talk today will focus on some observations about data citation and attribution from the life sciences perspective. The Protein Data Bank (PDB) is a data repository that I am involved with and I am going to use it an example of some of the things that are happening with data in the life sciences.
Let me start with the following observation. In terms of life sciences data repositories, the National Library of Medicine (NLM) is one of the largest in the field. In many ways, the NLM has done a very good job of providing data resources. It also allows people to deposit data and has some level of integration with the literature. However, it appears to me that what they are doing is not fully consistent with some of the data citation and attribution principles and best practices that have been discussed in this workshop so far.
For example, the ability to cite and attribute the data at the NLM is highly variable:
• Digital Object Identifiers (DOIs) are assigned in some cases, but are not used.
• Attribution is made through the metadata in most cases.
• Such attribution is typically through the associated literature reference, if it exists, or by a database identifier.
• The use of data repositories such as Dryad is compelling for the “long tail” problem, but not recognized by NLM.
• Data journals are on the horizon.
There is an interplay between data and publication that is very interesting in the life sciences. To people that maintain data repositories, their metric of success is being published in Nucleic Acids Research, since it brings prominence to these data resources through traditional publication. Similarly, I am an author of a paper about the PDB, which is cited yet I can guarantee that few people have ever read it. There is no reason to read it. It is just there to provide a conventional value metric for a database.
The PDB is a resource that is distributing worldwide the equivalent to one quarter of the Library of Congress each month. The PDB is one of the oldest data repository in biology, a 1 TB resource that is used by approximately 280,000 individuals per month, including an increasing number of school kids. There is absolutely no way that when this resource was founded over forty years ago that scientists ever believed that school children would be using that data.
______________________
1 Presentation slides are available at http://sites.nationalacademies.org/PGA/brdi/PGA064019.
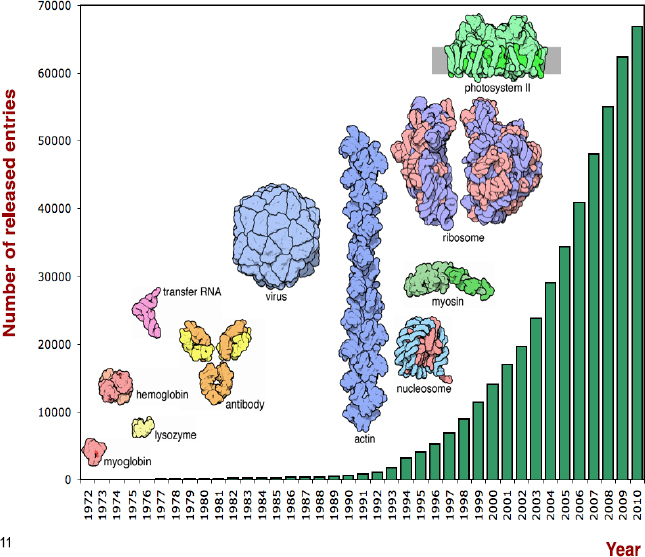
FIGURE 6-1 Protein data bank repository growth over time.
The main point in this diagram is that PDB data volumes are increasing and that the complexity of the data is increasing dramatically as well. Therefore, how we define data, how we cite them, and what granularity we use is changing all the time.
Another point I would like to make is that people are doing more with the data. The following is our big metric of success that we use when we go to funding agencies. We have grown the user base considerably and people are spending more time working with the data than they ever did before. For example, statistics show that the number of visits and page views is growing faster than the number of unique visitors, implying that users are spending more time on the site.
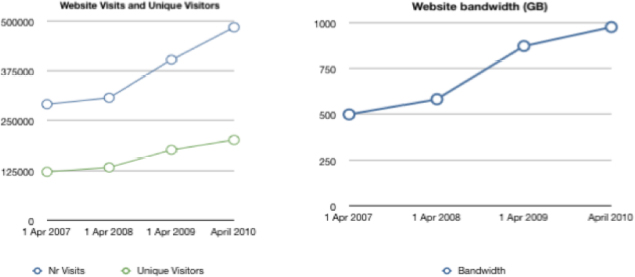
FIGURE 6-2 Web site visitors and bandwidth.
Then there is the question of what are valuable data and what are not? Figure 6-3 provides an example related to H1N1 pandemic data. Effectively, associated PDB data were hardly used at all but suddenly, during that pandemic, the data became highly accessed. We cannot really tell when data are going to be valuable, so, how do we decide what to keep and what not to keep?
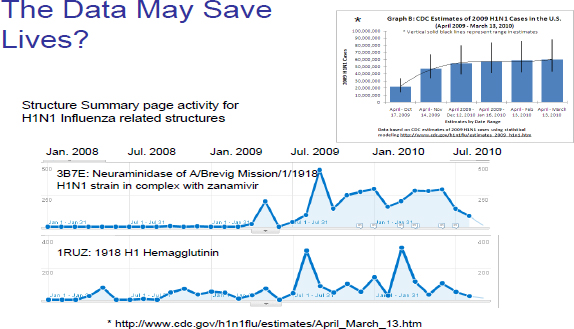
FIGURE 6-3 Data related to the H1N1 pandemic.
Source: Centers for Disease Control.
Let me now talk about data attribution and citation. We spend about 25 percent of the PDB’s budget on remediating data that we already have. This has introduced issues related to our support of multiple versions, which we also have handled. There is the so-called copy of record, which is the version that actually went with the publication which we also always make
available. The community of depositors of the data also requested that scientists cannot publish their articles unless the supporting data are deposited.
We do cite DOIs but, as I said earlier, no one uses them. Database identifiers are preferred. This exemplifies the kind of problems that we are facing and I think it is a sociological matter. Take the example below of a molecule. It is a receptor on the cell surface called CD4. It is important for many reasons, but one of them is that there is a protein on the HIV virus that binds to it and so it has been identified as the causal agent of HIV infection. When this structure came out, we were not using DOIs.
FIGURE 6-4 Image of 1CD4.
Figure 6-4 is known in the biology community everywhere as CD4 because this is the document identifier that it got when it was assigned by a group of scientists at Yale University. That is the identifier we use for character identifiers in the PDB. So what happened is that a group of scientists from Columbia University did a better job on this dataset and then called it in the database 2CD4. The Yale group then went back and did some more work. As a result, they created another dataset and called it 3CD4. But the problem was that 3CD4 had already been given to someone else.
This actually caused angst in the community and it has absolutely no relevance, whatsoever. It is just an identifier.
If there were different sources of those data and we could not be clear on what the copy of record was, it would be a problem but, in this case, it is quite clear. So, rather than thinking about data and journals, it is the idea of trying to bring all this together into a seamless connection. The journal is just one view on the data in some ways and we have been working on this with a number of journals.
Previously, when people did not cite the data, the authors could not use a standard mechanism to find out who has been using the dataset. Now they can because we scan all the open access literature. We can point out to people all of the references that a particular piece of data has in
the open access literature. What is even more interesting is that we can then see in each of those articles what else is being cited.
This page intentionally left blank.






